Volume 1 Issue 4 - November 2016, Article ID, AIBM.MS.ID.555569, 5 pages
http://dx.doi.org/10.19080/aibm.2016.01.555570Evaluation of Resistance Determinants and Clonal Relationship of Extended-Spectrum Beta-Lactamase Positive Gram-Negative Bacterial Strains by PCR and Raman Spectrophotometry
Gülnur TARHAN1*, Tülin DEMİR2, Uğur TAMER3, Merve ERYILMAZ3 and Mümtaz DADALI2
1Department of Medical Microbiology, Adiyaman University, Turkeys
2Ahi Evran University Training and Research Hospital, Turkey
3Department of Analiytical Chemistry, Gazi University, Turkey
Submission:September 22, 2016; Published: November 30, 2016
*Corresponding author: Gülnur Tarhan, Department of Medical Microbiology, Adiyaman University, Faculty of Medicine, Siteler Mahallesi, Atatürk Bulvar, No: 411 Adiyaman, Turkey, Tel: 90505 9448568; Emailgulnur.tarhan@yahoo.com
How to cite this article: Gülnur T, Tülin D, Uğur T, Merve E, et al.Evaluation of Resistance Determinants and Clonal Relationship of Extended-Spectrum Beta-Lactamase Positive Gram-Negative Bacterial Strains by PCR and Raman Spectrophotometry. Adv Biotech & Micro. 2016; 1(4): 555570. DOI: 10.19080/AIBM.2016.01.555570
Abstract
Background: ESBLs are enzymes which are capable of hydrolysing penicillins, broad-spectrum cephalosporins and monobactams. They are generally derived from TEM and SHV-type enzymes.
Materials and methods: We investigated the presence of TEM, SHV, CTX-M, OXA and IBC beta-lactamase genes by PCR in 300 E. coli, 130 Klebsiella spp., 100 P. aeruginosa ve 23 Enterobacter spp. isolated from various clinical specimens recovered from inpatient and outpatients. Additionally, epidemiological relationships were evaluated by Raman spectromicroscopy method.
Results: According to the PCR results; blaTEM (49.3%) and blaCTX-M (52.6%) were found at high rates, but blaIBC was not detected in any isolates. When PCR test results were evaluated according to the origin of bacteria, the highest rates of blaTEM (69.3%) ve blaCTX-M (79.3%) were detected in E. coli isolates, while the lowest rate (1%) was found in P. aeruginosa. With raman spectromicroscopy, it was identified 9 cluster in E. coli, 6 cluster in K. pneumoniae and K. oxytoca, 11 cluster in P. aeruginosa and 3 cluster in E. Cloacae
Conclusion: Although we obtained valuable results in the comparison of the raman spectrums in positive for resistance genes, there is a great necessity for standardisation of the studies prior to the usage of the test routinely.
Keywords: ESBL; PCR; Raman spectroscopy
Introduction
Extended-spectrum beta-lactamases (ESBL) were first described soon after the use of wide spectrum cephalosporins in the early 1980s. These enzymes are prevalent globally with a wide scale of variation between countries, hospitals wards and patient groups [1-8]. These are mainly described on plasmids but can be either plasmid or chromosomally-mediated causing resistance to ampicillin, carbenicillin, ticarcillin, cefalotin and cefamandole but have no effect on monobactam, cefamicin and susceptible to beta-lactamases inhibitors, cefoxitin and cefotetan [2,8-10]. ESBLs are derived from TEM-1, TEM-2 or SHV-1 gene loci by mutations [8] and over 400 types of different ESBLs have been identified with the common plasmid-encoded types including TEM, SHV, CTX-M and OXA [11].
TEM-beta-lactamases are the most common type of enzyme among Enterobacteriaceae and were also detected in P. aeruginosa [12-14]. While TEM-1 and TEM-2 could hydrolyse penicillin and first generation cephalosporins. SHV-1 beta-lactamases, generally encoded chromosomally in most strains, were first detected among K. pneumoniae and spread to other Enterobacteriacea species [13,15]. SHV-1 is resistant to ampicillin, ticarcillin and piperacillin but has no effect on oximinocephalosporins. CTX-M beta lactamase originates from chromosomal AmpC enzymes of Klyuvera ascorbata due to horizontal gene transfer and mutations and it shows 40% similarity with TEM and SHV enzymes. These are more active against cefotaxime and ceftriaxone than ceftazidime, even though point mutations can increase the activity towards ceftazidime [16]. Recently, it is reported that CTX-M15 is the most common enzyme globally. While most ESBLs are detected among Enterobacteriaceae, OXA types are commonly found in P. aeruginosa [13,14].
