Genetic and Phenotypic Variation of Calonectria pseudonaviculata Isolates Causing Boxwood Blight Disease in the Hyrcanian Forest of Iran
Parisa Khazaeli1*, Saeed Rezaee1, Mansoureh Mirabolfathy2, Hamidreza Zamanizadeh1 and Hadi Kiadaliri3
1Department of Plant Pathology, College of Agriculture and Natural Resources, Science and Research Branch, Islamic Azad University, Iran
2Iranian Research Institute of Plant Protection, Agricultural Research Education and Extension Organization (AREEO), Iran
3Department of Forestry, College of Agriculture and Natural Resources, Science and Research Branch, Islamic Azad University, Iran
Submission: October 06, 2018; Published: December 12, 2018
*Corresponding author: Parisa Khazaeli, Department of Plant Pathology, College of Agriculture and Natural Resources, Science and Research Branch, Islamic Azad University, Tehran, Iran.
How to cite this article: Parisa K, Saeed R, Mansoureh M, Hamidreza Z, Hadi K. Genetic and Phenotypic Variation of Calonectria pseudonaviculata Isolates Causing Boxwood Blight Disease in the Hyrcanian Forest of Iran. Agri Res& Tech: Open Access J. 2018; 19(1): 556081. DOI: 10.19080/ARTOAJ.2018.19.556081
Abstract
In the last decade, an outbreak of boxwood (Buxus sempervirens subsp. hyrcana) blight disease caused by the pathogen Calonectria pseudonaviculata has resulted in death and decline of boxwood across 52,000 hectares of the Hyrcanian forest in northern Iran. It is necessary when attempting to control the disease to determine the phenotypic and genetic variation, and the virulence of Iranian Calonectria pseudonaviculata isolates. The northern boxwood forests located in Mazandaran, Guilan and Golestan provinces were monitored from June to September of 2014 and isolates of C. pseudonaviculata collected. Morphological characters of conidia, conidiophores, colony patterns and radial growth rates of 75 isolates were recorded and the virulence of 35 selected isolates was evaluated for their pathogenicity in the greenhouse. Partial btub, CaM, and H3 genes of the Iranian C. pseudonaviculata isolates were sequenced. Boxwood blight disease was observed in all surveyed boxwood forests located in northern Iran and C. pseudonaviculata isolates were obtained from all surveyed areas. A wide variation of virulence was recorded among the tested isolates (range of 0.2-39.2%) on B. sempervirens subsp. hyrcana. The most virulent isolate (MN107) was collected from Neka, Mazandaran province. Multilocus sequencing of Iranian C. pseudonaviculata isolates placed all isolates in one group.
Keywords: Calonectria pseudonaviculata; Hyrcanian forest; Genetic variation
Abbrevations: PAUP: Phylogenetic Analysis Using Parsimony; TBR: Tree-Bisection-Reconnection; DSI: Disease Severity Index; GPS: Global Positioning System; H3: Histone; TL: Tree Length; CI: Consistency Index; RI: Retention Index; RC: Rescaled Consistency Index; HI: Homoplasy Index; MCMC: Markov Chain Monte Carlo; ML: Maximum Likelihood; TUB: ß-tubulin; HMG: High Mobility Group; CaM: Calmodulin; MP: Maximum Parsimony; BI: Bayesian Inference; PCA: Potato Carrot Agar
Introduction
Boxwood (Buxus sempervirens subsp. hyrcana) trees are evergreen trees and shrubs endemic to the Hyrcanian forest areas of northern Iran growing in Guilan, Mazandaran and Golestan provinces (Figure 1). Boxwood is considered a unique and genetically valuable forest resource for Iran [1]. Boxwood can be damaged by several diseases but the most severe of these is Boxwood blight, causing widespread destruction in Iran and on forest trees or cultivars in many countries throughout Europe and in the Asia Pacific, and in numerous states of the USA [2]. Boxwood blight symptoms were first observed in a nursery in Hampshire, England in 1994 [3]. Several years later similar symptoms were observed on Buxus in New Zealand [4]. The fungal pathogen causing Boxwood blight was originally described in the literature as Cylindrocladium pseudonaviculatum [5], and soon after as Cylindrocladium buxicola [6]. It is now most correctly known by the teleomorph name Calonectria pseudonaviculatum. Boxwood blight disease was first discovered in Iran in 2010 and the causal agent confirmed in 2012 from the forests of Mazandaran and Guilan provinces [7-9]. Two years later the disease had spread to the Eastern Golestan province of the Hyrcanian forest [10]. Molina et al. [11], estimated that by 2014 approximately 40,000 hectares of boxwood trees had been infected in northern Iran and it is estimated that this could now be closer to 52,000 hectares of the total extent of approximately 70,000 hectares.
