Mitochondrial DNA Indicate that Kenyan Indigenous Goat (Capra hircus) Population is Diverse
Jacqueline N Kivila1,2, Elijah K Githui2*, Nelson O Amugune1, John M Githaiga1 and Zedekiah A Okwany2
1University of Nairobi, Department of Genetics, Kenya
2Institute of Primate Research, Molecular Genetics Lab, Kenya
Submission: September 11, 2018; Published: September 26, 2018
*Corresponding author: Elijah Kem Githui, Institute of Primate Research, Molecular Genetics Lab, P. O. Box, 24481-00502, Karen, Nairobi 00502, Kenya; Tel: 723539103; Email: kegithui@yahoo.com
How to cite this article: Jacqueline N K, Elijah K G, Nelson O A, John M G, Zedekiah A O. Mitochondrial DNA Indicate that Kenyan Indigenous Goat (Capra hircus) Population is Diverse. Agri Res & Tech: Open Access J. 2018; 18(1): 556044. DOI: 10.19080/ARTOAJ.2018.18.556044
Abstract
Kenyan indigenous goat populations are not fully characterized, and this can result in erosion of their unique genotypes through indiscriminate cross breeding. Kenyan goat breeds are morphologically classified as Small East African goats (SEA), Galla goats and their crosses. This study aimed at genetic characterization of Kenyan regional goat breeds using mitochondrial DNA. 28 goat blood samples of the SEA were obtained from Narok and Kajiado counties and DNA extracted. PCR amplification was carried out using primers flanking mitochondrial D-loop where 24 out of the 28 samples amplified successfully (Narok 13, Kajiado 11) and the 800 base-pair product was sequenced. Phylogenetic analysis of 44 sequences of Kenyan indigenous goats in this study (SEA) combined with data previously submitted to Genbank for Galla goats in Garissa and Isiolo counties of Kenya, showed that the majority of Kenyan indigenous goats are not distinct but rather are of diverse structure with isolated pockets including an early branching lineage. The analysis of genetic diversity gave a weak negative Tajima’s D value (-0.371332) indicating that the population genetic drift is at equilibrium. Haplogroup analysis showed that these goat populations belong to group A and G. Median-joining haplotype network analysis showed no variation among different regions. The information obtained from this study can serve as a guideline in breeding programs combining improved productivity with conservation of unique traits.
Keywords:Indigenous goats; Kenya; Genetics; Traits
Abbrevations: SEA: Small East African; DNA: Deoxyribonucleic Acid; EDTA: Ethylene Di-amine Tetra-Acetic Acid
Introduction
Indigenous goat (Capra hircus) populations contribute greatly to food security, particularly among pastoralist communities in arid and semi-arid regions of sub-Saharan Africa. These locally adapted livestock species are primarily reared for their provision of meat, milk and skin that provide secure investment for the resources-poor farmers [1]. In Kenya as well as in other parts of Africa, indigenous goats are also used for religious and other cultural ceremonies such as payment of dowry [2]. Kenyan indigenous goat inherent characteristics of tolerant to dehydration and preference for browse over a wide range of feeding habits [3], help them adapt to the vast arid and semi-arid regions of the country.
The current genetic diversity of indigenous goat populations bears the molecular signature of historical lineages and demographic expansions. Studies of archaeology show that Capra hircus was domesticated from the wild goat, bezoar (Capra aegagrus) in the Fertile Crescent [4], and this have been confirmed by phylogenetic studies using mitochondrial DNA [5] and nuclear DNA [6]. Currently, there are more than three hundred (300) different breeds of Capra hircus distributed throughout the world [7]. Livestock policy initiative (IGAD LPI) estimates 27.7 million goats in Kenya [8]. The main goat breeds kept for meat in Kenya include the Galla goat, Small East African (SEA) goat and the crosses of these two. The Galla goat, also known as Boran/Somali goat are indigenous in northern Kenya arid lands while Small East African goats is spread out in semi-arid lands in Northern and Southern Kenya. These goats are potentially useful in stocks upgrading breeding program due to their characteristic’s resistance to dehydration, a preference for browse and a wide range of feeding habits [3].
