Is Genetic Variability of Coastal Species (Cakile Maritima) Due to Human Effect?
Gandour M1,2, Taamalli W3 and Abdelly C1
1Laboratory of extremophiles plants, Center of Biotechnology of Borj-Cedria, Tunisia
2Faculty of sciences and technology of Sidi Bouzid, University of Kairouan, Tunisia
3Laboratory of olive biotechnology, Center of Biotechnology of Borj-Cedria, Tunisia
Submission: November 17, 2016; Published: December 13, 2017
*Corresponding author: Gandour M, Laboratory of extremophiles plants, Center of Biotechnology of Borj-Cedria, Tunisia,, Email: gandourmed@yahoo.fr
How to cite this article: Gandour M, Taamalli W, Abdelly C. Is Genetic Variability of Coastal Species (Cakile Maritima) Due to Human Effect?. Agri Res &Tech: Open Access J. 2017; 12(5): 555862. DOI: 10.19080/ARTOAJ.2017.12.555862
Abstract
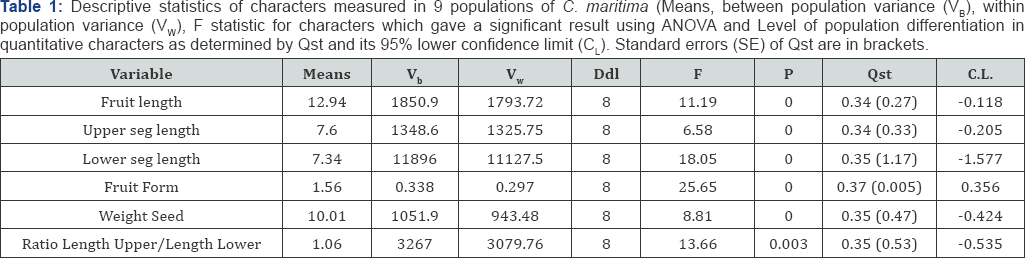
Human impact on natural habitat has significantly increased in the past few decades. Understanding how plant populations respond to such a phenomenon is an important aim of plant ecological research. In the present study, we use phenotypic and biochemical markers to examine the impact of habitat disturbance on the genetic diversity and population structure of Cakile maritima. Both phenotypic and biochemical markers showed high levels of genetic differentiation within and between populations. Populations exhibited strong spatial structure as indicated by the large Pop-φST value of 0.31 (P<0.001) estimated with AMOVA, Indirect estimation of gene flow for the overall population is Nm=0.55. The estimates of population differentiation in quantitative traits (Qst) varied widely among the six quantitative traits (from 34% for fruit length and upper segment length to 37% for fruit form with an average value of 0.35 among all populations].
significant correlation (r=0.74, P=0.026) was found between the two matrices of pair-wise genetic differentiations (quantitative trait values and genetic differentiation). Conversely no correlation was found between genetic differentiations (Fst) and geographic distances based on the Mantel test indicating that the genetic distance did not necessarily correlate with the geographical distance. Despite the fragmentation process, sustain of high genetic diversity suggests that there are some ecological factors determining population structure.
Keywords: Cakile maritime; Genetic variation; Habitat fragmentation; Qst; Fst
Introduction
Understanding how plant populations respond to human impact on natural habitat is an important aim of plant ecological research. Kelly et al. (2003) has shown that plant populations strongly vary in space and time in response to the disturbance in their natural’s habitats. Because the continuous disturbance, the eventual fates of all populations is extinctions [1]. Thus, the regional dynamics of plant species are usually a matter of both local extinction and colonization of new unoccupied sites [2,3]. Today in many ecosystems, fragmentation and degradation of habitats have increased the extinction risk of populations of plant species adapted to specific environmental conditions [4,5]. In particular, spectacular recent growth of urbanism and tourism in coastal areas were widely identified as the most important factor causing coastal plant decline [6-8]. Tourism has had a range of negative direct and indirect impacts on native flora including contributing to the decline of some rare and threatened plant species and communities [9,10]. In this context, direct effects of tourism and recreation on flora may include vegetation clearance for resorts, roads and other infrastructure [11].
Thus, development of hotel and Edinburgh’s shoreline for commercial and leisure purposes has led to the disappearance of much of coastal species habitat and the plant’s subsequent decline by reducing both habitat area and the connectivity of the remaining population fragments identified three ways in which fragmentation can lead to reduction of genetic variation:
(i) Increased losses of populations from genetic drift,
(ii) Increased rates of inbreeding and
(iii) Reduced gene flow between remaining population fragmentations.