The genetic diversity of 17 isolates of C. pseudonaviculata from the UK and one isolate from New Zealand have been analyzed using AFLP, and results showed that the isolates collected from the UK and New Zealand are genetically homogenous [6]. Genetic diversity of a collection including 234 Calonectria isolates from 15 countries and four continents using multilocus phylogenetic analysis indicated that there are two genetic clades (G1 and G2). The isolates in the G1 clade were identified as C. pseudonaviculata sensu stricto and those in the G2 clade identified as C. henricotiae sp. nov. [2].
To better understand the movement of boxwood blight throughout all Hyrcanian forests during 2013-2017, we conducted a comprehensive review of genetic and phenotypic diversity of C. pseudonaviculata collected from the Hyrcanian forest of Iran. The genetic diversity of Iranian C. pseudonaviculata isolates was determined using multilocus sequencing, β-tubulin (βtub), Calmodulin (CaM) and Histone (H3) genes, and their pathogenicity tested on boxwood saplings in the greenhouse.
Materials and Methods
Sample collection and fungal isolates
From June to September of 2014, 254 samples were collected from infected seedlings, bushes and boxwood trees throughout the Hyrcanian forest across different altitudes and climatic regions and the location recorded using a handheld Global Positioning System (GPS). A total of 254 samples were collected from 75 locations across three provinces (Figure 1): Mazandaran (Liresar, Sisangan, Chalandar, Toskatook, Kohnesara, Neka, Sorkhkola, Afrachal, Savadkooh, Chaeebagh, Vachat, Ghaemshahr, Amol, Behshahr), Guilan (Astara, Kalat, Gisoom, Roodsar, Siahkal, Bijarkenar) and Golestan (Bandargaz-Livan sharghi). Infected plant tissues collected were surface sterilized using sodium hypochlorite (1% concentration) for 1 min and then washed in sterile water. The pure isolates were cultured on Potato Dextrose Agar (PDA; Merck, Germany) and Malt Extract Agar (MEA; Merck, Germany) media using the single spore method [12].

Morphological characterization
The colony morphology and growth rate of 75 isolates were studied on PDA and MEA at 25 °C for 10 days. For spore production the isolates were grown on Potato Carrot Agar PCA (carrot pieces 20g, potato pieces 20g, agar 20g per liter) under near-UV light at 25 °C. Morphological characteristics of 20 isolates recorded included length, shape and the number of septa of conidia. The length and vesicle characterization of the stipe were observed using a compound microscope (Olympus CX21) and measured using a software image analyzer system (Leica Qwin) with 40 measurements obtained per structure.
Growth pattern and growth rate
All isolates were grown at 25 and 30°C as these are considered the most appropriate temperatures to distinguish C. pseudonaviculata from closely related species. In addition, the growth rate of 20 representative isolates from the Hyrcanaian forest were examined at 5, 10, 15, 20, 25, 28 and 30 °C. All isolates were created for these measurements by placing a 5 mm diameter agar plug from 7-day-old colonies on to the centre of a 90mm PDA plate. Three replicates for each isolate were produced and diameters of each colony were measured after 7 days.
Pathogenicity assay
The pathogenicity assay of the isolates was conducted in greenhouse conditions using 35 representative isolates chosen based on geographical coordinates and altitude of the sampling location, and variation in growth rate.
Six-year-old healthy boxwood saplings approximately 40- 50cm high were supplied from the Sisangan nursery forest in Mazandaran province. Saplings were replanted in 3 litre pots filled with sterilized forest soil which was prepared by filling an oven container with the moistened forest soil and setting the temperature in the range of 85-90 °C for 30 minutes and the temperature of the centre of the soil pile was checked to reach 80 °C. To test whether the plants had been previously infected with C. pseudonaviculata the saplings were kept in conditions favorable for development of disease symptoms (95% relative humidity and 23-25 °C) for two weeks. A total of 108 saplings were selected for this experiment.
