Survival of Probiotic E. Coli and Ent. Faecalis in the Human Host after Oral Intake: Results from in Vitro and in Vivo Studies
Trudy M Wassenaar1*, Massimo Marzorati2,3, Claudia Beimfohr4, Alexander Siegl4 and Kurt Zimmermann5
1Molecular Microbiology and Genomics Consultants, Germany
2University of Ghent (LabMET), Belgium
3ProDigest BVBA, Belgium
4Vermicon AG, Germany
5SymbioPharm GmbH, Germany
Submission: February 9, 2017;Published: March 15, 2017
*Corresponding author: Trudy M Wassenaar, Molecular Microbiology and Genomics Consultants, Tannenstrasse 7, 55576 Zotzenheim, Germany, Tel: +49-6701-8531; Fax: +49-67-1-901803, Email:trudy@mmgc.eu
How to cite this article: Edward C. Addiction Needs Neuroscience, Gene Sequencing and Hard Research in Seeking A Cure. Glob J Add & Rehab Med. 2017; 1(5): 555592. DOI:10.19080/AIBM.MS.ID.555592
Abstract
The aim of this study was to determine if commercially available strains of probiotic bacteria belonging to the species Ent. faecalis and E. coli were able to survive passage through the human stomach and colonise the gastrointestinal tract (GIT). Survival of bacteria following exposure to gastric pH levels was assessed using a dynamic in vitro model resembling conditions in the stomach as well as the SHIME® model more closely mimicking the upper GIT environment (stomach and small intestine) during fed conditions.
Viability of both Ent. faecalis DSM 16431 and E. coli DSM 17257 decreased during acid exposure. However, subsequent exposure to simulated small intestine conditions resulted in no further decrease. A human volunteer took a single dose of probiotic Ent. faecalis, after which live bacteria were determined for 6 days in faeces by means of targeted cultivation and identification of colonies using species- and strain-specific PCR primers. Detection of the strain’s DNA by PCR in stool samples was positive for 4 days. This duration of colonisation was much shorter compared to previously determined human colonisation by E. coli DSM 17257.
In conclusion, probiotic E. coli and Ent. faecalis are susceptible to gastric pH, which reduces their viability with several logs. However, sufficient numbers survive to colonise the gut, so that the bacteria are detected in the stool for several days (Ent. faecalis) and multiple weeks (E. coli) following a single dose.
Keywords: Probiotics; GIT; Passage; Colonisation; Human gut; Survival
Abbreviations: GIT : Gastro-Intestinal Tract; SHIME: Simulation of the Human Intestinal Microbial Ecosystem; Ent: Enterococcus
Introduction
According to the definition proposed by the FAO, probiotics are living organisms that, when ingested in sufficient amounts, are beneficial for the host [1]. For probiotic bacteria to produce the desired beneficial effect, they presumably have to colonise the human intestine for a short period at least, although some would argue probiotics don’t have to be alive to be beneficial [2]. Indeed, bacterial lysates are also marketed for presumed beneficial health effects, but their effects are different from those of products based on living bacteria, even when comparing identical strains and species, as the human body treats abiotic bacterial components more likely as food particles than as members of the intestinal microbiota. In vitro studies suggest that peripheral blood mononuclear cells exposed to bacterial components, such as
LPS- and CpG-motif containing oligonucleotides or bacterial lysates respond differently compared to exposure to live probiotic bacteria [3]. For viable probiotic bacteria, surviving the passage through the gastro-intestinal tract (GIT) is essential in order to produce beneficial effects to the host. A classical strategy to improve the viability of probiotic strains is to deliver the bacteria in capsules to protect the cells during stomach passage, (e.g. Mutaflor, containing E. coli strain Nissle 1917). Products can also contain suspensions of viable cells. How well the latter survive passage through the human GIT was addressed here by the use of two in vitro models, a simple model resembling passage through the GIT and a more sophisticated model mimicking the in vivo conditions at a fed stage as closely as possible, using the SHIME technology platform.
