Assessment of Breeding Potential of Cultivated Okra (Abelmoschus esculentus L. Moench) for Selecting Donor Parent Aiming at Enation Leaf Curl Virus Disease Tolerance in Eastern India
Ayam Pushparani Devi1, Swadesh Banerjee1, Praveen Kumar Maurya1, Tridip Bhattacharjee1, Imtinungsang Jamir2, Soumitra Chatterjee3, Asit Kumar Mandal2, Subrata Dutta2 and Arup Chattopadhyay1*
1 Department of Vegetable Science, Bidhan Chandra Krishi Viswavidyalaya, India
2Department of Plant Pathology, Bidhan Chandra Krishi Viswavidyalaya, India
3 Department of Agricultural Economics, Bidhan Chandra Krishi Viswavidyalaya, India
Submission: November 11, 2018; Published: November 29, 2018
*Corresponding author: Arup Chattopadhyay, Department of Vegetable Science, Bidhan Chandra Krishi Viswavidyalaya, Mohanpur-741252, Nadia, West Bengal, India
How to cite this article: Ayam P D, Swadesh B, Praveen K M, Tridip B, Arup C,etc al. Assessment of Breeding Potential of Cultivated Okra (Abelmoschus esculentus L. Moench) for Selecting Donor Parent Aiming at Enation Leaf Curl Virus Disease Tolerance in Eastern India . Agri Res& Tech: Open Access J. 2018; 18(5): 556072. DOI: 10.19080/ARTOAJ.2018.18.556072
Abstract
Enation leaf curl virus (ELCV) disease is becoming a menace in major okra growing regions of the tropics, causing substantial loss in marketable yield. A strategic breeding program is urgently required to evolve tolerant line(s) against ELCV disease. No developed varieties in India were identified free from the attack of this disease. However, advanced breeding lines of okra have potential to broaden the genetic variability may improve tolerance to this disease. Out of 560 germplasm screened over two consecutive years, 20 genotypes of okra were infected with ELCV disease, and were evaluated for tolerance at periodic interval under field condition and to assess their genetic diversity through multivariate analysis. Infection of the disease started at 35 days after sowing (DAS) and found maximum at 80 DAS. Based on percent disease index (PDI) values, genotypes were categorized as highly tolerant (10/OKYVRES-5 and 10/OKYVRES-10), moderately tolerant (10/OKYVRES-11 and IC112449), moderately susceptible (IC105675, IC411692, IC417885, IC433445, IC433533, IC433532 and BCO-1) and highly susceptible (10/OKYVRES-6 and IC411880). The cumulative progress of disease using area under disease progress curve (AUDPC) varied greatly between susceptible and tolerant genotypes. Based on diversity analysis, twenty genotypes were grouped into five clusters. Four characters PDI of ELCV, node at 1st flowering, fruit length and days to 1st flowering contributed maximum towards divergence. Based on averages, D2 statistics, principal component analysis and ELCV disease severity, five genotypes ‘2014/OKYV RES-5’, ‘2014/OKYV RES-10’, ‘2014/OKYV RES-11’, ‘IC433532’ and ‘IC411692’ appeared to be promising candidates for further utilization in ELCV disease resistant breeding programme.
Keywords: ELCV; Genetic diversity; Okra; Principal component analysis; Percent disease index
Abbrevations: ELCV: Enation Leaf Curl Virus; YVMV: Yellow Vein Mosaic Virus; FYM: Farm Yard Manure; DAS: Days After Sowing; PDI: Percent Disease Index; AUDPC: Area Under the Disease Progress-Curve; PCA: Principal Component Analysis
Introduction
Okra (Abelmoschus esculentus L. Moench.) is a vegetable crop of great social and economic importance in the tropics, subtropics and hot regions of temperate zones [1]. It is believed to be the native of South Africa and the first recorded reference was by the Egyptians in 1216 A.D. [2]. A putative ancestor (A. tuberculatus) being native to Uttar Pradesh in India, suggests the Indian origin whereas, the presence of another putative ancestor (A. ficulneus) in East Africa, suggesting northern Egypt and Ethiopia as its geographical origin [3]. Okra is an often-cross-pollinated crop as the natural out crossing occurs in a range of 4.00 to 19.00% [4], with the maximum of 42.20% [5].
Okra cultivation is concentrated exclusively in the developing countries of Asia and Africa with poor productivity, especially in African countries (2.25t/ha) compared with any other region. India ranks first in the world with a productivity of 11.44t/ha [6]. West Bengal, a state of eastern India, covers a coveted position in okra acreage (75.45 thousand hectares) and production (882.39 thousand tonnes). Okra has great potential as foreign exchange earner and accounts for about 60% of the export of fresh vegetables from India [7].