Three replicates of 6-year-old plants were inoculated with each isolate in a completely randomized design. Inoculation was performed by dipping leaves into a conidial suspension (2×104conidia/ml) of C. pseudonaviculata for less than 1min and three control plants were treated by dipping with sterilized distilled water [13]. Inoculated plants were covered with transparent plastic bags and a piece of wet cotton inserted inside the plastic bag to provide saturated humidity (95% relative humidity) at a temperature range of 23 °C to 25 °C for 21 days in greenhouse. Disease severity was evaluated according to Peever et al. [14], using the Disease Severity Index (DSI) as follows: 0 = without disease symptoms, 1= 1-25% leaf area covered with lesions, 2= 25-50% leaf area covered with lesions, 3= 50-75% leaf area covered with lesions, 4= 75-100% leaf area covered with lesions (at 100% all leaves are dead).
The Disease Severity Index (%) was determined according to Wheeler, 1969 [15], and data were analyzed using one-way ANOVA and the means were compared using Duncan’s multiple range test, at P = 0.01 level of significance. All statistical analyses were done using SPSS (version 16.0). Ryan–Joiner test in Minitab v.16.2.4. was used to evaluate the normality and homogeneity of the data, respectively. Data where necessary were normalized by using the formula √(x-1).
DNA extraction
Fungal isolates were cultured on MEA medium and incubated for 7 days at 25 °C, and mycelium scraped from the plates and used for DNA extraction. The mycelium was ground in liquid nitrogen and the DNA extracted using the CTAB method with some modifications [16]. The powdered mass was mixed with pre-heated (62 °C) 600ml of CTAB extraction buffer (10mM Tris HCI pH 8.0, 20mMEDTA, 1.4 M NaCI and 2% CTAB) per tube and incubated at 62 °C for 45min in a water bath with occasional mixing by gently inverting the tube. 2μl of 10mg/ml Proteinase K was added and incubated at 37 °C for 30 min. 700μl of chloroformisoamyl alcohol (24:1) was added, invert for 10min and the tubes were centrifuged at 13000rpm for 10 min. The aqueous phase was transferred to another tube and 2μl RNAse was added, incubated at 37 °C for 30 min. 400μl of chloroform-isoamyl alcohol (24:1) was added, inverted for 10 min and the tubes were centrifuged at 13000rpm for 10 min. The aqueous phase was transferred to another tube and 180μl cold isopropanol was added, incubated for 15min at -20 °C and after that centrifuged at 13000rpm for 10min. The supernatant was discarded, and the pellet was suspended with 70% ethanol and then centrifuged at 13000rpm for 10min and two ethanol washes were given and centrifuged for 10min at 13000rpm. The pellet thus obtained was air-dried and sterile nuclease free water was added to dissolve the DNA and stored at -4 °C for further study. DNA concentration was quantified using a Nano Drop ND-1000 spectrophotometer (Isogen Life Science, Netherlands).
Multilocus DNA sequencing
Partial sequences of β-tubulin (βtub), Calmodulin (CaM), and Histone H3 genes of 20 representative isolates of C. pseudonaviculata were selected based on variation in colony patterns, growth rate at optimum temperature, pathogenicity and altitude of sample locations (Table 2). These partial sequences were amplified by the Bt2a/Bt2b, Cal228F/Cal737R and H3-1A/ H3-1B primer pairs, respectively [17,18].
Reactions were performed in a total volume of 25μl containing 2.5μl 10XPCR buffer (Fermentas, Germany), 0.5μl of dNTP (10mM), 0.75μl MgCl2 (50mM), 1μl of each primer (10pmol/ml), 0.25μlU Taq DNA Polymerase (5unit/ml) (Fermentas, Germany) and 50ng genomic DNA. DNA amplification was performed with a BioRad thermocycler, with an initial denaturation step of 94 °C for 2min followed by 35 cycles of 94 °C for 1min., annealing temperature for 30s, 72 °C for 30min and a final extension at 72 °C for 5min. Amplicons were sequenced in both directions using the amplification primers and the Sanger sequencing method at Macrogen (Seoul, Korea).