During the fragmentation process, distances between populations increase, progressively hampering the arrival of immigrants from surrounding populations until complete isolation is reached. The genetic consequences caused by habitat fragmentation are not yet well understood and further investigation is needed [12,13]. One such susceptible species is the Cakile maritima; a characteristic plant of Tunisian Coastal dune vegetation [14].
C. maritima was probably one of the first members of reinvasion floras after successive glaciations [15]. It grows in sandy habitats along the North Atlantic Ocean, the Mediterranean Sea coasts, the Canary Island and southwest Asia [16]. In these surf and wind. According to, in Tunisia, this species is frequent along the coast from north to south. However these populations have been severely disturbed by urbanization and infrastructure developed for tourists in the past decade. Thus we asked what effect the fragmentation of coastal vegetation might have had on the populations genetics of the C. maritima populations now isolated from each other. To answer this question, we studied the genetic structure and variability in nine C. maritima populations that differ in the intensity of urbanism and tourism development.
Materials and Methods
Plant material collection and studied characters
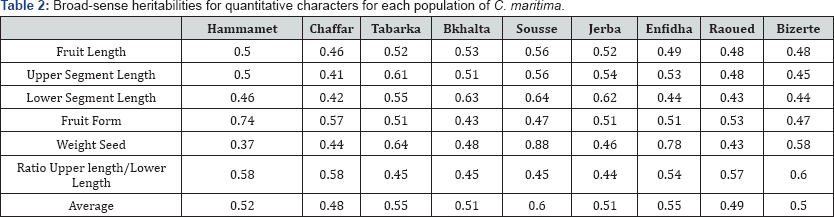
The plant material consisted of nine Tunisian populations of C. maritima. From each population, seed material from 15 randomly chosen individuals were collected and used as a source of genotypic data. Five morphological characters (total fruit length, lower segment length, upper segment length, fruit type, seed weight) judged discriminate traits for Cakile [17] and one ratio were measured for twenty siliques per individual (Table 2) so that 300 siliques per population.
distances between consecutive group of plants were kept to estimate the degree of fragmentation. In fact, distance inferior to 100m between groups was considered as weakly deteriorated area (class 1), distance between 100m to 300m between groups was considered as a sign of area moderately deteriorated (class 2), distance between 300m and 700m between groups indicates a deteriorated area (class 3) and distance superior to 700 m between groups of individuals were considered as a sign of highly deteriorated area (class 4).
Statistical analysis
Variation in phenotypic traits was analysed after running nested ANOVA including population and family (nested within population) as random effect. varcomp was used to estimate population differentiation Qst VB/(VB+VW), where VB is the proportion of genetic variance distributed among populations and VW is the proportion of genetic variance distributed within populations [18-19]. For each of the nine populations one-way ANOVA was performed to estimate broad-sense heritability, h2 = VG/(VG+VE). The within-genotype variance estimated the environmental variance VE and the among-genotype variance provided an estimate of the genetic variance VG. Principal component analysis was conducted on standardized original values using the Multi-Variate Statistical Package (MVSP, Kovach, 1999). Spearman's correlations between variance within each population and the state of origin sites were calculated following appropriate procedures. It is worth noting that Spearman's rank correlation is the best-known procedure for studying the degree of relationship between 2 variables when there is sub-normality in both pairs of variables.
Seed protein extraction and data analysis
Seed storage proteins were extracted from 0.8g of seeds with 800νl of Tris-HCl buffer (1M pH 8.8) and were centrifuged at 9100g for 10 min (centrifuge sigma; rotor 121154). The supernatant of each population was kept and was stored at -20 °C until analysis. A volume of 30νl protein extract was added to equal volume of treatment buffer and boiled at 95 °C for 10 min. Thereafter, 15νl of the crude extract was directly analysed by SDS-bis-Polyacrylamide Gel Electrophoresis using mini gel 8% (w/v) and a low range weight SDS Standards of BIORAD as a marker. Electrophoresis was carried out at 20mA for 2 hours. Gel were stained with 0.5% coomassie Brillant Blue (CBB) R2 50 for about 1 hour then distained in acetic acid-methanol- water (7:20:73 volume ratio) for overnight. The number of bands revealed in each lane of the gel was counted and analysed using gel-pro-system (version 3.1). Only those bands obtained clearly reproducible after three repetitions were scored.