Genetic characterization and tracing of gene flow among these Kenyan indigenous goat populations has been scanty. Migration is sometimes used synonymously with gene flow especially when migration is the driving force in genetic drift [9], can reduce, or conversely, increase an individual’s offspring’s fitness [10,11]. Genetic mutations data has been used to successfully trace the ancestry and extent of gene flow of domestic animals [12]. One of the most commonly used genetic marker is the mitochondrial DNA (mtDNA) because it evolves faster than nuclear DNA. However, mitochondrial DNA has no molecular recombination as is maternally inherited. Molecular phylogeny of genus Capra indicate that geographically neighboring species tend also to be close relatives [5], and the different breeds arose by ancient hybridization between different ancestral wild goats [6,13]. This study aims at genetic characterization and evaluation of gene flow among Kenyan indigenous goat regional breeds using mitochondrial control (D-loop) region.
Materials and Methods
Sample population
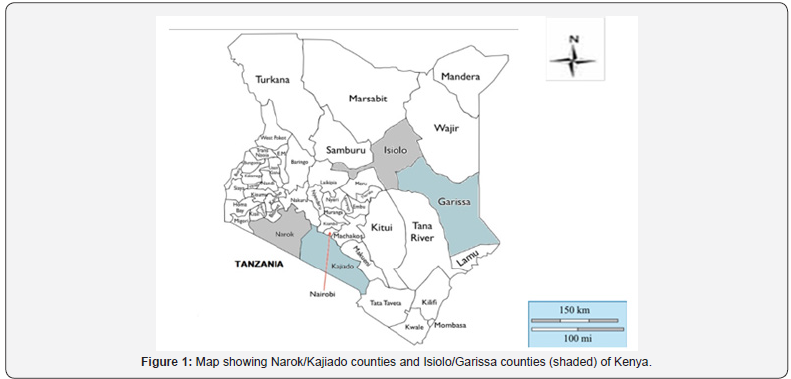
The sampling regions were selected on the basis of regional pastoral communities’ demographic distribution in the country. Small East African (SEA) goats reared by the Maasai pastoralists were sampled from Narok and Kajiado counties of Kenya while Galla goats (also known as Somali or Boran goats) reared by the Somali and Boran pastoralists were sampled from Garissa and Isiolo counties of Kenya (Figure 1).
Ethical clearance
This study is a continuation of a research project that was passed by University of Nairobi Veterinary School, Kabete and by National Commission of Science and Technology, Kenya (Nacosti). Ethical guidelines in the humane treatment of animals were followed, in conjunction with University of Nairobi Veterinary School animal care guidelines.
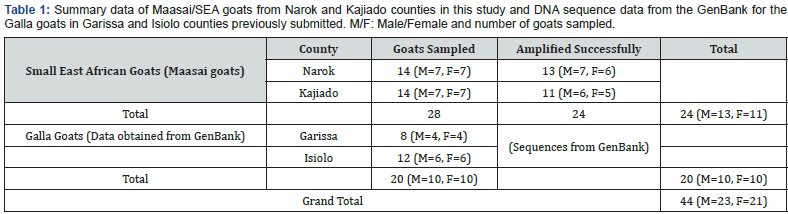
Blood sample collection and DNA analysis: A total of 28 blood samples representative of the Small East African (Maasai) goat breeds were obtained by a veterinary officer in Narok and Kajiado. Half of these goats sampled were male while the others were females (Table 1). The goats were restrained manually and blood obtained by jugular venipuncture into 5ml capacity vacutainer tubes containing 100μl of anticoagulant Ethylene diamine tetra-acetic acid (EDTA). The blood samples were then stored in a cool box, transported to the laboratory and stored at -20 ˚C.
DNA extraction: DNA was extracted from whole blood. 200μl sample of whole blood was incubated in 30μl of proteinase K, 200μl of digestion buffer and 70μl of 10% SDS (Sodium Dodecyl Sulphate) at 56 ˚C for 10 minutes. The DNA was extracted from the lysate by the standard Phenol: Chloroform method [14] and precipitated from the aqueous phase by adding 2 volumes of absolute ethanol. The pellet was then washed twice in 100% ethanol, air-dried for about 20 minutes and dissolved in 100μl of nuclease free water. The quality of DNA was analyzed by electrophoresis in 1% Agarose gel in 1X TAE buffer (40mM Trisacetate, 1mM EDTA, pH 8.0). The respective tubes with DNA were labeled appropriately and stored at –20 °C.