These are commonly spread with plasmid and transposon yielding resistance to aminopenicillin and ureidopenicillin. They have the ability to hydrolize oxacillin, cloxacillin and methicillin [13,14]. Inhibitor-resistant beta-lactamases (IRT), a variant due to mutations of SHV and TEM, has no ESBL activity and can not hydrolyze third generation cephalosporins and they are resistant to SAM and AMC but susceptible to TPZ [13]. IRT was commonly found among E. coli but also reported among other Enterobacteriaceae [14,15]. IBC-1, integron related Class A wide spectrum beta-lactamases, is highly resistant to ceftazidime, intermediate susceptible to cefotaxime, cefepime, aztreonam and less susceptible to clavulanic acid and piperacillin-tazobactam compared to other wide spectrum beta-lactamases [15,17]. It was first detected in E. cloacae and then in E. coli [11,18,19]. Recently, IBC-2, a variant of IBC-1 has been detected among
P. aeruginosa
strains. Additionally, non-TEM and non-SHV ESBLs such as PER, VEB, GES, TLA were reported [13].New approaches of bacterial identification have been considered recently for the rapid and accurate identification of bacteria. Vibrational spectroscopic techniques, infrared (IR) and Raman spectroscopy (RS), are commonly used in chemistry, since vibrational information is specific to the chemical bonds and symmetry of molecules. The mechanism of these systems is based on an intense beam of laser in the visible or infrared or ultraviolet region focused on the sample and detecting the scattered beam to get information about the vibration modes of the sample molecules. RS is a powerful molecular fingerprinting technique by which the molecule and bacteria can be identified through the interaction of coherent light and the sample’s molecules. It has recently gained popularity as an attractive approach for the biochemical characterization, rapid identification, and accurate classification of a wide range of bacterial species and strains [20-24]. This method is in clinical use in some advanced microbiology laboratories in recent years with the advantages of the ease for sample preparation, faster test results, reproducibility and higher discrimination power compared to other phenotypic and genotypic methods.
Studies have demonstrated that Raman spectra generated from bacterial and fungal colonies give sufficient information to identify and differentiate microorganisms and also for biofilm detection [18,20-22]. Raman signals obtained from bacterial samples suffer from weakness and a huge background. Surface enhanced Raman spectroscopy (SERS) is the most common and widely used way to amplify the weak Raman signal is to attach the sample to a metallic rough surface [18,20,23].
In this study, we aimed to determine the frequencies of beta-lactamase genes, TEM, SHV, CTX-M, OXA ve IBC by PCR and epidemiological clonal relationship by Raman spectromicroscopy among ESBL-positive E. coli, K. pneumoniae, Enterobacter spp. and P. aeruginosa recovered from various clinical specimens.
Materials and Methods
Bacterial strains
Between January 2009 and December 2012, a total of 553 non-duplicated ESBL-positive strain [300 E. coli, 130 Klebsiella spp. (88 K. pneumoniae, 42 K. oxytoca), 100 P. aeruginosa ve 23 Enterobacter spp. (20 E. cloacae, three E. aerogenes) recovered from clinical specimens of the patients admitted to Ahi Evran University Research and Training Hospital, Kirsehir, Turkey were included in the study. Identification to species level was carried out using the VITEK-2 Compact automated system (bioMérieux, France) and conventional biochemical tests.