Statistical analysis
The 0/1 matrix obtained from scoring the seed storage proteins markers was subject to analysis of molecular variance (AMOVA) using the Manual Arlequin ver 3.1 package. AMOVA is based on pairwise squared Euclidean distances among seed storage proteins phenotypes, and allows the molecular variance to be partitioned among several hierarchical levels, analogous to conventional analysis of variance [20]. The significance of variance components at each hierarchical level is tested with a permutation procedure. From the variance components, a ^ST statistic can be calculated, which is a good estimator of Fst. ^ST is defined as the proportion of total molecular variance that is due to differences among the units of a given level. Here, we partitioned the total variance into components attributable to differences among populations and among individuals within populations. Pairwise population Fst was used to produce a dendrogram using neighbour joining (DARwin 5.0.148 program). Arlequin program was also used to compute a correlation between pairwise population Fst and genetic distances calculated from quantitative traits (Mantel test [21]) in one hand and between pairwise population Fst and geographic distance (distance measured along the coast or distances measured as a straight line) in other hand. Principal Coordinates Analysis was used to ordinate relationships among individuals and among regions using the MVSP program.
Results
Genetic differentiation
Different electrophoretic patterns were seen for the populations of Cakile maritima. The protein bands analysed were found in the high, average and low molecular weight regions. In total, 14 to 17 bands per population were detected in SDS-PAGE electrophoregrams, ranging from 13 to 150KDa. The degree of variation of each population was analysed by comparing the different phenotypes. Variation in at least one band was taken to indicate two different phenotypes. A population was considered polymorphic when at least one specimen, showed a different phenotype to the rest. Populations were composed of different genotypes. Populations exhibited strong spatial structure as indicated by the large Pop-φST value of 0.31 (P<0.001) estimated with AMOVA, suggesting that about 31% of seed storage protein occurred between populations.
Indirect estimation of gene flow for the overall population is Nm=0.55. The matrix of genetic distance values using pairwise Fst coefficient was built to generate a dendrogram showing seed protein relationship between Tunisian populations of C. maritima. Three groups were identified. First group enclosed populations of Bekalta, Sousse and Jerba, which seem to be similar, the second group included populations of Hammamet, Enfidha, Raoued, Bizerte and Chaffar, and exceptionally the last group was represented with a single population (Tabarka). Genetic distance between population pairs was only weakly associated with geographical distance, as indicated by the small and non-significant Mantel correlation between the two distance matrices. In line with the dendrogram, principal coordinate analysis was performed in order to determine the genetic relationships among the populations with minimum distortion.
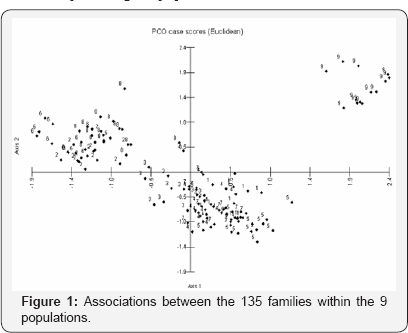
The populations were plotted on principal coordinates 1 and 2, accounting for 22.35% and 14.82% of the variation and together explaining 37.18% of the total variation (Figure 1). Tabarka population (no. 9) in the upper right quarter of the plot is from extreme North West of Tunisia. Three populations (no. 2, 6 and 8) from Sahal and south region are grouped in the upper left quarter of the plot (Figure 1). Chaffar population (no. 3) is between the upper left and lower left quarters. Bizerte, Enfidha, Hammamet and Raoued population (no. 1,4,5 and 7) are in the lower right quarter; they also form a group in the dendrogram and resemble each other more than other population.
Quantitative genetic variation
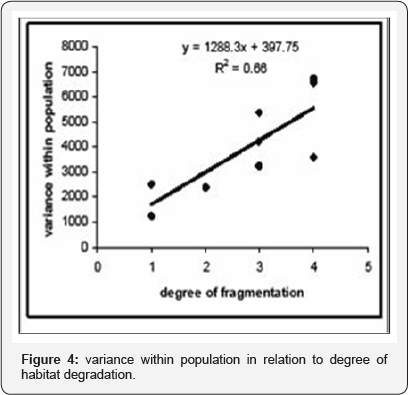
Population means and coefficients of variation for six quantitative traits are shown in Table 2. Population values differed significantly in each trait and over all traits analysed simultaneously. ANOVA also revealed highly significant differences (P<0.001) in each pairwise population comparison. Variation within groups, although different for some traits (as revealed by Levene's test), showed no consistent pattern across the groups. Variance within each population was highly correlated with degree of habitat fragmentation (r =0.81, P=0.0022) (Figure 4). Heritability of all traits was high, ranging 0.48-0.60 when averaged over nine populations, but with striking differences among populations. Estimates of population differentiation in quantitative traits (Qst) varied widely among the six quantitative traits (Table 1). Values of Qst varied from 34 % for fruit length and upper segment length to 37 % for fruit form with an average value of 0.35 among all populations (Table 2).
