Prospects of Novel Species of Oral Veillonella in Human Saliva
Citra F Theodorea1, 2, Izumi Mashima13,4 and Futoshi Nakazawa1*
1Department of Oral Microbiology, Health Sciences University of Hokkaido, Japan
2Department of Oral Biology, Universitas Indonesia, Indonesia
3Postdoctoral Fellow of Japan Society for the Promotion of Science, Japan
4Department of Oral Biology, University at Buffalo, The State University of New York, USA
Submission: August 3, 2017; Published: August 22, 2017
*Corresponding author: Futoshi Nakazawa, MS, Ph D, Department of Oral Microbiology, School of Dentistry, Health Sciences University of Hokkaido, 1757 Kanazawa, Ishikari-Tobetsu, Hokkaido, 061-0293, Japan, Tel +81-133-23-2484; Fax: +81-133-23-1385; Email: nakazawa@hoku-iryo-u.ac.jp
How to cite this article: Citra F T, Izumi M, Futoshi N. Prospects of Novel Species of Oral Veillonella in Human Saliva. Adv Biotech & Micro. 2017; 5(4): 555667. DOI: 10.19080/AIBM.2017.05.555667
Abstract
Although 13 species of bacteria have been established in the genus Veillonella, only 6 have currently been isolated from human oral cavities (V. atypica, V denticariosi, V. dispar, V. parvula, V rogosae, and V tobetsuensis). Oral Veillonella have played a central role as early colonizers that establish multispecies oral biofilms that cause many oral infectious diseases in humans such as periodontitis and dental caries. Our previous study isolated 1,609 strains (confirmed by PCR with the genus-specific primer set) of the genus Veillonella from saliva samples of 107 Thai children. Although 167 strains could not be classified under any oral Veillonella species, 1,442 strains were linked to the 6 oral Veillonella species by PCR with species-specific primer sets. To reveal the phylogenetic characteristics of these unclassified Veillonella strains, PCR-based amplifications and sequence analyses of rpoB genes were performed for 23 representative strains. When the phylogenetic tree was examined based on the obtained sequence data, these 23 strains formed a cluster distinct from the 13 established species of the genus Veillonella with a robust bootstrap value. Thus, the phylogenetic study demonstrated the prospects of several novel species of oral Veillonella in human saliva.
Keywords: Novel species; Oral biofilm; Oral Veillonella; Phylogenetic tree; rpoB gene
Abbreviations: PCR: Polymerase Chain Reaction
Introduction
The genus Veillonella consists of small, strictly anaerobic, gram-negative cocci lacking flagella, spores, and capsules that are characterized by their ability to obtain energy from short- chain organic acids [1,2]. Although 13 species have currently been established in the genus VeiIIoneIIa, only 6 species (V. atypica, V denticariosi, V dispar, V parvula, V rogosae, and V. tobetsuensis) have been isolated from human oral cavities [2-9]. The main habitats of the oral Veillonella are tongue biofilms, dental biofilms, buccal mucosa, and saliva [10-12]. Oral Veillonella has been found in severe early childhood caries [13], apical root canals [14], and dental tubules [15]. Additionally, oral Veillonella are also predominantly found in saliva [16] and subgingival biofilm specimens [17] from patients who have chronic periodontitis.
It was reported that oral Veillonella play a central role as early colonizers that establish multispecies oral biofilm communities [18]. Oral biofilms are known to cause many oral infectious diseases, such as periodontitis and dental caries in humans [2,19]. It was reported that oral VeiIIoneIIa, found throughout the entire oral cavity, comprise as much as 10% of the bacterial community initially colonizing the enamel [20, 21], and often form biofilms with the Streptococcus species [22]. Some studies have also shown that during the formation of early dental plaque, the ratio of Veillonella to Streptococcus species varied in mixed-microbial colonies [23].