Cultivated okra is mostly susceptible to a large number of begomoviruses having overlapping host rang. One of the reasons for overlapping host range could be the polyphagous nature of the vector whitefly (Bemisia tabaci) and diverse cropping system prevalent in the country. Yellow vein mosaic virus (YVMV), leaf curl virus (LCV), and enation leaf curl virus (ELCV) together cause serious yield losses in okra [8]. The loss in yield, due to YVMV and/ or ELCV in okra is ranging from 30% to 100% depending on the age of the plant at the time of infection [9]. In India, ELCV was first reported from Bangalore (Karnataka) during the early 1980s; causes yield loss up to 80-90% [9]. The characteristic symptoms of ELCV include leaf-cupping/curling, vein-thickening, decrease in leaf surface area, and twisting of petioles. Moreover, the infected plants become severely stunted with fruits being small, deformed, and unsuitable for marketing [10]. Infection at the early growth stages causes no yield of fruit. This disease is emerging to be as future menace of okra cultivation in the Gangetic plains of West Bengal. The occurrence of ELCV disease is severe in certain locations in certain seasons and accordingly screening of breeding populations is required to be done in the hot-spot areas. Host genetic resistance to viruses is one of the most practical, economical and environment-friendly strategies for reducing yield loss in okra. This will help breeders to identify major genes controlling known physiological basis of resistance to ELCV. Paucity of resistance source to this virus in cultivated varieties has forced breeders to look into the new sources of resistance. Therefore, efforts were made to screen okra germplasm against this disease in the Gangetic plains of West Bengal to identify some of the good sources of tolerance against ELCV disease.
Genetic diversity is the base for survival of plants in nature and for crop improvement. Diversity in plant genetic resources provides opportunity for plant breeders to develop new and improved cultivars with desirable characteristics, which include both farmer-preferred traits (high yield potential, large seed, etc.) and breeder-preferred traits (pest and disease resistance and photosensitivity, etc.). Presence of genetic diversity within and between crop plant species permits the breeders to select superior genotypes either to be directly used as new variety or to be used as parent in hybridization programme. Diversity between two parents is essential to realize heterosis and to obtain transgressive segregates. Hence, identification of suitable donor parent among cultivated species of okra is a prerequisite for the development of tolerant recombinant line(s) to ELCV disease. Considering the lack of research done in the Gangetic plains of eastern India, the present investigation was undertaken to identify ELCV disease tolerant/resistant lines and to analyze the genetic divergence of collected materials based on important quantitative traits including ELCV disease severity to identify diverse parents for hybridization program.
Materials and Methods
Plant materials and field growing
Initially 560 okra germplasm were collected from ICARNBPGR, New Delhi, India and were screened against viral diseases at one of the hot spots of the country under the research field of All India Coordinated Research project on Vegetable Crops, Bidhan Chandra Krishi Viswavidyalaya, Kalyani, Nadia, West Bengal, India following augmented design during Kharif seasons of 2015 and 2016. Only 20 out of 560 genotypes were infected with enation leaf curl virus. Okra plants either expressed YVMV or ELCV symptom in a population at a time. The experimental site was thoroughly ploughed to a depth of 20-30cm to get fine tilth with the help of tractor followed by harrowing. The pre-soaked seeds of 20 genotypes were sown after treatment with thiram @ 3g/kg seed in pre-irrigated plots having unit size of 3.0m × 2.1m following randomized block design with three replications during Kharif season, 2017. Spacing of 50cm × 30cm was maintained to accommodate 42 plants per plot. Well decomposed FYM @10 tonnes /ha was applied at the time of final land preparation. Recommended dose of inorganic fertilizers @ 120:60:50 (N:P:K) kg/ha was applied. Urea, single super phosphate and muriate of potash were used as source of N, P and K, respectively. The entire quantity of phosphorus, potassium and half of nitrogen were applied during final land preparation. Remaining N fertilizer was applied in two equal splits (one quarter each) at 30 DAS and at flowering, respectively. Subsequent irrigation was given as and when necessary. Hand weeding was done to control the weeds as and when necessary. Harvesting of fruit was done manually by bending the pedicel with jerk. Tender pods were harvested on every alternate day. Scheduled package of practices was followed to raise the good crop [11].
Experimental data recording
Observations were recorded from 15 randomly selected plants of each plot in each replication. The characters studied were plant height (cm), days to first flowering, days to 50% flowering, node at first flowering, number of ridges/fruits, fruit length (cm), fruit diameter (cm), fruit weight (g), number of fruits per plant, fruit yield per plant (g) and PDI of ELCV (%) at 80 DAS. Fifteen randomly selected tagged fruits of marketable maturity (7 days after anthesis) were sampled from the selected plants per replication to record the observations on fruit characters.
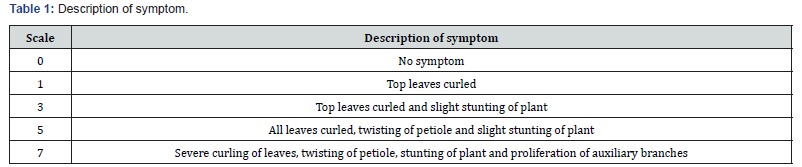
Twenty genotypes were grown without any protective cover of systemic insecticides to take data on percent disease index (PDI). PDI of twenty genotypes was recorded replication wise at five stages at an interval of 15 days starting from 20 days after sowing (DAS) up to 80 DAS. Cupping of leaves and petiole bending of any form in the plant was treated as disease incidence. The PDI was expressed as percentage taken from all the 42 plants in a replication by using disease severity scale (0-7) for single plant through visual evaluation. The following disease severity scale of ELCV developed by Alegbejo [12], was adopted as a reference (Table 1).