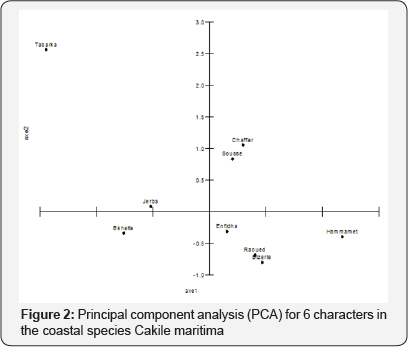
Principal component analysis can determine which of the characters most strongly contributes to the PC. The analyses reduced the original 6 characters in the experiment to 3 derived variable or components. The first 3 PCs with eigen values >1 explained 90.89% of the variation among the 9 populations. PC1, which is the most important component, explained 45% of the total variation and was positively related to total fruit length and lower segment length, is therefore a weighted average of these two characters; PC2 accounted for 29.03% of the total variation and the characters with the greatest weight on this component were upper segment length and ratio of the two segments length; PC3 was mainly related to seed weight. The entries in this study were grouped into 4 clusters based on average linkage. Cluster I consisted of one population, cluster II of two, cluster III of three and cluster IV of four populations. The first two PC scores were plotted to aid visualization of group differences (Figure 1).
Relationships between molecular and quantitative trait variation
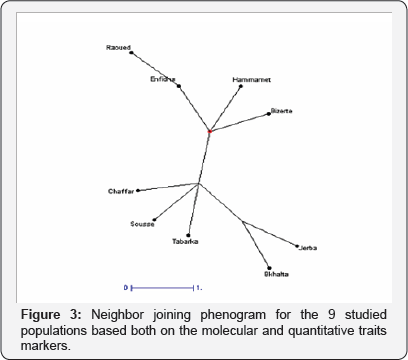
A test of the relationships between two matrices of pair wise genetic distances (quantitative trait values and seed storage protein allele frequencies) calculated from 9 populations revealed a highly significant relationship (r=0.74, P=0.026, Mantel test). Conversely no correlation was found between genetic distances (Fst) and geographic distances based on the Mantel test indicating that the genetic distance did not necessarily correlate with the geographical distance since the genetic distance among geographically distant populations was large and those among close populations were small. Although a common analysis of molecular data and morphological one was established (tree comparison methods), thus, it has demonstrated that morphological diversity paralleled molecular one Figure 3.
Discussion
Many animal and plant species have strongly declined during the last decades as a result of changes in land use [5]. Understanding the consequences of habitat change for population viability and long-term persistence of both animal and plant species is a major goal of conservation research [22-23]. One of the most species in Tunisian coast is Cakile maritima, but many of their habitats have been severely affected by human activities (urbanism, agriculture and tourism) and a fragmentation has occurred. The negative effects of habitat degradation may be exacerbated by the effects of habitat fragmentation that result in increased isolation and reduced size and density of plant populations [24].
Because of their greater sensitivity to habitat disturbance, environmental and demographic stochasticity, small populations face an increased risk of extinction [25]. Moreover, small and isolated populations may suffer from genetic erosion [26], and reduced genetic diversity has been found to reduce offspring fitness [27] and population viability [28]. However, many studies based on allozymes have shown that even narrowly restricted species may maintain high levels of diversity [29-31]. According to those authors, this study show that Cakile maritima maintain a high degree of genetic diversity. In fact, in the most variable populations each specimen was of a different phenotype. Interpopulation variability was also high and specimens with identical phenotypes were not found in different populations. Accordingly the analysis of these populations with morphological characters indicates a high degree of intra and inter-population variation.