The identification of Veillonella isolates at the genus level is relatively simple, but identification at the species level remains difficult because of the lack of conventional phenotypic and biochemical tests [24]. However, molecular methods based on the 16S rRNA gene sequencing have previously been used to identify Veillonella strains at the species level [25,26]. Unfortunately, recent studies have demonstrated that using 16S rRNA gene sequencing exclusively for identification purposes is unreliable because of the high levels of sequence conservation in the 16S rRNA gene sequence among several Veillonella species [5,8,9,25]. To overcome this difficulty, Beighton et al. [27] used the rpoB gene sequence to identify isolates of the genus Veillonella from the human tongue at the species level. Furthermore, our previous study established the first successful one-step PCR method using species-specific primer sets based on the highly variable region of the rpoB gene of 6 species of oral Veillonella (positions 2,500-3,100)[12]. This one-step PCR method is easier and more effective than the previous molecular methods described above in identifying oral Veillonella at the species level.
Our previous study examined the distribution and frequency of oral Veillonella species in the saliva of 107 Thai children with varying oral hygiene statuses [28]. PCR with the genus- specific primer sets confirmed 1,609 strains, isolated in the study, as members of the genus Veillonella [27]. Furthermore, 1,442 strains were identified by the one-step PCR method as V atypica, V denticariosi, V dispar, V parvula, V rogosae, or V. tobetsuensis. However, 167 strains could not be classified into any oral Veillonella species. This short communication seeks to report the phylogenetic position and diversity of 23 of the 167 unclassified Veillonella strains by phylogenetic analysis with the rpoB gene sequence.
23 unclassified Veillonella strains-S22108-3a, S34277-12b, S42329-13a, S31177-14a, S12034-12, S11011-11d, S44377-5b, S13062-15, S13054-9, S44377-5a, S34277-12a, S22118-11a, S33222-2c, S42329-8b, S43355-15, S34284-15, S14073-19a, S24170-5, S31180-9b, S14073-19c, S32198-4a, S42329-8a, S31177-14b-were selected as representative strains and cultured in BactoTM Brain Heart Infusion (Difco Laboratories, BD) broth supplemented with 5% (v/v) defibrinated sheep blood (BHI agar), hemin (10μg/mL), and menadione (5μg/mL) under anaerobic conditions with 80% N2, 10% CO2, and 10% H2 at 37 °C for 5d.
Genomic DNA was extracted from individual bacterial cells using an Insta Gene Matrix Kit (Bio-Rad) and PCR-based amplification and sequence analyses of rpoB gene was performed using previously described primers [27]. The sequences were determined by an ABI PRISM 310 Genetic Analyzer (Applied Biosystems) and were aligned and connected by using SEQMAN Pro of the LASERGENE 14 program (DNASTAR). The programs MEGALIGN including CLUSTAL W using the neighbor-joining (NJ) method [29] were used to compare sequences and to reconstruct an evolutionary tree. The rpoB sequences of the 23 unclassified Veillonella strains were aligned against the sequences of the representative Veillonella type strains retrieved from GenBank.