The bacteria used in these experiments were Enterococcus faecalis DSM 16431 (the constituent of Symbioflor 1), as an example of a Gram-positive probiotic species, and E. coli DSM 17257 (present in Symbioflor 2), as an example of Gramnegative probiotic bacteria. Randomised, blind clinical trials have shown that Symbioflor 1 treatment can improve conditions such as chronic recurrent bronchitis [4] and chronic recurrent hypertrophic sinusitis [5]; the latter study was positively evaluated in a subsequent systematic review [6]. Symbio flor 2 was compared to placebo in randomized double-blind clinical trials for treatment of IBD in adults [7] and children [8]. In all these studies positive effects were demonstrated, and it has been shown that the products elicit a humoral immunological response [9,10]. Nevertheless, a mechanistic explanation of their beneficial effects relies on at least temporal survival in the gut, and this had not been demonstrated.
The application of a precisely controlled continuous model presented here allowed a study of the processes occurring in the human gastrointestinal tract under controlled conditions. The findings were compared to an in vivo experiment, in which a human volunteer orally took a known dose of viable cells, after which survival of the bacteria was determined in stools over time. Highly specific PCR primers in combination with a molecular FISH probe targeting E. faecalis were used to ensure the detected bacteria were indeed identical to the probiotic strain.
Materials and Methods
The following probiotic products were used in this study: Symbioflor 1, containing a suspension of approximately 2x107 CFU mL-1 E. faecalis DSM 16440 (SymbioPharm GmbH, Herborn, Germany); Symbioflor 2 containing approximately 2x107 CFU mL-1 E. coli DSM 17252 (SymbioPharm GmbH, Herborn, Germany).
Gastric exposure model
Exposure to gastric conditions was first tested in a model described previously [11]. The model allows simulation of stomach, duodenal, jejunal and ileal conditions. Two batches of Symbioflor 1 Ent. Faecalis (numbers 1383 and 1412) and two batches of Symbioflor 2 E. coli (numbers 2115 and 2124) were used to test their viability under conditions simulating the stomach and ileum. Viability of bactteria in Ent. faecalis batch 1383 was 10 times lower than that of batch 1412 at the start of the experiment. For the two E. coli batches there was no significant difference in presence of viable cells. Bacteria were followed for 180min in stomach conditions while a fraction (1/10 per 5min) was passed on after 35min to reach the ileum conditions, which were measured from t=60min till t=360min. During gastric conditions the pH was lowered from 4.8 to 1.7, reaching pH 2.0 after 60min. Bile salts were increased in ileal conditions from 0 to 6mmol L-1 in 300min. Further details are described in [11]. Viability counts were performed as described below.
Simulation of the Human Intestinal Microbial Ecosystem (SHIME) model The SHIME model was used, which simulates the physiological conditions that occur in a typical human GIT. The tests were conducted under conditions simulating a full stomach.
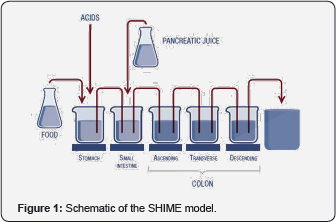
The reactor setup was adapted from the SHIME model as recently described [12]. The SHIME model consists of a succession of five reactors simulating the physiological conditions of the different parts of the human GIT, as visualized in Figure 1. The first two reactors simulate the stomach and the small intestine, and incubation of the probiotic bacteria was restricted to these steps. In the stomach, SHIME nutritional medium was used to simulate food intake (see below). Subsequently, standardized pancreatic and bile liquid was added for the simulation of small intestinal conditions. Incubation conditions, retention time and pH were chosen in order to resemble in vivo conditions in the upper part of the gastrointestinal tract under fed conditions.