PCR amplification and sequencing: The DNA samples were amplified in 25μl reactions containing 1 unit of DNA Taq polymerase, 2mM of MgCl2, 0.4mM of each dNTP and 1μl of each primer, at 10pMol. Primers flanking the mitochondrial control region (D-loop) were applied: Ibex forward primer I - 5’ ACATGAATTGGAGGACAGCCAGTC-OH 3’ and Ibex reverse primer II - 5’ CTGTAATGCCCATGCCTACC-OH 3’. After 5 minutes of initial denaturation at 92 °C, 35 cycles were run with the following parameters: denaturation at 92 °C for 1min, annealing at 50 °C for 45 seconds, extension at 70 °C for 1minute and a final extension at 70 °C for 3 minutes. 24 out of the 28 DNA samples amplified successfully (Table 1). The amplification product was verified on 1% Agarose gels stained with Ethidium bromide observed under UV light. The fragment was excised from the gel, solubilized in sodium iodide solution then bound to (silica) column in the gene clean procedure. Bound DNA was eluted in 30μl nuclease-free ddH2O.
Gene cleaned DNA was sequenced at Macrogen Inc., Denmark using Sanger di-deoxy sequencing method. Each analyzed sample was independently sequenced three times and the raw sequences selected for non-ambiguous consensus. The consensus sequences were deposited in the NCBI nucleotide database (GenBank Accession ID: MF101245 – MF101258). Previous data for Galla goats found in Garissa and Isiolo counties were downloaded from the Genbank [15].
Phylogenetic analysis
DNA sequences were aligned by Clustal-W program in Bioedit (version 7.05) and saved as a fasta file in MEGA 7 program [16]. The phylogenetic tree was constructed from the aligned nucleotide sequences and the relationships inferred by the neighbour-joining method [17], at bootstrap 1000 replicates [18]. Haplotype diversity, nucleotide diversity, gene flow and genetic differentiation for the goat populations were analysed using the DnaSP program version 5.01 [19]. Gene flow estimates were computed using the sequence data following method by Hudson et al. [20]. Haplotype network analysis was also constructed using the Median-joining method in PopART program [21]. Molecular clock analysis of present day phylogeographic goat breed clusters in relationship to the ancestral wild goats was conducted using Rel Time method [22], as implemented in MEGA 6 [23].
Results
PCR amplification
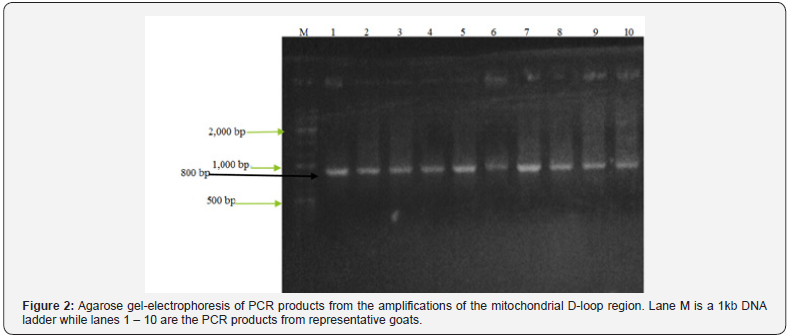
PCR amplification of mitochondrial D-loop invariably gave a product size of approximately 800bp (Figure 2) in all samples representative of indigenous Maasai goats from Narok and Kajiado counties of Kenya. The products were gene cleaned and submitted for custom sequencing.
Phylogenetic relationships of kenyan indigenous goat populations
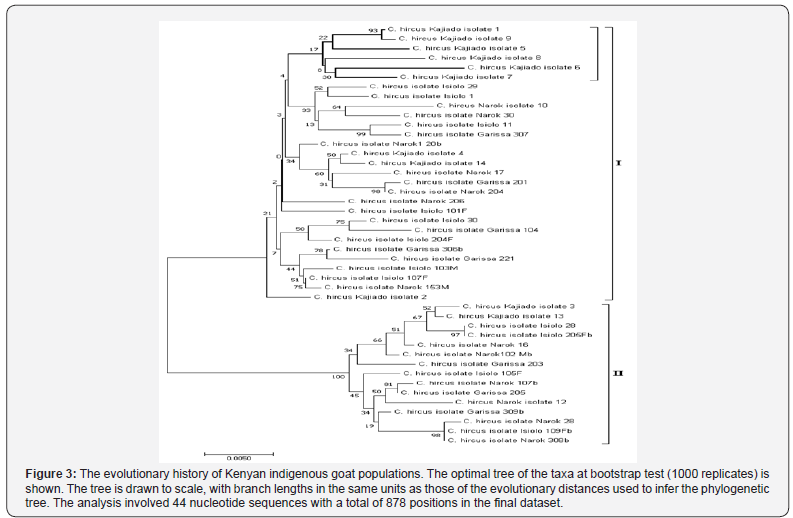
The evolutionary history of Kenyan indigenous goat populations was inferred using the Neighbor-Joining method implemented in MEGA 7. Phylogenetic tree (Figure 3) of the Kenyan goat populations’ sequence data shows two major distinct clusters (I and II) which did not correlate with morphological trait description. Cluster I shows a distinct sub-cluster (bold lines) of Kajiado goats while most of the other sub-clusters are a mixture of goats reared by the Maasai and the Somali/Boran pastoralists.