The following scale was used for evaluation of resistance/ susceptible reaction of the genotypes to this disease by Percent Disease Index (PDI) as suggested [13].
a. Highly tolerant (HT) PDI ≤ 15%,
b. Moderately tolerant (MT) 16 % < PDI ≤ 25%,
c. Moderately susceptible (MS) 26 %< PDI ≤ 45%,
d. Highly susceptible (HS) PDI > 45%
Number of plants infected in each entry was recorded and PDI was calculated with the following formula:
PDI=Numerical ratings/Highest grade of rating total number of plants of the entry examined X 100
Area under the disease progress-curve (AUDPC), was used to calculate relative spread of the disease among the different genotypes using the following the standard method of Campbell and Madden [14], as follows:
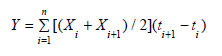
Y is the AUDPC, Xi is the disease incidence of the ith evaluation, and X i +1 is the disease incidence of the i +1st evaluation, t i +1 – ti is the number of days between two evaluations.
Recording of whitefly population during kharif season
The incidence of ELCV disease depends on the population buildup of the vector (Bemisia tabaci) and the presence of virus source. Whitefly populations were monitored from July to September and were recorded on five leaves, two each from lower, middle and one from upper canopy of the plants at 5.30 a.m. to 6 a.m. from 5 randomly selected tagged plants of each genotype at 7 days intervals after sowing.
Statistical analysis
D2 statistic [15], was used for assessing the genetic divergence of twenty genotypes for eleven quantitative traits mentioned above. The grouping of populations was done by using Tocher’s method as described by Rao [16]. Hierarchical cluster analysis was done with those same genotypes in order to observe the degree of association according to their characteristics that was expressed in dendrogram following Ward’s [17], method. Principal component analysis (PCA), as the method of identifying the factor dimension of the data, was used to summarize varietal information in a reduced number of factors (PDI of ELCV, node at 1st flowering, fruit length and days to 1st flowering) for selection of the best performing genotype(s).
Results
Screening of okra genotypes against ELCV disease under field condition
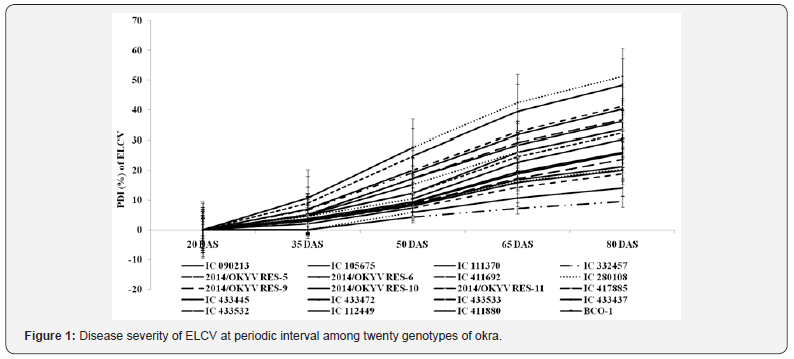
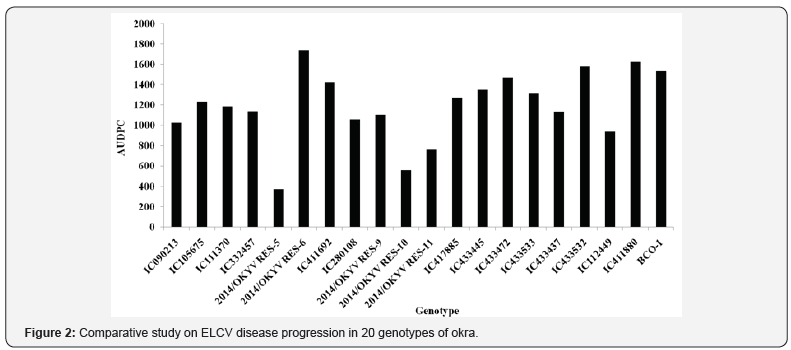
Reactions of different genotypes in terms of PDI values of ELCV differed at different DAS of okra. Genotypes showed no severity of ELCV disease at 20 DAS (Figure 1). As the growth of the plant progresses, the PDI values gradually increased from 35 DAS which varied from 0.00% to 10.71% and then showed variable responses from 50 DAS onwards up to 80 DAS. No further infection was observed beyond 80 DAS. The PDI values varied from 9.52% to 51.19% at 80 DAS (Figure 1). Among the genotypes, PDI values were lower (<15%) in 14/OKYVRES-5 and 14/OKYVRES-10 followed by 14/OKYVRES-11(<20%), and higher (>45%) in 14/ OKYVRES-6 followed by IC411880 at 80 DAS. The genotypes 14/ OKYVRES-5 and 14/OKYVRES-10 were categorized as highly tolerant, and 14/OKYVRES-11 and IC112449 were moderately tolerant. However, the genotypes IC105675, IC411692, IC417885, IC433445, IC433533, IC433532 and BCO-1 were moderately susceptible, and 14/OKYVRES-6 and IC411880 were highly susceptible according to the categorization scheme of Cao et al. [13]. The range of area under disease progress curve (AUDPC) of the different genotypes varied greatly between tolerant and susceptible groups (Figure 2). In the present study, the AUDPC values were found to be minimum (up to 800) within tolerant group and above 1500 within the susceptible group.