Antimicrobial susceptibility testing and ESBL screening
Testing of susceptibility to ampicillin (AMP, 10 μg), amikacin (AMK, 30 μg), amoxicillin–clavulanic acid (AMC, 20/10 μg), aztreonam (ATM, 30 μg), cefepime (FEP, 30 μg), cefotaxime (CTX, 30 μg), ceftazidime (CAZ, 30 μg), ceftriaxone (CRO, 30 μg), cefuroxime (CXM, 30 μg), ciprofloxacin (CIP, 5 μg), cotrimoxazole (SXT, 1.25/23.75 μg), fosfomycin tromethamine (FOF, 200 μg), gentamicin (GEN, 10 μg), imipenem (IPM, 10 μg), and piperacillin–tazobactam (TZP, 100/10 μg) (Oxoid Ltd, Basingstoke, UK) was determined by Kirby–Bauer disk diffusion test method in accordance with Clinical and Laboratory Standards Institute (CLSI) guidelines [24] and the VITEK-2 Compact system. E. coli ATCC 25922 and Pseudomonas aeruginosa ATCC 27853 were used as quality control strains. ESBL screening was performed by disk synergy test, and results were confirmed by cefotaxime, ceftazidime, cefotaxime–clavulanic acid (CTC, 30/10 μg), and ceftazidime–clavulanic acid (CZC, 30/10 μg) disks, in accordance with CLSI guidelines [24]. The minimum inhibitory concentration (MIC) for imipenem was determined by gradient strip method (bioMérieux, France) following the manufacturer’s instructions, for strains resistant or intermediately resistant to imipenem by disk diffusion test. Additionally, the MBL gradient strip (bioMérieux, France) was used to determine MBL production for the strains resistant or intermediately resistant to imipenem.
Identification of resistance genes by PCR
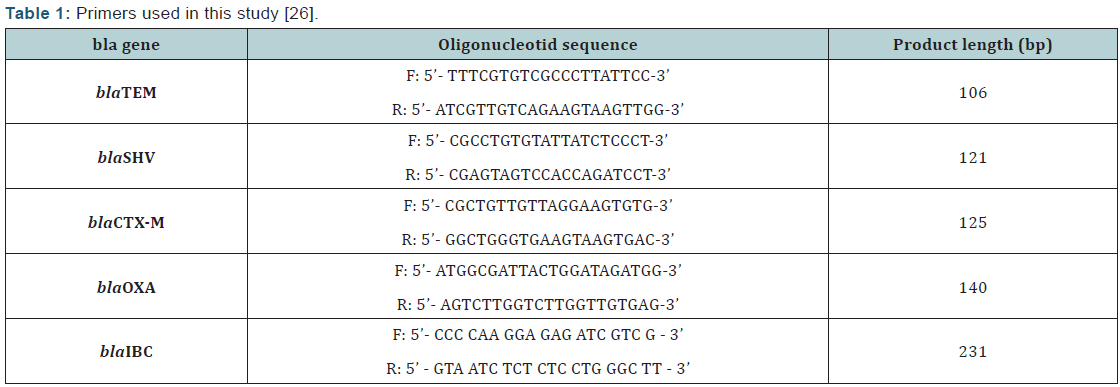
DNA templates for polymerase chain reaction (PCR) were obtained from isolates on nutrient agar. DNA extraction was performed by phenol:chloroform:isoamyl method [25]. The ESBL genes (blaIBC, blaOXA, blaCTX-M, blaTEM and blaSHV) were identified by PCR as previously described [26]. Primers used in this study were shown on Table 1. E. coli ATCC 25922, E. cloacae HT9 [blaIBC-1(+) control] (Strain was kindly provided by Eva Tzelepi, from Hellenic Pasteur Institute, Bacteriology Department), blaTEM-8, blaTEM-1 and blaSHV-3 and blaCTX-M-15 were used in each run.
Raman Spectromicroscopy
Preparation of bacterial cells: All strains were cultured in a nutrient broth at 37°C overnight, then 1 μL of the suspension was streaked on nutrient agar and incubated at 37 °C for 24 hours. Single colonies were harvested from the plates using an inoculating loop and suspended in 1.5 ml of deionized water. These aliquots were centrifuged at 5000 g for 3 min. The supernatant was discarded and pellet was used for the test. 10 μL of each of the suspensions was transferred to a lowfluorescence quartz microscope slide and allowed to dry at room temperature.