Genetic diversity analysis: Genetic diversity analysis of Kenyan indigenous goat population mitochondrial D-loop nucleotide data was analyzed in Tajima’s D test. The data showed a slightly weak negative value (-0.371332) reflecting slight departure from neutrality (Table 2).
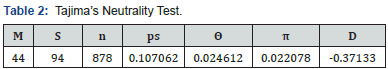
The analysis involved 44 nucleotide sequences of Kenyan indigenous goat populations. There were a total of 878 positions in the final dataset. Evolutionary analyses were conducted in MEGA7. m = number of sequences, n = total number of sites, S = Number of segregating sites, ps = S/n, Θ = ps/a1, π = nucleotide diversity, and D is the Tajima test statistic.
Gene flow and DNA divergence of maasai and somali goat populations: Gene flow and differentiation analysis was done using the DnaSP program version 5.10. The analysis of the 44 nucleotide sequences (878 positions in the dataset) showed 94 polymorphic sites and 42 haplotypes with estimated haplotype diversity of 0.998. The values obtained in the analysis are summarized (Table 3). The data show that there were no fixed differences between the Maasai and Somali goat population but had 53 shared mutations.
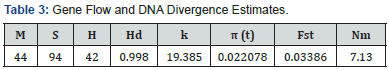
M = no. of sequences; S = no. of polymorphic sites; H = no. of Haplotypes; Hd = haplotype diversity; k = average no. of nucleotide differences; π(t) = nucleotide diversity; Fst and Nm represent the F statistic and the gene flow estimator. Analyses were conducted in dna SP software V5.
Haplogroup analysis and haplotype network of kenyan indigenous goat populations
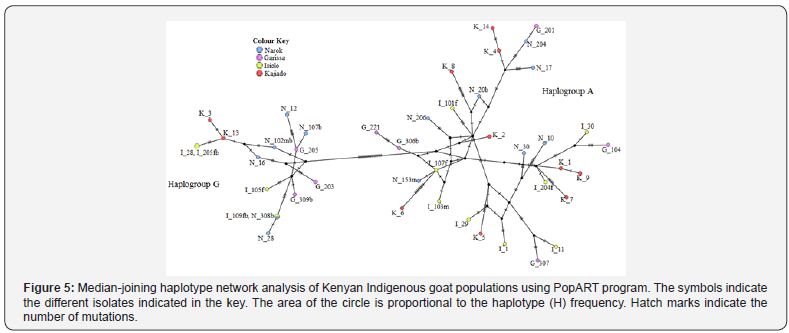
Haplogroup analysis of evolutionary genetic relationships of Kenyan indigenous Maasai and Galla goat isolates (Figure 4) was inferred using the Neighbor-Joining method using mtDNA control region in relation to reference sequences belonging to the six known phylogeographic haplogroups as described by Naderi et al. [24]. Analysis involving 66 nucleotide sequences combining work in this study and datasets from public database resources [25], show that Kenyan Maasai goat major cluster is haplogroups A and minor cluster is haplogroup G. To visualize genetic relationships forty-two haplotypes of Kenyan indigenous goat were analyzed using median-joining haplotype network (Figure 5) implemented in poPART.
The results of this analysis are closely related to the haplotype analysis tree (Figure 4) where haplogroup A cluster together and separate from the haplogroup G. The most common haplotype is represented by isolate I_109fb and N_308b (Figure 5) and is shared by two geographically separated individuals; Isiolo (Galla goat) and from Narok (Maasai goat). The haplotype network also shows that the haplotypes in haplogroup A are more diverse than the haplotypes in haplogroup G as illustrated by the number of hatch marks (mutations) between haplotypes.