In the fed state, the SHIME nutritional medium contains 1gL-1 Arabinogalactan, 2gL-1 Pectin, 1gL-1 Xylan, 4gL-1 Starch, 0.4gL-1 glucose, 3gL-1 Yeast extract, 1gL-1 Peptone, 4gL-1 Mucin and 0.5gL-1 Cysteine (Sigma-Aldrich, Schnelldorf, Germany). Further details are available elsewhere [13]. Under these conditions, the production of pepsin in the stomach increases and the pH decreases over time according to typical sigmoidalshaped curve from 4.5 to 1.8. Thus, 2gL-1 Pepsin was gradually added over a time period of 10min. while the pH was gradually decreased from 4.2 to 1.8 over an incubation time of 90 min. Samples were taken at 45min and 90 min. Intestinal pH values are naturally neutralized with bicarbonate ions secreted by the pancreas, resulting in a pH gradient in the small intestine from pH 5 in the duodenum to pH 7 or higher in the small intestine. Bile salt concentrations under fasted conditions are typically in the range of 5-6mmolL-1 [14].
Approximately 1-2L of pancreatic juice is secreted into the duodenum over a 24h period, while bile salt excretion totals about 600mL/day. This translates to approximately 50-100mL secretion per hour. The transit time through the small intestine varies little for liquids and solids, different doses or between fasted and fed status: the general residence time is in the range of 2-3h [15]. Based on these considerations, the following conditions were applied in the simulated small intestine: to the medium, NaHCO3 was added to give 745mmolL-1, together with gradual addition of 6gL-1 Bile (Oxgall, BD Biosciences, Heidelberg, Germany), corresponding with an initial bile salt concentration of 9mmolL-1, decreasing to 4.5mmolL-1 in 100min. Pancreatin (0.9gL-1) was added (Sigma Aldrich, Schnelldorf, Germany), as a representative mixture of enzymes secreted by the pancreas. The pH was increased in 20 min to a stable value of pH 6.8. An incubation period of 60min, 120min and 180min were sampled. The experiments were performed in triplicate. Viability counts were performed on Brain Heart Infusion (BHI) Agar plates (BD, Heidelberg, Germany) incubated at 37 °C for 24hrs.
Human exposure experiment
Under supervision of a physician, a healthy volunteer (male, BMI 23, 65 yrs of age), took 10ml (10 times the recommended dose) on a full stomach, corresponding to 2x108 CFU of the Ent faecalis strain. The volunteer was fully informed and had given written consent. Stool samples were collected on a daily basis for 8 days and analysed via PCR, cultivation, and fluorescence in situ hybridisation (FISH) as follows. For PCR, DNA was extracted from the stool samples using High Pure PCR Template Preparation Kit (Roche Diagnostics, Mannheim, Germany) and amplified using the highly strain-specific primers targeting gene EFS1_2450 of Symbioflor 1 Ent. faecalis bacteria, described by [16]. Stool samples were also culturally analysed for the presence of Ent. faecalis on CATC agar plates, incubated for 4 hr at 35 °C, followed by 20hr at 37 °C. The PCR mentioned above was used to identify Symbioflor 1 Ent. faecalis bacteria. FISH experiments were performed on stool samples for the simultaneous visualization of Enterococcus spp. and Ent. faecalis using the 16S rRNA targeting probes Enc131 and Efs129, respectively as described before [17]. After one month the same volunteer took a dose of 50ml (1x109 CFU) after which stool samples were again analysed for 8 days.
Results and Discussion
Survival of both Ent. faecalis and E. coli was tested in an in vitro model resembling the gastric environment, mimicking conditions in the stomach, followed by conditions simulating the ileum. Two batches of each product were tested. The results obtained with Symbioflor 1 Ent. faecalis batch 1412 and 1383 are shown in Figure 2. Viability of the bacteria decreased between 1 and 2 logs in 30min of exposure to stomach acid conditions, so that few viable cells were detected after 180min. However, after 60 min the simulation of conditions applying in the ileum showed that enough bacteria had survived to slowly increase their numbers (Figure 2).