Preparation of the solutions: Delta Nu Examiner Raman Microscopy System (Deltanu Inc., Laramie, WY) was used in the analysis of bacterial strains. Parameters were set on the programme of 20x objective, 30 μm laser spot area, 220 mW laser power and 60 seconds of data collection time. Spectrums were obtained between 200-2000 cm-1 range. CTAB-stable golden nanorods were used in this study. Gold nanorods were synthesized with the minor revisions in the technique of the nucleus magnification. Nucleus solution was prepared with 7.5 ml CTAB and 250 μl AuCl4. Afterwards, 600 μl of NaBH4 solution that was previously prepared in the ice bath was added to the mixture quickly and left in room temperature for 30 minutes prior to use. Magnification solution was prepared using 4.75 ml CTAB, 500 μl AuCl4 and 60 μl AgNO3 and 100 μl ascorbic acid. 10 μL of nucleus solution was added to magnification mixture and left in room temperature for an hour prior to use.
Analysis and SERS measurement
Aluminium surface of the TLC paper was used for SERS measurement surface. SERS spectrum for each bacterial strain was obtained by dropping 2 μl of bacterial suspension onto TLC paper and interacting to the paper with 1 μl of gold nanorod solution. Median values of the signals from at least three different analysis area were evauated. Data analysis were performed by MATLAB version 7.1 (Mathworks, Natick, MA).
Statistical analyses
Cluster analysis of the spectral sets were performed by SpectraCell RA software (River Diagnostics). MATLAB version 7.1 (The MathWorks, USA) programme was used for histogram and correlation matrix. The similarities between spectrums were calculated using Pearson correlation coefficient analysis (R2) and was multiplied by 100 to express percentage. Data were analyzed using SPSS software 15.0 (SPSS, Inc., Chicago, IL, USA). Comparisons of categorical variables were done using Chisquare tests, although Fisher’s exact test was used when data were sparse. Significance was set at p < 0.05 using two-sided comparisons.
Results