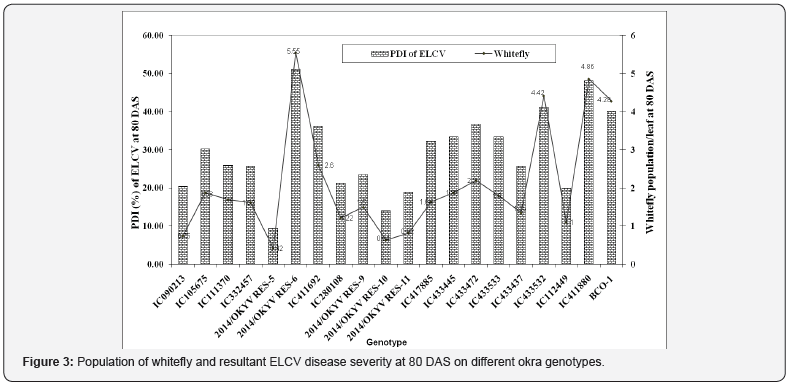
The population dynamics of whitefly/leaf was monitored among okra genotypes tested under study (Figure 3). Population of whitely differed among okra genotypes at 80 DAS. The population of whitefly was found maximum in 14/OKYVRES-6 (5.55/leaf) followed by IC411880 (4.86/leaf), IC433532 (4.42/leaf) and BCO- 1 (4.28/leaf). However, whitefly population remained minimum in 14/OKYVRES-5 (0.42/leaf) followed by 14/OKYVRES-10 (0.64/ leaf) and 14/OKYVRES-11 (0.82/leaf).
Genetic diversity of the genotypes through multivariate analysis
The basic step for crop improvement depends on characterization and identification of cultivars. The present study aimed at analyzing the genetic divergence of 20 genotypes employing 11 important quantitative characters, plant height, node at 1st flowering, days to 1st flowering, days to 50% flowering, ridges/fruit, fruit length, fruit diameter, number of fruits/plants, fruit weight, fruit yield/plant and PDI of ELCV. Twenty genotypes were classified into five clusters on the basis of D2 values (Table 2). Cluster I had the largest with seven genotypes, followed by cluster II (4 genotypes) whereas, Cluster III, IV and V had three genotypes each. Thus, formation of cluster with different genotypes indicated diversity among the genotypes.
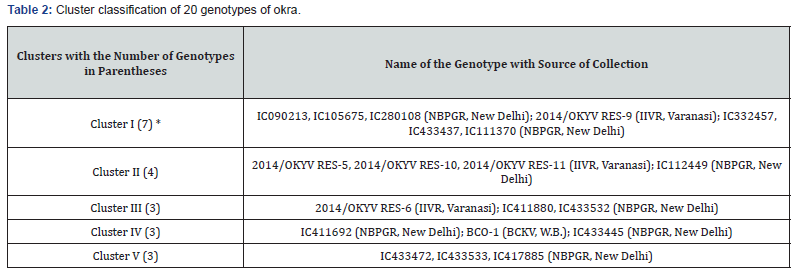
*Figures in parentheses indicate number of genotypes.
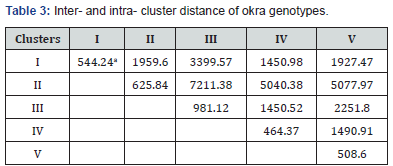
aBold diagonal values indicate intra-cluster distance; the remainder of values indicate the inter cluster distances.
The intra-and inter-cluster distance represent the index of genetic diversity among clusters (Table 3). The range of intracluster distance was from minimum of 464.37 in fourth clusters to maximum of 981.12 in the third cluster. This apparently indicated that cluster III have genotypes that are relatively distant from each other than the other clusters which have lower D2 distances. The maximum inter-cluster distance of 7211.38 was observed among the second and third cluster indicating large genetic differences among genotypes of these two clusters. Minimum inter-cluster distance of 1450.52 was observed between the III and IV clusters indicating significantly lesser genetic differences among the genotypes of these two clusters.