Phylogeographic divergence of goat breed lineages
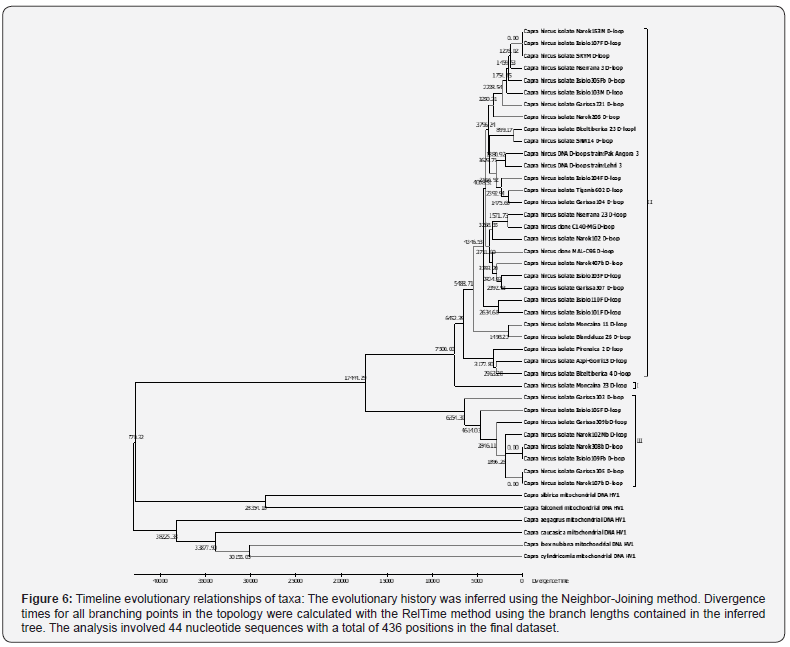
Molecular clock analysis of present day phylogeographic goat breed clusters in relationship to the ancestral wild goats show that the old stock goat breeds arose approximately 6000 years ago and that the majority of present day isolates are less than 3000 years old (Figure 6, clusters II, III). Based on the data analyzed, these isolates seem to arise from old divergent lines of mountain goat, Moncaina isolate and a cluster of divergent lineage observed in Kenyan isolates (Lineage I and cluster II) that separated approximately 17000 years ago. In this time scale, the wild goat lineages and the stock that gave rise to domestic goat separated approximately 47,000 years ago.
Discussion
The transportation and exchange of domestic animals has been closely connected to trade and human relocations since the beginning of animal domestication. This explains why there is a weak phylogeographic structure for the currently more than 300 different breeds of Capra hircus distributed throughout the world [7]. The results (Figure 3 & Figure 4) of this study indicate that Kenyan indigenous goat populations are intermixed with no distinct phenotypes as previously defined by morphology [26]. The mixed genotypes of Kenyan indigenous goat populations likely came through trade, human relocations or by dowry gifts in certain cultural practices. Analysis of indigenous goat populations in Kenya, however observed a sub-cluster within Kajiado goats. This could indicate a geographic barrier between Narok and Kajiado regions (Figure 1) which might restrict trade or movement of these animals. In fact, vast sections of forested/ rugged landscape exist along the Eastern Rift Valley escarpments that divide these two counties and could contribute to limitations in gene flow between the two populations. This is evident in these results where some distinct isolated pockets of populations are observed (Figure 3).

Genetic diversity among the Kenyan goat populations was assessed in the Tajima’s D neutrality test [27,28]. The analysis of the dataset showed a near neutrality, though weak negative value, -0.371332 (Table 2). A neutral Tajima’s D is expected in a population that does not have any demographic changes such as population expansion/contraction and migration. Values close to zero indicate that the nucleotide diversity is near neutrality and the population from which the samples were obtained is almost in equilibrium with respect to drift. Positive selection is expected to give a significantly negative Tajima’s D in a population that does not have any demographic changes going on. The Tajima’s D estimate observed in the current study was comparable with other recent studies [15,29], of Kenyan indigenous goat populations which also reported non-significant Tajima’s D values. This might also be an indication of population expansion after a recent bottleneck as previously reported [15]. For goats, a neutral estimate is consistent with a demographic population expansion [30], expected following domestication from a few founder individuals.