A total of 553 ESBL-producing Gram-negative clinical bacterial isolates (300 E. coli, 130Klebsiella spp. (88 K. pneumoniae, 42 K. oxytoca), 100 P. aeruginosa ve 23 Enterobacter spp. (20 E. cloacae, 3 E. aerogenes) recovered from various infection sites were included in the study. Of all strains tested 402 were recovered from urine, 71 from tracheal aspirate, 48 from skin and soft-tissue infection. Overall the isolates tested; rates of strains carrying only SHV, TEM, CTX-M and OXA gene loci was 19 (3.4%), 63 (11.4%), 97 (17.5%) and 8 (1.4%), respectively. Out of 533 ESBL-producing strains 154 (27.8%) harbored none of the gene loci. blaCTX-M (n=291; 52.6%) was the most common enzyme type followed by blaTEM (n=274; 49.5%) and blaSHV (n=70; 12.6%) among all isolates tested.
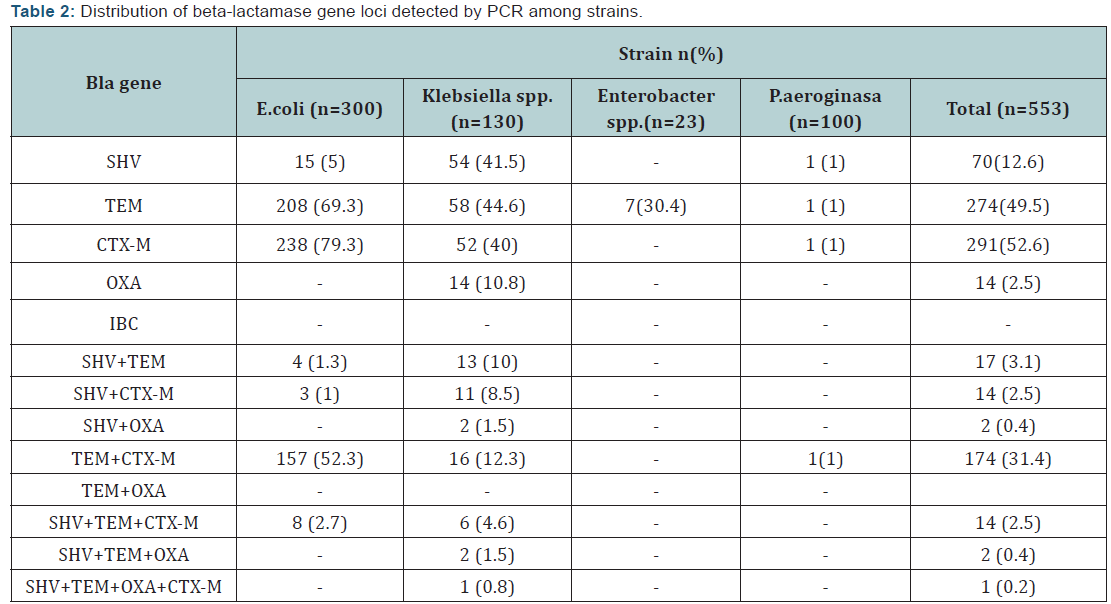
While blaCTX-M (79.3%) followed by blaTEM (69.3%) gene loci were frequent among E. coliM isolates, blaTEM (44.6%) and blaSHV (41.5%) gene loci were the frequent among Klebsiella spp. In this study, blaIBC was not detected in any of the strains. The most co-existence of the gene loci was blaTEM and blaCTX-M with 31.4% of the strains. blaSHV, blaTEM, blaCTX-M gene loci was found in 1% of the P. aeruginosa strains but blaOXA gene was not detected. blaTEM positivity was 30.4% for Enterobacter spp. and blaSHV, blaCTX-M and blaOXA gene loci was not detected. Overall the E. coli isolates tested (n=300), 157 was positive for blaTEM+blaCTX-M gene and eight was positive for SHV+TEM+CTX-M. In a K. pneumonae isolate recovered from aspirate (0.8%), all four gene loci except IBC were detected. Presence and distribution of blaSHV, blaTEM, blaCTX-M, blaOXA and blaIBC gene was shown on Table II and III.
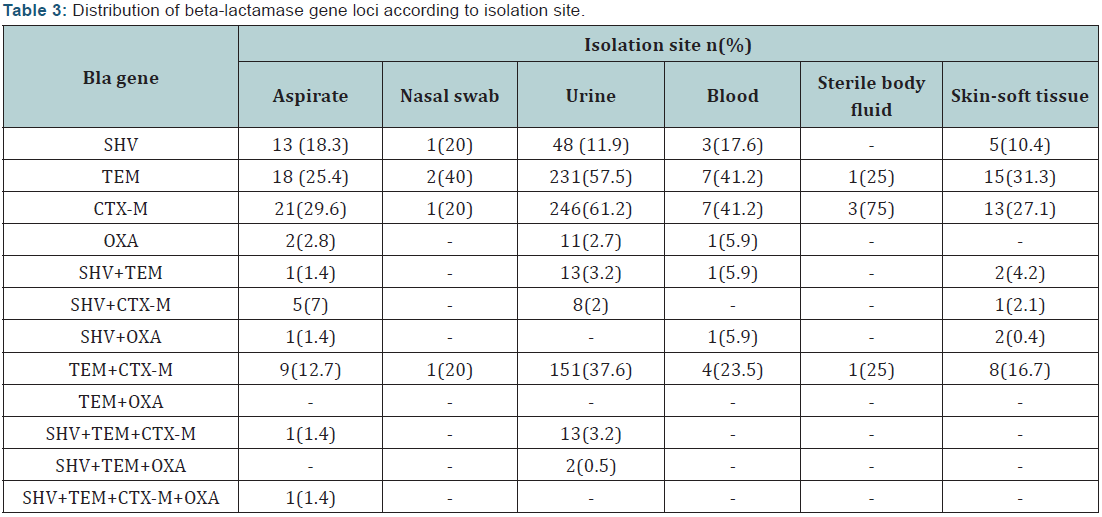
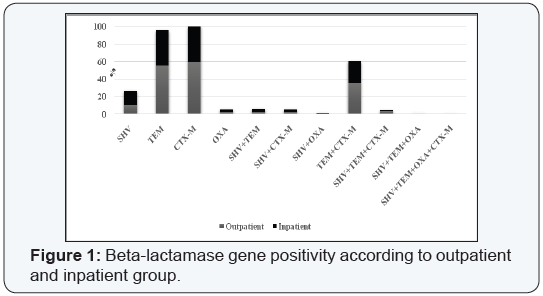
While among urinary isolates the common gene loci was CTX-M (61.2%) followed by TEM (57.5%); 41.2% of the bloodstream isolates were positive for TEM/CTX-M. Any gene loci was not detected among throat and ear samples. Evaluation of the gene loci positivity according to the patient admission, SHV, followed by CTX-M positivity were the most common gene loci among both inpatient and outpatient group (Figure 1, Image 1).