In fact, the Qst calculated in each character was significantly different from zero. Among other factor, population variability is related to the method of reproduction (preferably out-crossing). This has been shown in numerous species and is reflected by the positive association between the amount of out-crossing and genetic variation [32]. According to, between population diversity values <20% are usually characteristic of out-crossing species, while those > 50% would apply for inbreeding species. The high degree of intra-population variability in some of the populations of this study might be attributed to a certain degree of out-crossing (65% according) to a phenomenon observed in other species [32] or probably due in part to continuous germination through the year [33], which creates a mixed age structure.
It can be also explained by assuming that such long-distance dispersal events have occurred and have led to diversification [34]. Zohary 1965 suggest that the high degree of variation in species able to colonize different habitat could be the result of hybridisation and recombination of different genomes of common origin. From an early analysis of phenotypic and biochemical (seed oil content) diversity of Tunisian Cakile populations in 2001 published it appear not only a significant difference between population but also an important ecogeographic effects. In recent years, the habitats of Cakile maritima have been devastated by human activities, and this has led to massive elimination of population of this species. Our results show that these effects of reduced population size, although significant, are too small to affect genetic diversity and maybe too small to affect seed dispersal ability. This finding is in agreement with sensitivity analyses of mechanistic wind dispersal models, which showed that seed dispersal distances are not very sensitive to intraspecific variation in seed terminal velocity [35]. Thus, habitat fragmentation does not affect plant seed dispersal ability by reducing the size of isolated populations.
Conclusion
For species featuring high levels of genetic differentiation between populations, the loss of a population can cause the irreversible loss of genetic variation that is not shared with the other populations [36-37]. Based on our data, the extent of the differences between the various populations seems to be weakly correlated to human impact differences, corrected for a putative sea seed transport. Nevertheless, the loss of its natural habitat due to excessive and unplanned tourism development can have consequences in the long term. Thus, it will be important to precise the genetic character of the population differences, using neutral molecular markers, and assessing the heritability of the main traits [38-41]. In a second step, the ecological significance of the phenotypic differences will be studied through fitness assessment
References
- Lande R, Engen S, Saether BE (2003) Stochastic population dynamics in ecology and conservation. Oxford University Press, Oxford, UK.
- Hanski I (1999) Metapopulation Ecology. Oxford University Press, Oxford, UK.
- Eriksson O, Ehrlen J (2001) Landscape fragmentation and the viability of plant populations. In: Silvertown J, Antonovics J (Eds.), Integrating ecology and evolution in a spatial context. Blackwell, Oxford, UK, pp. 157-175.
- Pitman NCA, J0rgensen PM (2002) Estimating the size of the world's threatened flora. Science 298(5595): 989.
- Baillie JEM, Hilton-Taylor C, Stuart SN (2004) Red List of Threatened Species. A Global Species Assessment. IUCN, Gland, Cambridge, UK.
- Mayol J (1995) Changements socio-economiques et conservation de la flore dans les lles de la mediterranee. Ecol. Medit. 21: 337-344.
- Mus M (1995) Conservation of flora in the Balearic Islands. Ecol Medit p. 21.
- Sih A, Jonsson BG, Luikart G (2000) Habitat loss: ecological, evolutionary and genetic consequences. Trends Ecol Evol 15(4): 132-134.
- Liddle M (1997) Recreation Ecology: the Ecological impact of Outdoor Recreation and Ecotourism. Chapman and Hall, London.
- Buckley R (1999) Tourism in the most fragile environments. Tourism and Recreation Research 25(1): 31-40.
- Kelly CL, Peckering CM, Buckley RC (2003) Impacts of Tourism on threatened plant taxa and communities in Australia. Ecological Management and Restoration 4(1): 37-44.
- Young AG, Merriam HG, Warwick SL (1993) The effects of forest fragmentation on genetic variation in Acer saccharum Marsh. (sugar maple) populations. Heredity 71: 277-289.
- Kang M, Jiang M, Huang H (2005) Genetic Diversity in Fragmented Populations of Berchemiella wilsonii var. pubipetiolata (Rhamnaceae). Ann Bot 95(7): 1145-1151.
- Pottier-Alapetite G (1979) Flore de la Tunisie Angiospermes- Dicotyledones; Apetales-Dialypetales (premiere partie).
- Davy AJ, Scott R, Cordazzo CV (2006) Biological flora of the British Isles: Cakile maritima. Journal of ecology 94(3): 695-711.
- Clausing G, Vickers K, Kadereit JW (2000) Historical biogeography in a linear system: genetic variation of Sea Rocket (Cakile maritima) and Sea Holly (Eryngium maritimum) along European coasts. Molecular Ecology 9(11): 1823-1833.