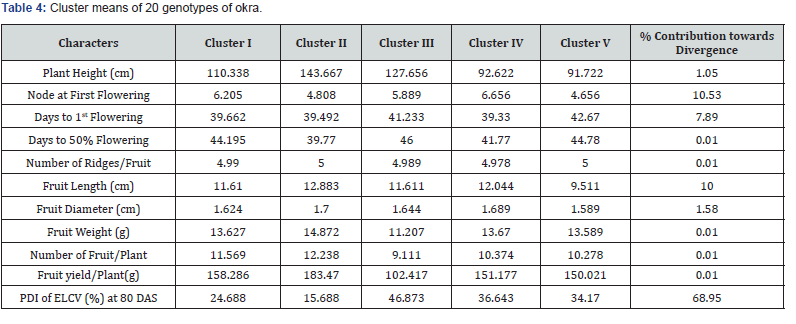
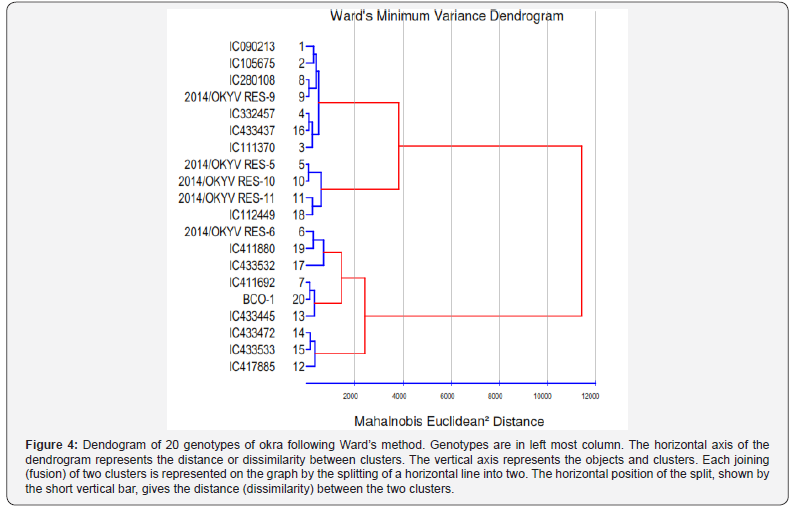
The character means were also worked out for the genotypes falling in these five clusters (Table 4). Cluster II was found to be the cluster consisting of genotypes having taller plant (143.66cm), longer fruit length (12.88cm), wider fruit diameter (1.70cm), more fruit weight (14.87g), maximum number of fruits/plant (12.23) and high fruit yield/plant (183.47g). The genotypes of this cluster took minimum days to produce 50% flowering (39.77) and exhibited low severity of ELCV disease (15.68%). The least mean value for fruit yield/plant (102.41g), number of fruits/plant (9.11) and fruit weight (11.20g) was represented by Cluster III. The highest ELCV disease severity was also recorded in Cluster III (46.87%) followed by Cluster IV (36.64%). Top four characters namely, PDI of ELCV (68.95%), node at 1st flowering (10.53%), fruit length (10.00%) and days to 1st flowering (7.89%) contributed maximum towards divergence. In further study of dendrogram following Ward’s [17], method by using squared Euclidean distance, it became clearly evident that there was high diversity among the okra genotypes along with strong relationships among the genotypes (Figure 4).
Principal component analysis (PCA)
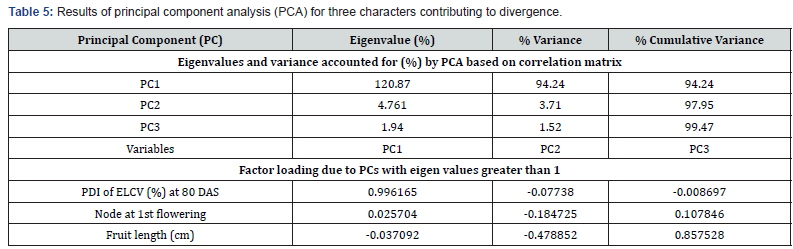
The PCA was performed to obtain a simplified view of the relationship between the characters PDI of ELCV, node at 1st flowering, and fruit length which explained nearly 100% contribution towards divergence, and variable loadings for components PC1 (PDI of ELCV), PC2 (node at 1stflowering) and PC3 (fruit length) were determined (Table 5). These components were selected because their eigenvalues exceeded 1.0 and explained 99.47% of total variance. The first component (PC1) explained 94.24% of total accounted for variance in which an increase of ELCV disease severity leads to formation off lowering at higher node and decrease in fruit length. The second component (PC2) explained an additional 3.71% of the variance in which lesser node of flowering was associated with decrease in fruit length along with less severity of ELCV disease. The third component (PC3) explained another 1.52% of the variance in which higher fruit length was associated with flowering at higher node along with less severity of ELCV disease.