Phylogenetic analysis
The software program DNASTAR’s SeqMan NGen was used for editing and assembling the sequence files. Single locus data sets and a combination of multilocus data sets of three gene sequence regions of the studied isolates were recovered, and these were compared with sequences retrieved from GenBank by aligning using CLUSTALX2 [19]. Phylogeny of the combined aligned data sets of sequences was analyzed by Bayesian Inference (BI), Maximum Likelihood (ML) and Maximum Parsimony (MP). Suitable models for the Bayesian analysis were first selected using models of nucleotide substitution for each gene, as determined using Mr Modeltest version 3.6 [20]. The Bayesian analyses [Mr Bayes v. 3.2.1] [21], were run from random trees for 4,000,000 generations and sampled every 100 generations. ML analysis was conducted in MEGA v. 5.05 software [22], performing a heuristic search with 1000 replicates. The MP analysis was performed with PAUP* (Phylogenetic Analysis Using Parsimony) v. 4.0b10 [23]. Trees were inferred by using the heuristic search option with TBR (Tree-bisection-reconnection) branch swapping and 1,000 random sequence additions. Tree length [TL], consistency index [CI], retention index [RI], rescaled consistency index [RC], and homoplasy index [HI] were calculated. The robustness of the equally most parsimonious trees was evaluated by 1,000 bootstrap replications, resulting from a maximum parsimony analysis, each with 10 replicates of random stepwise addition of taxa. The Markov chain Monte Carlo (MCMC) sampling approach was used to calculate posterior probabilities. Chains were analyzed with random starting trees for 4,000,000 generations. The remaining trees after burning-in [24], were used to build a 50 % majority rule consensus tree where the percentage of the remained best trees supporting a branch represents the Bayesian posterior probabilities. The resulting trees were printed with Tree view [25], and Tree Graph 2. Cylindrocladiella peruviana CPC 5614 (AY725736) was used as an out-group.
Results
Distribution and morphological characteristics
Calonectria pseudonaviculata was obtained from all infected plants and sample locations. The disease caused by C. pseudonaviculata was spread throughout the Hyrcanian forest from the western region to eastern areas bordering the Caspian Sea (Figure 1). Based on morphological characteristics, all 75 isolates collected during the study were identified as C. pseudonaviculata. Colony pattern of Iranian isolates on PDA was dark brown at the centre fading through to light brown with a pale luteous halo. Conidia were cylindrical, rounded at the both ends, 1-septate, 48-68 (47.3) × 4-6 (4.8) μm. Stipes were long (90-140μm), hyaline, the extension terminating in a broadly ellipsoid papillate vesicle, the widest part was above the middle.
Radial growth rate
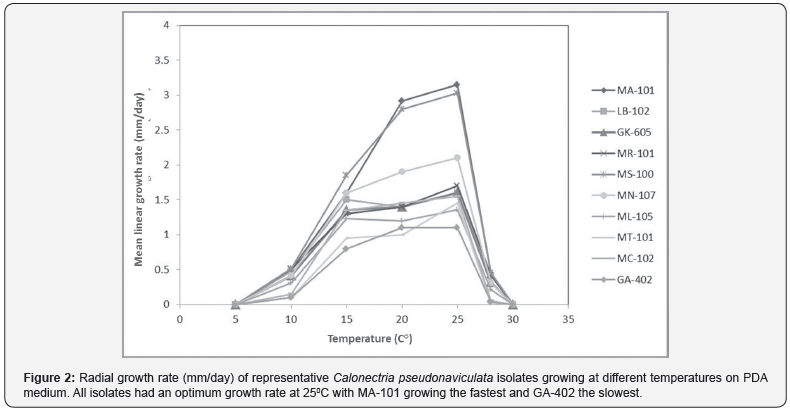
The isolates differed markedly in their growth rate (Figure 2) with growth rate was reduced at 10 °C and 28 °C and ceasing entirely at 5 °C and 30 °C. Optimum temperature for mycelium growth was 25 °C for all isolates with isolates grouped into three groups according to mycelium growth rate at 25 °C for 7 days on PDA: slow growth rate group (0-12mm) included GA402, GB104, MT101; moderate growth rate group (13-23mm) included GA408, GK605, GR102, LB102, MC100, MC102, MCh102, MD100, MG101, MK105, ML105, MR101, MV101; and the fast growth rate group (more than 23 mm) included: MA101, MK104, MM12, MN107 (Figure 3).