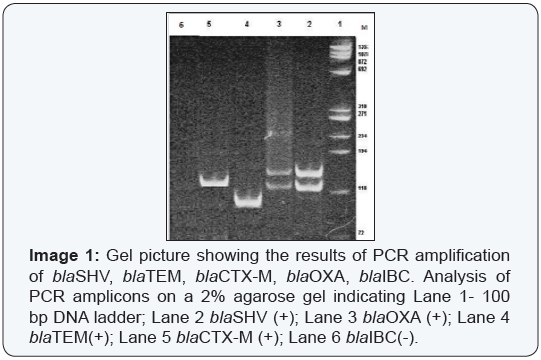
In the second part of the study, subtype analysis of the strains positive for any of beta-lactamase gene loci by PCR were evaluated using Raman spectromicroscopy with five different concentration. Five μL of bacterial suspension was added to 1X nanoparticule suspansion prepared as 100 μL, 125 μL, 150 μL, 175 μL, 200 μL. Raman spectromicroscopy did not show marked difference in the spectrum among the prepared solutions in some strains. As shown in Figure 2, it is observed that the best bacterial concentration was 5 μL bacteria added to 100 μL of nanoparticule suspension. Same results were achieved in the following four repeated tests performed with same samples and same test conditions. Concentration of 5 μL bacterial suspension to 100 μL of nanoparticule suspension was used for the evaluation of the strains positive for any of blaSHV, blaTEM, bla CTX-M and bla OXA gene loci.
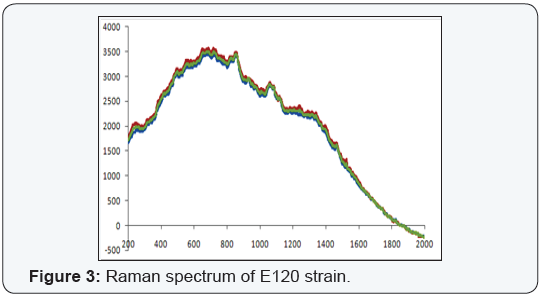
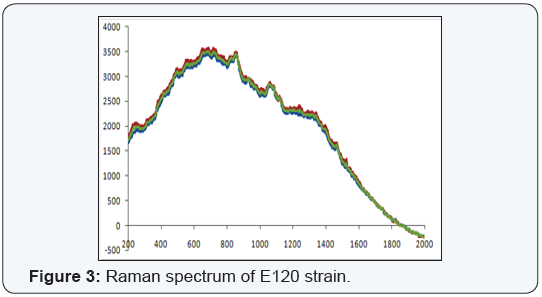
The raman spectrums of the strains with lack of gene loci, E. coli E120, was shown on Figure 2 and positive for three (blaSHV, blaTEM and blaCTX-M) of the five gene loci, E. coli E113, E. coli E66 ve K. pneumoniae K9 strains were shown on Figure 3-5. Weak signal was detected in E120 strain that any of the gene loci was not detected. A strong and discriminative spectrum was detected among E. coli E113, E. coli E66 and K. pneumoniae K9 strains that were positive for blaSHV, blaTEM and blaCTX-M by PCR. All bacterial strains showed similar results in this evaluation (Figure 3-6).