Discussion
Environmental factors, viral strain and the biotype of vector affected the host resistance responses. These highly different and heterogeneous responses to viral pathogen is the result of that specific pathogenic strains for certain hosts may have developed only in certain parts of the region and are not found elsewhere or these hosts may only be susceptible where a number of environmental factors such as temperature, rainfall, inoculums and vector are conductive to disease expression coincide. Presence or absence of virulent strains of the viruses, population of viruliferous vectors, environmental conditions and load of inoculum can influence the ELCV disease tolerance reaction of okra. Combination and cumulative effect of these factors have significant influence on spread of this disease. The advantage of using AUDPC values over the single severity assessment is that the AUDPC reflects disease progress throughout the entire growth period [18]. AUDPC uses multiple evaluations and do not rely on transformations thus lead to more accurate phenotypic evaluation and moreover, simple to calculate [14]. The higher the AUDPC values, more was the susceptibility of the genotype. In this investigation, the cumulative progress of disease using AUDPC, which varied greatly between susceptible and tolerant groups, indicating low cumulative disease progress in tolerant genotypes. Three genotypes 14/OKYVRES-6, IC411880, IC433532 and BCO- 1 were found to be the most attractive genotypes to whitefly infestation. Under the Gangetic plains of West Bengal, these three genotypes also showed high severity (>40.00 % PDI) of enation leaf curl virus (ELCV) which is transmitted by insect vector, whitefly. In general infestation of whitefly has no direct damage to okra crop but two viruliferous whiteflies are sufficient enough to transmit ELCV disease in a single plant [19]. The least infested genotypes 14/OKYVRES-5 and 14/OKYVRES-10 exhibited less ELCV disease severity (<15.00 % PDI) under field condition as the vector population did not cross economic threshold limit. Thus, a positive relationship between whitefly infestation and ELCV disease severity was observed. This finding agreed well with the observation of Venkataravanappa et al. [19].
The composition of clusters II, III and IV revealed that these clusters comprised of heterogeneous geographic origin indicating that the genotypes were distributed amongst the different clusters randomly irrespective of their geographic origin. The grouping of genotypes from heterogeneous geographic origin in one cluster as observed in this cluster analysis could probably be due to the free exchange of breeding material from one place to another or due to unidirectional selection pressure practiced by the breeders in couture the promising genotypes. This study thus brought out the fact that there is no parallelism between genetic diversity and geographical divergence in okra. The present results are in conformity with the findings of Reddy et al. [20], and Seth et al. [21], who also observed the pattern of distribution of okra genotypes from diverse geographical region into different clusters was random.
Maximum inter-cluster distance indicate that the genotypes falling in these clusters had wide diversity and can be used for hybridization programme to get better recombinants in the segregating generations as indicated by Kalloo et al. [22]. Low level of intra-cluster distances was indicative of narrow genetic variation within the cluster. Genotypes of same cluster would not yield desirable recombinants. Genetic divergence of 20 okra genotypes assessed with Mahalanobis D2 statistics revealed that the inter-cluster distances were larger than the intracluster distances indicating a wider genetic diversity between genotypes of cluster with respect to trait under consideration. The analytic observations on the mean yield and its contributing traits performance of different clusters concluded that Cluster II appeared to be the most promising cluster to get the high yielding genotypes with minimum severity of ELCV disease. Cluster mean based estimations are very useful in targeting the genotypes for breeding programme, as they prevent the tedious efforts of screening the inferior germplasm lines. Hence, genotypes from desirable clusters could be directly used for final field evaluation in advanced breeding experiments. Inter crossing among okra genotypes with outstanding mean performance was also suggested for yield improvement of the crop [21,23].
The characters PDI of ELCV, node at 1st flowering, fruit length and days to 1st flowering contributing maximum towards divergence may be used in selecting genetically diverse parents for hybridization programme to exploit either maximum heterosis or to execute efficient selection in the segregating generation. The present study corroborated the findings of Priyanka et al. [24], who also observed fruit length, node at 1st flowering and days to 1st flowering, respectively as the most contributory characters to the total variation observed.
The choice of parents for hybridization depends on genetic diversity of parents. Precise information on the nature and degree of genetic divergence would help the plant breeder in choosing the selective parents for hybridization. The expression of heterosis is influenced by genetic diversity of parents. It is general belief that more diverse the parents within overall limits of fitness, the greater are the chances of obtaining higher amount of heterosis expression in the F1s and a broad spectrum of variability in segregating generations [25]. Cress [26], demonstrated that ‘genetic diversity is necessary for significant heterosis but not sufficient to guarantee it’. Several reports are available to show that hybrids between genetically diverse parents manifest greater heterosis than those between more closely related parents [27,28]. In fact, such a conclusion is based upon a rather restricted range of genetic diversity and may not hold over the entire range of divergence encountered in a species. In general, the level of heterosis increases with the increase in parental diversity up to some limit and decreases with further increase in parental diversity owing to crossability barriers. Thus, maximum heterosis occurs at an optimal or intermediate level of parental diversity. Further, the occurrence of heterosis cannot be predicted on the basis of genetic divergence alone [29]. Apart from the high degree of divergence and the mean yield performance of genotypes should also be given due consideration. The best combination of parents for improvement in various economic characters can be recommended on the basis of per se performance of the genotypes and inter-cluster divergence.