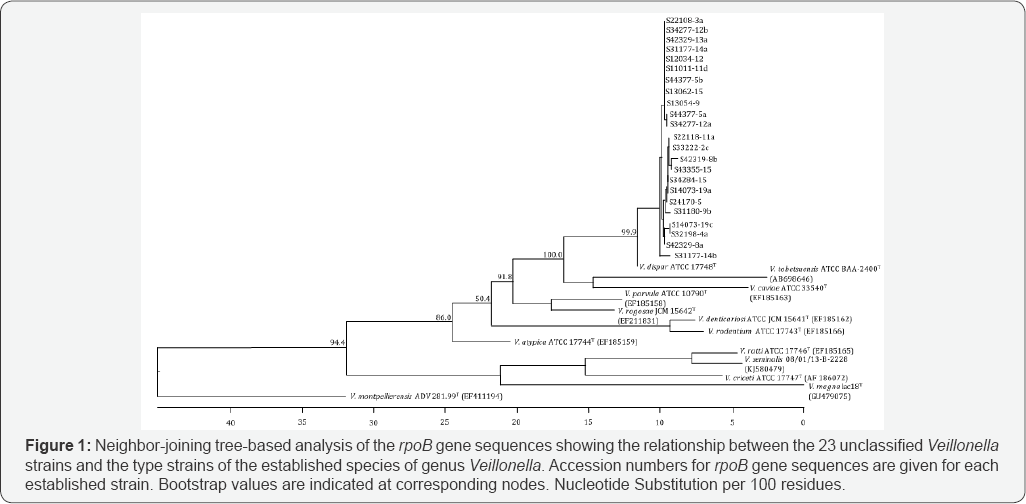
Alignment of the rpoB gene sequences showed 97.6-100% similarity among the sequences of the unclassified 23 Veillonella strains. These results indicate that these 23 strains are closely related. The evolutionary tree obtained for the type strain of the established species of genus Veillonella and the unclassified 23 Veillonella strains is shown in Figure 1. Based on the tree, the 23 unclassified Veillonella strains formed a cluster distinct from the established Veillonella species. Although the most closely related species was V. dispar, the robust bootstrap value of 99.9% indicated that these 23 strains formed a taxon distinct from V dispar.
Conclusion
This study demonstrated prospective novel oral Veillonella species in human saliva. It is expected that the new perspective of oral biofilm communities at early stages would be investigated by this discovery of novel Veillonella species.
References
- Carlier JP (2015) Veillonella. Bergey's Manual of Systematic of Archaea and Bacteria, Wiley online library, USA, pp. 1-11.?
- Delwiche EA, Pestka JJ, Tortorello ML (1985) The Veillonellae: gram negative cocci with a unique physiology. Annu Rev Microbiol 39: 175193.
- Rogosa M (1965) The genus VeiIIoneIIa IV. serological groupings, and genus and species emendations. J Bacteriol 90(3): 704-709.
- Jumas-Bilak E, Carlier JP, Jean-Pierre H, Teyssier C, Gay B, et al. (2004) Veillonella montpellierensis sp. nov., a novel, anaerobic, Gram-negative coccus isolated from human clinical samples. Int J Syst Evol Microbiol 54(Pt 4): 1311-1316.
- Byun R, Carlier JP, Jacques NA, Marchandin H, Hunter N (2007) Veillonella denticariosi sp. nov., isolated from human carious dentine. Int J Syst Evol Microbiol 57(Pt 12): 2844-2848.
- Arif N, Do T, Byun R, Sheehy E, Clark D, et al. (2008) Veillonellarogosae sp. nov., an anaerobic, Gram-negative coccus isolated from dental plaque. Int J Syst Evol Microbiol 58(Pt 3): 581-584.
- Kraatz M, Taras D (2008) Veillonella magna sp. nov., isolated from the jejunal mucosa of a healthy pig, and emended description of Veillonellaratti. Int J Syst Evol Microbiol 58(Pt 12): 2755-2761.
- Mashima I, Kamaguchi A, Miyakawa H, Nakazawa F (2013) Veillonellato betsuensis sp. nov., an anaerobic, gram-negative coccus isolated from human tongue biofilms. Int J Syst Evol Microbiol 63(Pt 4): 1443-1449.
- Aujoulat F, Bouvet P, Jumas-Bilak E, Jean-Pierre H, Marchandin H (2014) Veillonella seminalis sp. nov., a novel anaerobic Gram-stain-negative coccus from human clinical samples, and emended description of the genus Veillonella. Int J Syst Evol Microbiol 64(Pt 10): 3526-3531.
- Arif N, Sheehy EC, Do T, Beighton D (2008) Diversity of Veillonella spp. from sound and carious sites in children. J Dent Res 87(3): 278-282.
- Hughes CV, Kolenbrander PE, Andersen RN, Moore LV (1988) Coaggregation properties of human oral Veillonella spp.: relationship to colonization site and oral ecology. Appl Environ Microbiol 54(8):1957- 1963.