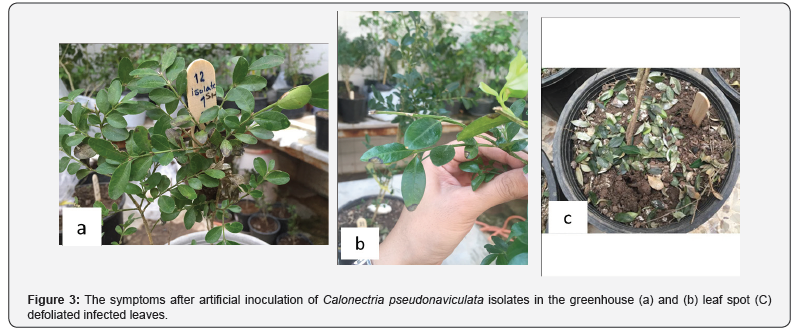

Pathogenicity
Symptoms of boxwood blight disease appeared after 12- 15 days, and after 21 days the disease symptoms included dark brown spots on the upper side of the leaves, elongated spots on the young twigs and leaf shedding that the pathogenicity of different isolates was significantly (P<0.01) different, and placed in three groups according to their DSI: low pathogenicity (0-13%), medium pathogenicity (13.4%-22.1%) and high pathogenicity (24.2%-39.2%).
Among the 35 tested isolates, MN-107, which was isolated from Neka in north-eastern part of the Mazandaran province, caused the greatest disease severity, while isolates MK-104, MK-101 and MA-105 obtained from Savadkoh-Sorkhkolah and Afrachal at the south-eastern part of the Mazandaran province, showed the least disease severity. All isolates from the Mazandaran province were represented across all DSI groups. The isolates collected from Guilan province, located west of Caspian Sea border line, were highly pathogenic. Based on the ANOVA there was no correlation between pathogenicity of isolates and altitude of the origin sample and differences were not statistically significant at a probability of 0.01.
Phylogenetic analysis
Trimmed sequences of β-tubulin, Calmodulin (CaM) and Histone H3 genes of the 20 representative isolates were matched between 99 to 100 percent to the registered sequences of C. pseudonaviculata in NCBI. Phylogenetic analysis was conducted using sequences of the β-tubulin (360bp), Calmodulin (447bp) and Histone H3 (403bp) genes individually (not shown) and combined (Figure 4). The divergence among βtub sequences was too low to allow separation between C. henricotiae and C. pseudonavicula, but those of CaM and H3 sequences were more variable and allowed reliable separation of these two species. Therefore, phylogenetic trees were constructed using combined βtub, CaM and H3 sequences for the selected 20 isolates obtained in this study, and another 14 sequences downloaded from GenBank along with a sequence of Cylindrocladiella peruviana as an out-group taxon.
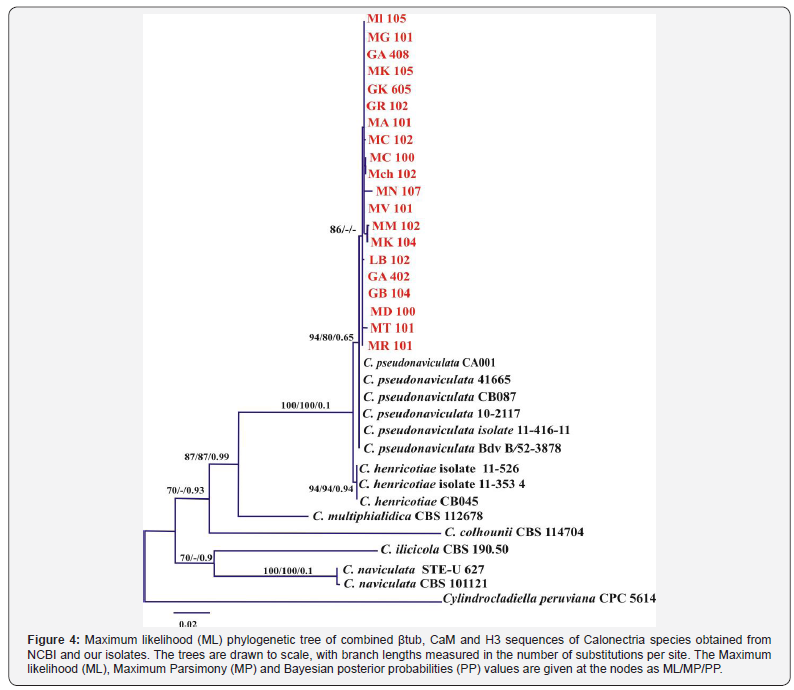
The combined βtub, CaM and H3 dataset yielded 1203 characters (including gaps). Dirichlet base frequencies and the HKY+ I model for βtub, HKY+ G model for CaM and GTR+G model were selected for H3 and set in Mr Bayes. The Maximum Likelihood (ML) analysis of the concatenated alignment resulted in a best tree with a log-likelihood. The parsimony analysis indicated that 782 characters were constant, 202 variable characters parsimonyuninformative and 219 characters parsimony-informative. After a heuristic search using PAUP, three most parsimonious trees were obtained (tree length = 682 steps, CI = 0.834, RI = 0.794, RC = 0.662, HI = 0.166). The Maximum Likelihood analysis resulted in a tree with the same topology and clade as the MP and BA trees (Figure 4).