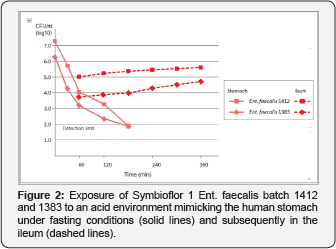
That Symbioflor 1 Ent. faecalis bacteria were able to survive passage through the GIT has thus far only been shown in the pig host [18]. In that study the bacteria were administered by a tube directly in the stomach, at a dose 10 times that of the daily recommended dose for humans, after body-weight correction. Using a number of in vitro tests, Christoffersen and colleagues compared Symbioflor 1 with four other Ent. faecalis strains: pathogenic V583 (a well-researched VanA-resistant clinical isolate), two commensal faecal isolates from babies (called strain 62 and 158B) and a cheese strain called LMGT3208 [19]. After 30min exposure to human gastric juice of pH 1.5, viability of all strains was severely impaired, and even when the pH was artificially raised to 5, exposure to the digestive enzymes present in gastric juice damaged all 5 strains. Of the tested strains, Symbioflor 1 was least capable to tolerate acid exposure, and grew poorly in BHI of pH 4.5 compared to the other strains. Exposure to human duodenal juice for 30 min also affected viability, but in this test Symbioflor 1 performed better (91% survival) than the other four strains [19]. These results suggest that Symbioflor 1 is not well equipped to survive in the human stomach, and this is confirmed by our results.
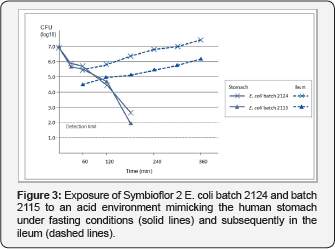
The results obtained with Symbioflor 2 E. coli batches 2124 and 2115 are presented in Figure 3. Compared to Ent. faecalis, the E. coli bacteria were better able to survive exposure to stomach pH, which is in accordance with a generally moderate acid tolerance for this species. The E. coli bacteria also reached higher levels in the ileum stimulation. The experiments resulting in Figures 2 & 3 were conducted simulating stomach conditions normally characterized by persistent low pH values. The SHIME in vitro model we next applied modelled physiological conditions that are more typical of a fed status. The applied stomach conditions lasted for 90min, followed by conditions simulating the small intestine for another 180min. The survival of the two different products is shown in Figure 4. Under the tested conditions, fewer bacteria were killed in the stomach compared to the previous model, while their numbers were relatively stable in the conditions simulating the upper GIT
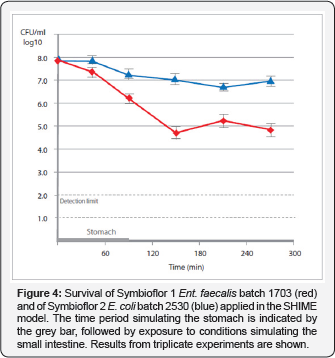
We consider the outcome of the SHIME model realistic, as oral uptake by a human volunteer indeed resulted in viable Symbioflor 1 Ent. faecalis in the stools (see below). A human volunteer study had already demonstrated that a single dose of Symbioflor 2 E. coli was sufficient to result in long-term colonisation, as demonstrated by strain-specific detection of the bacteria in the stools of six volunteers [20], with a single dose 2 to 20 times that of the daily recommended dose. In contrast, E. coli Nissle 1917 colonises the adult gut relatively poorly [21]. When seven adult volunteers took that probiotic for one week, the E. coli strain could only be detected in the stools of 4 individuals [22]. This poor colonisation is unlikely due to the killing activity of stomach acid, as the capsules sufficiently protect the bacteria against this (M. Marzorati, unpublished data). Instead, E. coli Nissle 1917 may just not be fit enough to outcompete residual microbiota in the adult gut. The latter is suggested by the fact that newborns are easily colonized [23]. Thus, there are marked differences between the two commercially available E. coli probiotic strains.