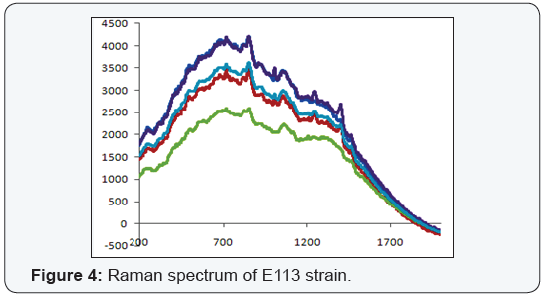
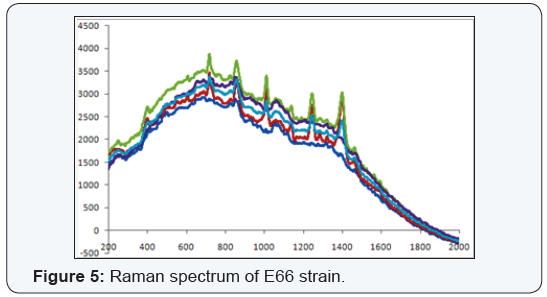
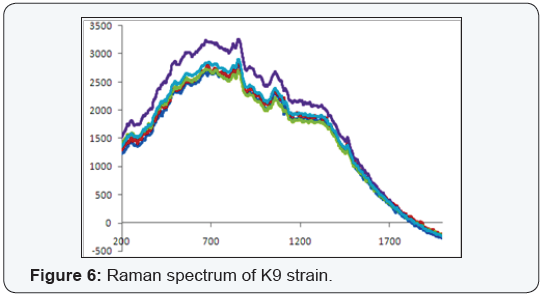
Bacterial typing with Raman spectrum revealed that nine cluster among E. coli strains, six cluster among Klebsiella spp, 11 cluster among P. aeruginosa and three cluster among E. cloacae were detected. Cluster analysis was not performed for E. aerogenes strains due to the low number of the strains.
Discussion
Extended-spectrum beta-lactamase producing and multidrug-resistant Gram-negative bacteria related infections are one of the major growing concerns worldwide. Thus, rapid and accurate detection of resistance mechanism and determinants has critical importance in the control of the infections and determining the treatment options [2].
In this study, a total of 553 ESBL-positive Gram-negative bacteria recovered from various infection sites were evaluated for beta-lactamase gene presence including bla IBC, bla TEM, bla SHV, bla CTX-M by PCR and raman spectroscopy. Among all isolates tested blaTEM 49.3% and blaCTX-M 52.6% gene loci was detected but bla IBC presence was not detected. Evaluation of the gene loci presence by bacterial strains revealed that bla TEM (69.3%) and bla CTX-M (79.3%) genes were higher among E. coli and blaSHV (41.5%) and bla TEM (44.6%) gene was higher among Klebsiella strains. While 1% of P. aeruginosa strains were positive for blaSHV, bla TEM, bla CTX-M genes, bla OXA was not detected. The most co-existence of the gene loci was blaTEM and bla CTX-M with 31.4% of the strains. Of 300 E. coli strain tested 165 were positive for bla TEM and bla CTX-M.
In a study conducted by Paterson et al. [11] including 455 K. pneumoniae strains recovered from 12 hospitals from seven countries were evaluated for ESBL presence and among ESBLpositive strains 67.1% were positive for SHV, 16.4% for TEM, 23.3% for CTX-M. This is worrisome, especially in Turkey where ESBL prevalence is very high [6,25]. In the past decade blaCTX-M gene has replaced blaSHV and blaTEM genes in Canada, Europe and Asia as the most common ESBL type in these bacteria similar with our findings. The CTX-M beta-lactamases are now widespread in both nosocomial and community-acquired pathogens. The blaTEM gene has a high frequency compared to bla SHV gene which is similar to our finding.
Raman spectroscopy is a new method in the determination and typing of infectious microorganisms. Although it has been used for a long time for the chemical characterization of different materials, it has just lately been applied to the study of biological samples in order to provide a rapid identification and discrimination of pathogenic organisms [18, 20- 22]. Data concerning issue is scarce worldwide. In our study, epidemiological typing with Raman spectrums was performed for the strains found to be positive for any of the five gene loci and found that E. coli has nine cluster, K. pneumoniae and K. oxytoca has six cluster, P.aeruginasa has eleven cluster and E. cloacae has three clusters. Due to the low number of the strains E. aerogenes strains were not included for typing. Comparison of the raman spectrums of the strains showed that higher spectrums were detected among strains positive for resistance genes. There is a great necessity for standardisation of the studies prior to the usage of the test routinely. Although, this method gives valuable information, it doesn’t seem that this method could be alternative to PFGE, gold standart for epidemiologial typing and it should be reevaluated with a reference method. Further studies should be conducted for better understanding and standardised the method itself.