The combined dataset from βtub, CaM and H3 genes sequence analysis successfully resolved that all isolates from the present study were located in the same clade within C. pseudonaviculata with ML bootstrap = 94%, MP bootstrap = 80% and Posterior Probability = 0.65 and were separated with high bootstrap support (ML and MP bootstrap = 100% and PP = 0.1) from C. henricotiae isolates (Figure 4).
C. pseudonaviculata isolates collected from boxwood trees from the forests of Mazandaran, Guilan and Golestan provinces resided within one group with 99 bootstrap support. No polymorphisms were found between the btub, CaM and H3 sequences of isolates of the same clade.
Discussion
In the present study, 75 isolates of C. pseudonaviculata were obtained from different geographical locations and altitudes from across the Hyrcanian forest of Iran. Twenty representative isolates showed 99-100% similarity with isolates from Europe and America based on multilocus sequencing of the partial btub, CaM and H3 genes. C. henricotiae species was not identified amongst the Iranian isolates. In a previous study, Henricote and Culham (2002) determined genetic diversity of 17 C. pseudonaviculata isolates from England and one isolate from New Zealand by sequencing ribosomal 5.8S RNA, ITS, ß-tubulin (TUB), high mobility group (HMG) of the MAT1-2-1 mating type genes (HMG-MAT1-2-1) along with AFLP. Their results showed that all isolates were genetically homogeneous. Two hundred and thirty-four isolates from 15 countries and four continents have been separated into two genetic clades (G1 and G2) using ITS, tubβ, MAT-1-2-1, Calmodulin and Histone H3 sequences [2]. Clade G1 has been confirmed as C. pseudonaviculata sensu stricto and genetic diversity wasn’t observed among C. pseudonaviculata isolates in this clade. The G2 clade was proposed as a new species, Calonectria henricotiae sp. nov. [2].
Isolates collected in this study were separated in the three categories: slow, moderate and rapid growth according to their diameter growth at different temperatures. The isolates were similar in terms of morphological characters. Mycelium growth of all isolates ceased at 5 °C and 30 °C, which were described as the criterion temperatures for distinguishing C. pseudonaviculata from other species of this genus [6].
There was no correlation between growth rate, genetic diversity and geographic regions of the collected isolates. The results of the pathogenicity test showed that the majority of high growth rate isolates were more virulent, but the fast-growing and slow-growing isolates did not help differentiate the three different pathogenicity groups.
There was no correlation between disease severity of isolates and the altitude of region from which isolates had been collected. All isolates collected from Guilan province where the boxwood blight outbreak was first observed, had a higher disease severity index. Also, the isolates collected from the areas where the prevalence and disease severity of boxwood blight were more, showed more virulence.
Conclusion
In conclusion it seems that the lack of genetic diversity among Iranian isolates is possibly due to the recent introduction of this pathogen, and the interaction between the pathogen and its specific host which is limited to one variety, B. sempervirens var. hyrcana, endemic to the Hyrcanian forests. This host species has grown in the region for thousands of years and during this time has regenerated locally and consequently has a low genetic diversity [26]. Our preliminary findings suggest that the population of C. pseudonaviculata in Iranian boxwood forests is likely to have low genetic diversity and this may be related to its recent introduction and possibly the introduction of a single or few isolates. Further work and in particular population studies are required to determine if this is the case. This will require more intensive collection and analysis of isolates within and outside of Iran.
Acknowledgements
We gratefully acknowledge the technical assistance of Mr. Hajmansoor and Mr. Ahangaran for their support in the laboratory and greenhouse, respectively.
References
- Jalili A, Jamzad Z (1999) Red Data Book of Iran, a preliminary survey of endemic, rare and endangered plant species of Iran. Iranian Research Institute of Forests and Rangelands. pp. 748.