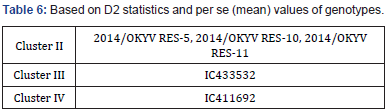
Based on D2 statistics and per se (mean) values of genotypes for yield the following 5 genotypes were selected from three clusters for inter-group crossing of the genotypes which would produce desirable recombinants (Table 6).
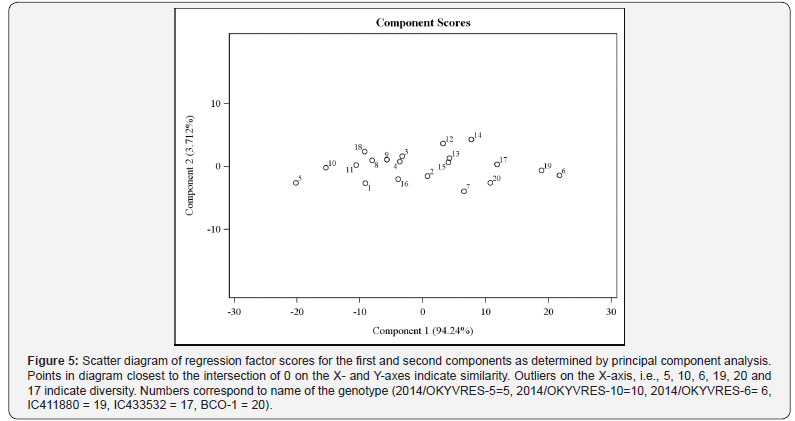
The PCA was also used to determine relationships among okra genotypes of Indian origin and to identify diverse parents for hybridization programme [21,30]. There are no clear guidelines to determine the importance of a trait coefficient for each principal component. Johnson and Wichern [31], regard a coefficient greater than half of the coefficient, divided by the square root of the standard deviation of the eigenvalue of the respective principal component, as significant. Accessions in close proximity were perceived as being similar in PCA; accessions that are further apart were more diverse (Figure 5). The differences observed in the data, and summarized in the PCA, indicated accessions 2014/ OKYVRES-5, 2014/OKYVRES-10, 2014/OKYVRES-11, 2014/ OKYVRES-6, BCO-1, IC433532 and IC411692 were quantitatively dissimilar from others. The remainder of genotypes had similar features forming a separate cluster [32].
Conclusion
Okra production in the Gangetic plain of eastern India is threatened by high severity of ELCV which varied from 9.52 % to 51.19 % at 80 DAS. Genotypes were logically grouped into 5 distinct clusters emphasizing the relative contribution of various quantitative characters to total variability. This study revealed that there is no parallelism between genetic diversity and geographical divergence in okra. Three principal components [Percent disease index (PDI) of ELCV, node at 1st flowering, and fruit length] had eigen values >1 and together accounted for 99.47% of total variation. Five donor parents 2014/OKYVRES-5, 2014/OKYVRES-10, 2014/OKYVRES-11, IC433532 and IC411692 were selected based on averages, multivariate analysis and ELCV disease severity and they should, therefore, be introduced into okra breeding programs to enhance tolerance against this disease. Thus, introduction of tolerant lines as such in disease prone areas and/or their subsequent utilization in ELCV disease resistant breeding program would be effective to sustain the productivity level of the crop in eastern India.
Acknowledgment
Authors wish to acknowledge ICAR-NBPGR, New Delhi, India and ICAR-IIVR, Varanasi, India for providing genetic materials and partial financial support to conduct this study.
References
- Duzyaman E, Vural H (2003) Evaluation of pod characteristics and nutritive value of okra genetic resources. Acta Horticulturae 598(1): 103-110.
- Lamont W (1999) Okra a versatile vegetable crop. Horticultural Technology 9(2): 179-184.
- Charrier A (1984) Genetic resources of the Genus Abelmoschus Med. (Okra). International Board for Plant Genetic Resources; IBPGR Secretariat, Rome.
- Kumar S, Dagnoko S, Haougui A, Ratnadass A, Pasternak D, et al. (2010) Okra (Abelmoschus spp.) in West and Central Africa: potential and progress on its improvement. African Journal of Agricultural Research 5(25): 3590-3598.
- Kumar N (2006) Breeding of Horticultural Crops. New India Publication Agency. New Delhi, pp. 173-177.
- Anonymous (2017) Indian Horticulture Database. National Horticulture Board. Government of India, Gurgaon, India.
- Singh B, Singh PM, Sanwal SK, Pal AK (2014) Standardization of costeffective hybridization technique for hybrid seed production in okra (Abelmoschus esculentus). Indian Journal of Agricultural Sciences 84(9): 1111-1114.
- Venkataravanappa V, Reddy CNL, Jalali S, Reddy MK (2013) Molecular characterization of a new species of begomovirus associated with yellow vein mosaic of bhendi (okra) in Bhubhaneswar. Europran Journal of Plant Patholology 136(4): 811-822.