Nucleotide diversity between the Maasai and Somali goat populations showed variation in polymorphic sites with 53 shared mutations but no fixed difference (Table 3). There were 94 nucleotide polymorphisms and 42 haplotypes from 878 sites. The number of polymorphisms and haplotypes was high (haplotype diversity = 0.998) indicating that Kenyan indigenous goat populations are extremely diverse. Exchange of genetic material between populations is the major determinant of population structure [31,32]. The commonly used parameters for gene flow between populations are Fst and Nm. The F statistic (Fst) is a standardized measure of the genetic variance among populations and is used to solve for Nm, which is the number of migrants entering a population successfully per generation. When Nm>0.5, it means that genetic structure is determined mostly by gene flow due to migrations and when Nm<0.5, significant genetic differentiation can result due to genetic drift. Migration of more than one individual between populations every other generation (Nm>0.5), would mean that different alleles at a locus in the two populations would not become fixed by genetic drift. The results of this study suggest that gene flow among the local populations of Capra hircus is high (Nm=7.13) indicating that the representative Kenyan goats have a random mating between populations.
The analysis of haplogroup diversity based on mitochondrial D-loop sequences of Kenyan indigenous goat breeds showed that the goats cluster into two different haplogroups, A and G (Figure 4 & Figure 5) consistent with majority of global geographic domestic goat breeds [24,30]. The frequency of distribution of these haplogroups in this study in which haplogroup A is the major component also concur with the reported patterns in other studies [24,33]. Haplogroup A is highly dominant all over the world. The B haplogroup is found mostly in whole Asia with a few individuals from sub-Saharan Africa and from Greece while haplotypes C and D are found across Asia and Europe. The F haplogroup is from Sicily and the G group is found in North Africa and in Middle East (Figure 4). The phylogeographic timeline lineages (Figure 6) is in agreement with this analysis where a limited number of ancestral lineages separated approximately 17000 years ago [4,30]. These further separated into different clusters 6000 years ago, most likely as a result of modern goat domestication and animal husbandry that started approximately 11000 years ago. The different isolates and their phylogeographic distribution observed today were evident 2000 to 30000 years ago.
Conclusion
Abundant genetic diversity observed in indigenous goat populations in Kenya represents a robust resource for selective breeding adapted to arid and semi arid regions of the country. The adaptive traits in these populations need to be preserved in the breed improvement programs as a strategy of conservation of genetic resources.
Conflict of Interest
All the authors “declare no conflict of interest.
References
- Huson HJ, Sonstegard TS, Silverstein J, Woodward-Greene MJ, Masiga C, et al. (2014) Genetic and Phenotypic Characterization of African goat populations to Prioritize Conservation and Production Efforts for Small-holder Farmers in Sub-Saharan Africa. Proceedings, 10th World Congress of Genetics Applied to Livestock Production.
- Henyambuga SW, Hanotte O, Hirbo J, Watts PC, Kemp SJ, et al. (2002) Genetic characterization of indigenous goat populations of sub-Saharan Africa using microsatelite DNA markers. Department of Animal Science and Production, Sokoine University of Agriculture, Sokoine, Tanzania, pp. 445-452.
- Devendra C, McLeroy GB (1982) Goat and sheep production in the tropics. Intermediate tropical agriculture series. Longman Group, UK, pp. 271.
- Zeder M, Vigne JD, Helmer D (2005) A view from the Zagros: new perspectives on livestock domestication in the Fertile Crescent. In: Vigne JD, Peters J, et al. (Eds.), The first steps of animal domestication. Oxbow Books, England, pp. 125-146
- Zeder M, Vigne JD, Helmer D (2005) A view from the Zagros: new perspectives on livestock domestication in the Fertile Crescent. In: Vigne JD, Peters J, et al. (Eds.), The first steps of animal domestication. Oxbow Books, England, pp. 125-146
- Pidancier N, Jordan S, Luikart G, Taberlet P (2006) Evolutionary history of the genus Capra (Mammalia, Artiodactyla): discordance between mitochondrial DNA and Y-chromosome phylogenies. Mol Phylogenet Evol 40(3): 739–749.
- Porter V (1996) Goats of the world. Farming Press, Ipswich, UK.
- IGAD (2011) Livestock Policy Initiative Working Paper No. 03-11.
- Slatkin M (1985) Gene flow in natural populations. Annu Rev Ecol Syst 16: 393-430.