We next tested the ability of Symbioflor 1 Ent. faecalis to survive passage of the human GIT in vivo and colonise the gut of a single volunteer, who took 10 ml on a full stomach (10 times the recommended dose), corresponding to 2x108 CFU Ent. faecalis. Stool samples were collected on a daily basis for 8 days. PCR analysis of the stool samples using strain-specific primers targeting Symbioflor 1 Ent. faecalis revealed that bacteria could only be detected at day 1 and 2, after which stools became and remained negative (data not shown). The detected bacteria during the first two days most probably reflected transiently passed cells only.
A month later, the same person took a 5 times higher dose (50 times the daily recommended dose), after which the bacteria could be detected for 4 days using PCR directly applied to DNA extracted from stool. At the same time, targeted cultivation of enterococci from these stool samples in combination with colony 8 identification by PCR was performed. By this method, Symbioflor 1 Ent. faecalis was detectable for 6 days, with a decrease from day 4 onwards, as shown in Figure 5. Note that the stool sample on day 0 was taken 5hrs after intake of the product. If only 5% of the swallowed bacteria survived passage through the stomach, these must have multiplied to at least their original numbers in the intestine, as indicated by the levels that were detectable in stools. The presence of viable bacteria in stool was corroborated by FISH- detection of live Enterococcus spp. and Ent. faecalis in the faeces at day 1 after intake of 50ml (data not shown).
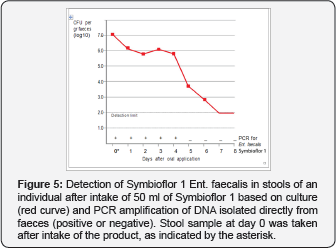
Compared to Symbioflor 2 E. coli, both survival in the stomach (as shown by the in vitro data) and colonisation in the human gut are weaker for Symbioflor 1 Ent. faecalis. The results show that, as the in vitro model predicted, sufficient bacteria survive the GIT passage to be excreted alive, while Ent. Faecalis bacteria do not persist longer than a few days, in contrast to Symbioflor 2 E. coli.
Conclusion
In conclusion, both probiotic Symbioflor products tested here are able to survive passage through the human GIT but, in contrast to E. coli, Ent. faecalis is only able to colonise the gut transiently.
conflict of interest
Trudy Wassenaar has worked as a consultant for Symbio Pharm GmbH. Kurt Zimmermann is a member of staff of SymbioPharm GmbH (producer of Symbioflor products). The other authors have no conflict of interest to declare.
References
- Food and Agriculture Organisation WHO (2001) Evaluation of health and nutritional properties of probiotics in food, including powder milk with live lactic acid bacteria. FAO & WHO Expert Consultation Report, Geneva, p.1-56.
- Caselli M, Cassol F, Calò G, Holton J, Zuliani G, et al. (2013) Actual concept of probiotics: is it more functional to science or business? World J Gastroenterol 19(10): 1527-1540.
- Giese T, Zimmermann K, Meuer SC (2003) Functional characterization of pro-biotic pharmaceuticals by quantitative analysis of gene expression. Arzneimittelforschung 53(5): 385-391.
- Habermann W, Zimmermann K, Skarabis H, Kunze R, Rusch V (2001) Einfluß eines bakteriellen Immunstimulans (humane Enterococcus faecalis Bakterien) auf die Rezidivhäufigkeit bei Patienten mit chronischer Bronchitis. Arzneimittelforschung 51(11): 931-937.
- Habermann W, Zimmermann K, Skarabis H, Kunze R, Rusch V (2002) Vrrminderung der Rezidivhäufigkeit bei Patienten mit chronisch rezidivierender hypertrophischer Sinusitis unter Behandlung mit einem bakteriellen Immunstimulans (Enterococcus faecalis Bakterien of humaner Herkunft). Arzneimittelforschung 52(8): 622-627.
- Vouloumanou EK, Makris GC, Karageorgopoulos DE, Falagas ME (2009) Probiotics for the prevention of respiratory tract infections: a systematic review. Int J Antimicrob Agents 34(3): 197. e1-10.