- Singh SJ (1996) Assessment of losses in okra due to enation leaf curl virus. Indian Journal of Virology 12(1): 51-53.
- Sanwal SK, Singh M, Singh B, Naik PS (2014) Resistance to yellow vein mosaic virus and okra enation leaf curl virus: challenges and future strategies. Current Science 106: 470-471.
- Chattopadhyay A, Dutta S, Bhattacharya I, Karmakar K, Hazra P (2007) Technology for Vegetable Crop Production. All India Coordinated Research Project on Vegetable Crops. Directorate of Research, West Bengal, India.
- Alegbejo MD (1997) Evaluation of okra genotype for resistance to okra mosaic virus. Proceeding of the 15th Annual Conference of Horticulture Society of Nigeria, held at the National Horticultural Research Institute. Ibadan: Nigeria, P. 60.
- Cao Bi-Hao, Jian-Jun Lei, Yong Wang, Chen Guo-Ju (2009) Inheritance and identification of SCAR marker linked to bacterial wilt resistance in egg plant. African Journal of Biotechnology 8(20): 5201-5207.
- Campbell CL, Madden LV (1990) Introduction to Plant Disease Epidemiology. Wiley-Interscience, New York, USA; pp. 532.
- Mahalanobis P (1936) On the generalized distance in statistics. Proceedings of the National Institute of Science, India. 12: 49-55.
- Rao CR (1952) Advance Statistical Methods in Biometrics. John Willey and Sons Inc, New York, USA; pp. 402.
- Ward JH (1963) Hierarchial grouping to optimize an objective function. Journal of the American Statistical Association 58(301): 236-244
- Roy A, Acharyya S, Das S, Ghosh R, Paul S, et al. (2009) Distribution, epidemiology and molecular variability of the begomovirus complexes associated with yellow vein mosaic disease of mesta in India. Virus Res 141(2): 237-246.
- Venkataravanappa V, Reddy CNL, Jalali S, Briddon RW, Reddy MK (2015) Molecular identification and biological characterization of a begomovirus associated with okra enation leaf curl disease in India. Europian Journal of Plant Pathology 141(2): 217-235.
- Reddy TM, Hari Babu K, Ganesh M, Chandrasekhar Reddy K, Begum H, et al. (2012) Genetic variability analysis for the selection of elite genotypes based on pod yield and quality from the germplasm of okra (Abelmoschus esculentus L. Moench). Journal of Agricultural Technology 8(2): 639-655.
- Seth T, Chattopadhyay A, Chatterjee S, Dutta S, Singh B (2016) Selecting parental lines among cultivated and wild species of okra for hybridization aiming at YVMV disease resistance. Journal of Agricultural Science and Technology 18: 751-762.
- Kalloo G, Singh VP, Dudi BS, Pratap PS (1980) Analysis of variation and genetic diversity in garden peas. Haryana Agricultural University Journal of Research 10: 540-546.
- Mallesh S, Shivshankar S, Muthaiah K, Prabhakar B, Vittal LM (2016) Genetic divergence study in okra [Abelmoschus esculentus (L.) Moench.]. Journal of Environmental Ecology 34(4b): 2163-2167.
- Priyanka VD, Thirupathi Reddy M, Begum H, Sunil N, Jayaprada M (2017) Genetic divergence analysis of inbred lines of okra [Abelmoschus esculentus (L.) Moench]. Int J Curr Microbiol App Sci 6(11): 379-388.
- Arunachalam V (1981) Genetic distance in plant breeding. Indian Journal of Genetics and Plant Breeding 41(2): 226-236.
- Cress CE (1966) Heterosis of the hybrid related to gene frequency difference between two populations. Genetics 53(2): 269-274.
- Moll, RH, Stuber CW (1974) Quantitative genetics empirical results relevant to plant breeding. Advances in Agronomy 26: 277-313.
- Singh SP, Sharma JR (1989) Genetic improvement of pyrethrum IV. Selective divergence and potential hybrid clones. Theor Appl Genet 78(6): 841-846.
- Matzinger DR, Werusman FA (1958) Four cycles of mass selection in a synthetic variety of an autogamous species, Nicotiana tabaccum L. Crop Science 8(2): 239-243.
- Reddy MT, Kadiyala H, Mutyala G, Hameedunnisa B, Reddy RSK, et al. (2013) Exploitation of hybrid vigour for yield and its components in okra [Abelmoschus esculentus (L.) Moench]. American Journal of Agricultural Science and Technology 1: 1-17.
- Johnson RA, Wichern DW (1988) Applied Multivariate Statistical Analysis. (2nd edn), John Wiley and Sons Inc, New York, USA; pp. 354.
- Sharma JP, Singh AK, Kumar S, Sharma N (2008) Genetic divergence studies in okra [Abelmoschus esculentus (L.) Moench]. Journal of Research, SKUAS & T 7(1): 1-8.






