- Vila C, Sundqvist AK, Flagstad O, Seddon J, Bjornerfeldt S, et al. (2003) Rescue of a severely bottlenecked wolf (Canis lupus) population by a single immigrant. Proceedings of the Royal Society B: Biological Sciences 270(1510): 91-97.
- Adams JR, Vucetich LM, Hedrick PW, Peterson RO, Vucetich JA (2011) Genomic sweep and potential genetic rescue during limiting environmental conditions in an isolated wolf population. Proc Biol Sci 278(1723): 3336-3344.
- Bruford MW, Bradley DG, Luikart G (2003) DNA markers reveal the complexity of livestock domestication. Nat Rev Genet 4(11): 900–910.
- Luikart G, Fernandez M, Mashkour M, England P, Taberlet P (2006) Origins and diffusion of domestic goats inferred from DNA markers. In: Zeder MA, Bradley DG (Eds.), Documenting Domestication: New Genetic and Archaeological Paradigms. University of California Press, Berkeley and Los Angeles, California, USA, pp. 181–208.
- Sambrook J, Maniatis T, Fritsch EF (1987) Molecular cloning. A laboratory manual. Cold spring harbor, 14th Printing. New York, USA. p. 34.
- Githui EK, Kibegwa FM, Kamau JM, Mutura SK, Okwany ZA, et al. (2016) Genetic relationships of indigenous goats reared by pastoralists in Kenya based on mitochondrial D-loop sequence. Anim Gen Res 59: 73- 80.
- Kumar S, Stecher G, Tamura K (2016) MEGA 7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7): 1870-1874.
- Saitou N, Nei M (1987) The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol Biol & Evol 4(4): 406-425.
- Felsenstein J (1985) Confidence limits on phylogenies: An approach using the bootstrap. Evolution 39(4): 783-791.
- Librado P, Rozas J (2009) DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25(11): 1451– 1452.
- Hudson RR, Boos DD, Kaplan NL (1992) A statistical test for detecting geographic subdivision. Mol Biol Evol 9(1): 138–151.
- Bandelt HJ, Forster P, Rohl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Bio Evol 16(1): 37-49.
- Tamura K, Battistuzzi FU, Billing-Ross P, Murillo O, Filipski A, et al. (2012) Estimating Divergence Times in Large Molecular Phylogenies Proceedings of the National Academy of Sciences 109(47): 19333- 19338.
- Tamura K, Stecher G, Peterson D, Alan Filipski A, Kumar S (2013) MEGA 6: Molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30(12): 2725–2729.
- Naderi S, Rezaei HR, Taberlet P, Zundel S, Rafat SA, et al. (2007). Large- Scale mtDNA Analysis of Domestic Goat Reveals Six Haplogroups with High Diversity. PLos One 2(10): e1012.
- NCBI (National Center for Biotechnology Information). The nucleotide database.
- Abate A, Wanyoike MM, Badamana MS, Abate AN (1989) Towards Improving Animal Production in the Rangelands. Proceedings of the 16th Grassland Congress, France 2: 1613.
- Tajima F (1989) Statistical methods to test for nucleotide mutation hypothesis by DNA polymorphism. Genetics 123(3): 585-595.
- Nei M, Kumar S (2000) Molecular Evolution and Phylogenetics. Oxford University Press, New York, USA, p. 352.
- Kibegwa FM, Githui KE, Jung’a JO, Badamana MS, Nyamu MN (2016) Mitochodrial DNA Variation of Indigenous Goats in Narok and Isiolo Counties of Kenya. J Anim Breed Genet 133(3): 238-247.
- Luikart G, Gielly L, Excoffier L, Vigne JD, Bouvet J, et al. (2001) Multiple maternal origins and weak phylogeographic structure in domestic goats. Proc Natl Acad Sci USA 98(10): 5927–5932.
- Wolf PG, Soltis PS (1992) Estimates of gene flow among populations, geographic races, and species in the Ipomopsis aggregata complex. Genetics 130(3): 639–647.
- Wright S (1931) Evolution in Mendelian populations. Genetics 16(2): 97–159.
- Joshi MB, Rout PK, Mandal AK, Tyler-Smith C, Singh L, et al. (2004) Phylogeography and origins of Indian domestic goats. Mol Biol Evol 21(3): 454-462.






