- Cody ML, Cody TWD (2004) Morphology and spatial distribution of alien sea-rockets (Cakile spp.) on South Australian and Western Canadian beaches. Australian Journal of Botany 52: 175-183.
- Spitze K (1993) Population structure in Daphnia obtusa: quantitative genetics and allozymic variation. Genetics 135(2): 367-374.
- Morgan KK, Hicks J, Spitze K, Latta L, Pfrender ME, et al. (2001) Patterns of genetic architecture for life-history traits and molecular markers in a subdivided species. Evolution 55(9): 1753-1761.
- Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: applications to human mitochondrial DNA restriction data. Genetics 131(2): 479-491.
- Mantel N (1970) The detection of disease clustering and generalized regression approach. Cancer Res 27(2): 209-220.
- Debinski DM, Holt RD (2000) A survey and overview of habitat fragmentation experiments. Conservation Biology 14(2): 342-355.
- Henl, K, Lindenmayer DB, Margules CR, Saunders DA, Wissel C (2004) Species survival in fragmented landscapes: where are we now? Biodiversity and Conservation 13(1): 1-8
- Lienert J, Fischer M (2003) Habitat fragmentation affects the common wetland specialist Primula farinosa in north-east Switzerland. Journal of Ecology 91(4): 587-599.
- Matthies D, Brauer I, Maibom W, Tscharntke T (2004) Population size and the risk of local extinction: empirical evidence from rare plants. Oikos 105(3): 481-488.
- Young A, Boyle OD, Brown T (1996) The population genetic consequences of habitat fragmentation for plants. Trends in Ecology and Evolution 11: 413-418.
- Leimu R, Mutikainen P, Koricheva J, Fischer M (2006) How general are positive relationships between plant population size, fitness and genetic variation? Journal of Ecology 94(5): 942-952.
- Frankham R (2005) Genetics and extinction. Biological Conservation 126: 131-140.
- Sharma IK, Jones DL, French CJ (2003) Unusually high genetic variability revealed through allozymic polymorphism of an endemic endangered Australian orchid, Pterostylis aff. picta (Orchidaceae). Biochemical Systematics and Ecology 31(5): 513-526.
- Lewis PO, Crawford DJ (1995) Pleistocene refugium endemics exhibit greater diversity than widespread congeners in the genus Polygonella (Polygonaceae). American Journal of Botany 82(2): 141-149.
- Ranker TA (1994) Evolution of high genetic diversity in the rare Hawaiian fern Adenophorus periens and implications for conservation management. Biological Conservation 70: 19-24.
- Hamrick JL, Linhart YB, Mitton JB (1979) Relationship between life history parameters and allozyme variation in plants. Ann Rev Ecol Syst 10: 173-200.
- Cordazzo CV (1994) Comparative population studies of four dominant plants of southern Brazilian coastal dunes. University of East Anglia, Thesis, Norwich.
- Gandour M, Hessini K, Abdelly C (2008) Understanding the population genetic structure of coastal species (Cakile maritima): seed dispersal and the role of sea currents in determining population structure. Genetics research 90: 167-178.
- Soons MB, Heil GW, Nathan R, Katul G (2004) Determinants of long-distance seed dispersal by wind grasslands. Ecology 85(11): 30563068.
- Hossaert M, Valero MM, Magda D, Jarry M, Cuguen J, et al. (1996) The evolving genetic history of a population of Lathyrus sylvestris: Evidence from temporal and spatial genetic structure. Evolution 50(5): 1808-1821.
- Chung MY, Epperson BK, Chung MG (2003) Genetic structure of age classes in Camellia japonica (Theaceae). Evolution 57(1): 62-73.
- Bussell J (1999) The distribution of random amplified polymorphic DNA (RAPD) diversity amongst populations of Isotoma petraea (lobeliaceae). Molecular Ecology 8(5): 775-789.
- Saunders DA, Hobbs RJ, Margules CR (1991) Biological consequences of ecosystem fragmentation: a review. Biological Conservation 5(1): 18-32.
- Thrall PH, Young AG,Burdon JJ (2000) An analysis of mating structure in populations of the annual sea rocket, Cakile maritima (Brassicaceae). Aust J Bot 48(6): 731-738.
- Zohary D (1965) Colonizer species in the wheat group. In: Baker HG, Stebbins GL (Eds.), The genetics of colonizing species. Academic Press, pp. 403-423.






