- Mashima I, Theodorea CF, Thaweboon B, Thaweboon S, Nakazawa F(2016) Identification of Veillonella Species in the Tongue Biofilm by Using a Novel One-Step Polymerase Chain Reaction Method. PLoS One 11(6): e0157516.
- Kanasi E, Dewhirst FE, Chalmers NI, Kent R, Moore A, et al. (2010) Clonal analysis of the microbiota of severe early childhood caries. Caries Res 44(5): 485-497.
- Baumgartner JC, Falkler WA (1991) Bacteria in the apical 5mm of infected root canals. J Endod 17(8): 380-383.
- Peters LB, Wesselink PR, Buijs JF, van Winkelhoff AJ (2001) Viable bacteria in root dentinal tubules of teeth with apical periodontitis. J Endod 27(2): 76-81.
- Takeshita T, Nakano Y, Kumagai T, Yasui M, Kamio N, et al. (2009) The ecological proportion of indigenous bacterial populations in saliva is correlated with oral health status. ISME J 3(1): 65-78.
- Heller D, Silva-Boghossian CM, do Souto RM, Colombo AP (2012) Subgingival microbial profiles of generalized aggressive and chronic periodontal diseases. Arch Oral Biol 57(7): 973-980.
- Periasamy S, Kolenbrander PE (2010) Central role of the early colonizer Veillonella sp. in establishing multispecies biofilm communities with initial, middle, and late colonizers of enamel. J Bacteriol 192(12): 2965-2972.
- Mashima I, Fujita M, Nakatsuka Y, Kado T, Furuichi Y, et al. (2015) The Distribution and Frequency of Oral VeiIIoneIIa spp. Associated with Chronic Periodontitis. Int J Curr Microbiol App Sci 4(3): 150-160.
- Aas JA, Paster BJ, Stokes LN, Olsen I, Dewhirst FE (2005) Defining the normal bacterial flora of the oral cavity. J Clin Microbiol 43 (11): 57215732.
- Diaz PI, Chalmers NI, Rickard AH, Kong C, Milburn CL, et al. (2006) Molecular Characterization of Subject-Specific Oral Microflora during Initial Colonization of Enamel. Appl Environ Microbiol 72(4): 28372848.
- Olson JC, Cuff CF, Lukomski S, Lukomska E, Canizales Y, et al. (2011) Use of 16S ribosomal RNA gene analyses to characterize the bacterial signature associated with poor oral health in West Virginia. BMC Oral Health 11: 7.
- Chalmers NI, Palmer RJ Cisar JO, Kolenbrander PE (2008) Characterization of a Streptococcus sp.-Veillonella sp. Community Micromanipulated from Dental Plaque. J Bacteriol 190(24): 81458154.
- Kolenbrander PE, Moore LVH (1992) The genus Veillonella. In: The Prokaryotes (2nd edn), Springer, New York, USA, pp. 2034-2047.
- Marchandin H, Teyssier C, Simeon De Buochberg M, Jean-Pierre H, Carriere C, et al. (2003) Intra-chromosomal heterogeneity between the four 16S rRNA gene copies in the genus Veillonella: implications for phylogeny and taxonomy. Microbiology 149(Pt 6): 1493-1501.
- Sato T, Sato M, Matsuyama J, Hoshino E (1997) PCR-restriction fragment length polymorphism analysis of genes coding for 16S rRNA in Veillonella spp. Int J SystBacteriol 47(4): 1268-1270.
- Beighton D, Clark D, Hanakuka B, Gilbert S, Do T (2008) The predominant cultivable Veillonella spp. of the tongue of healthy adults identified using rpoB sequencing. Oral Microbiol Immunol 23(4): 344347.
- Theodorea CF, Mashima I, Thaweboon B, Thaweboon S, Nakazawa F(2017) Molecular Detection of Oral Veillonella species in the Saliva of Children with Different Oral hygiene Statuses. Int J Curr Microbiol Appl Sci 6(7): 449-461.
- Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. MolBiol Evol 4(4): 406-425.






