- Enck P, Zimmermann K, Menke G, Klosterhalfen S (2009) Randomized controlled treatment trial of irritable bowel syndrome with a probiotic E.-coli preparation (DSM17252) compared to placebo. Z Gastroenterol 47(2): 209-214.
- Martens U, Enck P, Zieseniss E (2010) Probiotic treatment of irritable bowel syndrome in children. Ger Med Sci 8: Doc07.
- Jansen G, Deddens B, Wilkinson M, van der Waaij D (1993) Significant decrease of titres of circulating IgG after oral intake of a preparation of Enterococcus faecalis in a group of ten healthy volunteers. Infection 21(3): 193-194.
- Jansen GJ, Wildeboer-Veloo AC, van der Waaij D, Degener JE (1998) Escherichia coli as a probiotic? Infection 26(4): 232-233.
- Minekus M, Marteau P, Havenaar R, Huis in het Veld JHJ (1995) A multicompartmental dynamic computer-controlled model simulating the stomach and small intestine. ATLA 23: 197-209.
- Marzorati M, Possemiers S, Verhelst A, Cade D, Madit N et al. (2015) A novel hypromellose capsule, with acid resistance properties, permits the targeted delivery of acid-sensitive products to the intestine. LWT- Food Sci Technol 60(1): 544-551.
- Marzorati M, Verhelst A, Luta G, Sinnott R, Verstraete W, et al. (2010) In vitro modulation of the human gastrointestinal microbial community by plant-derived polysaccharide-rich dietary supplements. Int J Food Microbiol 139(3): 168-176.
- Dressman JB, Amidon GL, Reppas C, Shah VP (1998) Dissolution testing as a prognostic tool for oral drug absorption: immediate release dosage forms. Pharm Res 15(1): 11-22.
- Malagelada JR, Robertson JS, Brown ML, Remington M, Duenes JA, et al. (1984) Intestinal transit of solid and liquid components of a meal in health. Gastroenterol 87(6): 1255-1263.
- Rossmann FS, Racek T, Wobser D, Puchalka J, Rabener EM, et al. (2015) Phage-mediated dispersal of biofilm and distribution of bacterial virulence genes is induced by quorum sensing. PLoS Pathog 11(2): e1004653.
- Meier H, Koob C, Ludwig W, Amann R, Frahm E, et al. (1997) Detection of enterococci with rRNA targeted DNA probes and their use for hygienic drinking water control. Water Sci Technol 35(11-12): 437444.
- Ottendorfer D, Zimmermann K, Taborski B, Thoma W, Thoma H, et al. (1995) Immunologische Ex-vivo-Untersuchungen zum Wirkmechanismus von humanen Enterococcus faecalis-Bakterien (Symbioflor 1®) an Miniaturschweinen. Forsch Komplemetärmed 2(6): 302-309.
- Christoffersen TE, Jensen H, Kleiveland CR, D0rum G, Jacobsen M, et al. (2012) In vitro comparison of commensal, probiotic and pathogenic strains of Enterococcus faecalis. Br J Nutr 108(11): 2043-2053.
- Wassenaar TM. Beimfohr C, Geske T, Zimmermann K (2014) Voluntarily exposure to a single, high dose of probiotic Escherichia coli results in prolonged colonisation. Benef Microbes 5(4): 367-375.
- Schütz E (1989) Behandlung von Darmerkrankungen mit Mutaflor. Eine multizentrische retrospektive Erhebung. Fortschr Med 107(3): 599-602.
- Prilassnig M, Wenisch C, Daxboeck F, Feierl G (2007) Are probiotics detectable in human feces after oral uptake by healthy volunteers? Wien Klin Wochenschr 119(15-16): 456-462.
- Lodinova-Zadnikova R, Sonnenborn U (1997) Effect of preventive administration of a nonpathogenic Escherichia coli strain on the colonization of the intestine with microbial pathogens in newborn infants. Biol Neonate 71(4): 224-232.






