- Gehesquiere B, Crouch JA, Marra RE, Vanpoucke K, RYS F, et al. (2015) Characterization and taxonomic reassessment of the box blight pathogen Calonectria pseudonaviculata, introducing Calonectria henricotiae sp. nov. Plant Pathol 65(1): 37-52.
- Henricot B, Sierra AP, Prior C (2000) A new blight disease on Buxus in the UK caused by the fungus Cylindrocladium. Plant Pathol 49(6): 805.
- Ridley G (1998) New plant fungus found in Auckland box hedges (Buxus). Forest Health News 77: 1–2.
- Crous PW (2002) Taxonomy and pathology of Cylindrocladium (Calonectria) and allied genera. St Paul, MN, USA: The American Phytopathological Society.
- Henricot B, Culhum A (2002) Cylindrocladium buxicola, a new species affecting Buxus spp., and its phylogenetic status. Mycologia 94(6): 980- 997.
- Mirabolfathy M (2013) Outbreak of Boxwood tree leaf drop in Guilan and Mazndaran forests. Proceeding of 1st Iranian Mycological Congress, University of Guilan, Rasht, Iran, pp. 8.
- Mirabolfathy M, Ahangaran Y, Lombard L, Crous PW (2013) Leaf blight of Buxus sempervirens in northern forests of Iran caused by Calonectria pseudonaviculata. Plant Dis 97(8): 1121-1122.
- Rezaee S, Kia-Daliri H, Sharifi K, Ahangaran Y, Hajmansoor S (2013) Boxwood blight caused by Cylindrocladium buxicola in Tonekabon forest. Appl Entomol Phytopathol 80(2): 197-198.
- Khazaeli P, Rezaee S, Mirabolfathy M, Zamanizadeh H, Kiadaliry H (2015) Report of Boxwood blight extension to Golestan province forests. J Appl Entomol Phytopathol 83(1): 85-86.
- Molina JMG, Hamid AH, Sathyapala S, Altrell DO, Farahanirad H, et al. (2014) Iran Zagros and Caspian emergency project mission report. Food and Agriculture Organization of the United Nations.
- Booth C (1971) Methods in microbiology. Vol 4. Academic Press, London, UK, pp. 795.
- Henricot B, Gorton C, Denton G, Denton J (2008) Studies on the control of Cylindrocladium buxicola using fungicides and host resistance. Plant Dis 92: 1273-1279.
- Peever TL, Canihos Y, Olsen L, Ibanez A, Liu Y, et al. (1999) Population genetic structure and host specificity of Alternaria spp. causing brown spot of Minneola tangelo and rough lemon in Florida. Phytopathology 89(10): 851-860.
- Wheeler BEJ (1969) An Introduction to Plant Diseases, John Wiley and Sons Limited, London, UK, pp. 374.
- Nicholson P, Rezanoor HN, Simpson DR, Joyce D (1997) Differentiation and quantification of the cereal eyespot fungi Tapesia yallundae and Tapesia acuformis using a PCR assay. Plant Pathol 46: 842-856.
- Glass NL, Donaldson GC (1995) Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl Environ Microbiol 61(4): 1323-1330.
- Carbone I, Kohn LM (1999) A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 91(3): 553- 556.
- Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25(24): 4876-4882.
- Nylander JA, Ronquist F, Huelsenbeck JP, Nieves-Aldrey JL (2004) Bayesian phylogenetic analysis of combined data. Syst Biol 53(1): 47- 67.
- Ronquist FM, Teslenko M, Van der Mark P, Ayres DL, Darling A, et al. (2012) MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61(3): 539-542.
- Tamura K, Peterson D, Peterson N, Stecher G, Nei M, et al. (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10): 2731-2739.
- Swofford DL (2003) {PAUP*. Phylogenetic analysis using parsimony (* and other methods). Version 4}. Sinauer Associates, Sunderland.
- Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17(8): 754-755.
- Page RDM (1996) TreeView: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12(4): 357-358.
- Ghandehari V, Ahmadikhah A, Payamnoor V (2013) Genetic diversity of Buxus hyrcana populations in north of Iran using ISSR markers Genetic diversity of Buxus hyrcana populations in north of Iran using ISSR markers. Iranian Journal of Rangelands and Forests Plant Breeding and Genetic Research 21(1): 1-11.






