The use of some first line treatment antibiotics such as penicillin and trimethoprim/sulfamethoxazole seem inappropriate. Antibiotics resistance surveillance and the determination of molecular characteristics of ESBL isolates are primordial to ensure the judicious use of antimicrobial drugs. The prevalence of beta-lactamase producing isolates and their isolation from life-threatening infections, is increasing at an alarming rate worldwide. It was shown in this study that betalactamase producing E. coli strains are an emerging threat in hospitals and should be supervised by implementation of timely identification and strict isolation methods that will help reduce their severe outcomes and mortality rate in this patients. In conclusion, this study has confirmed the potential of Raman spectroscopy to identify bacteria.
References
- Bradford PA (2001) Extended spectrum beta-lactamases in the 21st century. Characterization, epidemiology and detection of this important rezistance threat. Clin Microbiol Rev 14(4): 933-951.
- Ramphal R, Ambrose PG (2006) Extended spectrum beta-lactamases and clinical outcomes: Current data. Clin Inf Dis 42(suppl 4): 164-167.
- Cornaglia G, Garau J, Livermore DM (2008) Living with ESBLs. Clin Microbiol Infect 1: 1-2.
- Bourjilat F, Bouchrif B, Dersi N, Gros Claude JD, Amarouch H, et al. (2011) Emergence of extended-spectrum beta-lactamases-producing Escherichia coli in community-acquired urinary infections in Casablanca, Morocco 5(1): 850-855.
- Hawser SP, Bouchillon SK, Hoban DJ, Badal RE, Hsueh PR, et al. (2009) Emergence of High Levels of Extended-Spectrum-β-Lactamase- Producing Gram-Negative Bacilli in the Asia-Pacific Region: Data from the Study for Monitoring Antimicrobial Resistance Trends (SMART) Program, Antimicrob Agents Chemother 53(8): 3280-3284.
- Bayraktar B, Toksoy B, Bulut E (2010) Detection of bla(CTX-M) betalactamase genes in extended-spectrum beta-lactamase producing gram-negative bacteria. Mikrobiyol Bul 44(2): 187-196.
- Paterson DL, Bonomo RA (2005) Extended-Spectrum B-Lactamases: a Clinical Update. Clin Microbiol Rev 18(4): 657-686.
- Paterson DL, Hujer KM, Hujer AM, Yeiser B, Bonomo MD, et al. (2003) International Klebsiella Study Group: Extended-Spectrum beta- Lactamases in Klebsiella pneumoniae Bloodstream Isolates from Seven Countries: Dominance and Widespread Prevalence of SHV-and CTX-M-Type beta-lactamases. Antimicrobial Agents and Chemotherapy 47(11): 3554-3560.
- Taslı H, Bahar IH (2005) Molecular Characterization of TEM and SHV-Derived Extended Spectrum Beta Lactamases in Hospital Based Enterobacteriaceae in Turkey. Jpn Journal of Infect Dis 58(3): 162-167.
- Stürenburg E, Mack D (2003) Extended Spectrum Beta Lactamases Implications for The Clinical Microbiology Laboratory, Therapy and Infection Control. J Infect 47(4): 273-295.
- Shah AA, Hasan F, Ahmed S, Hameed A (2004) Characteristics, Epidemiology and Clinical Importance of Emerging Strains of Gram Negative Basilli Producing Extended Spectrum Beta Lactamases. Res Microbiol 155(6): 409-421.
- Lansdell PM, Hamouda A, Amyes SG Boyes J, Hamouda A (2012) Molecular epidemiology and antimicrobial resistance pattern of extended-spectrum-β-lactamase-producing Enterobacteriaceae in Glasgow, Scotland. J Antimicrob Chemother 67(3): 573-577.
- Kartali G, Tzelepi E, Pournaras S, Kontopoulou C, Kontos F, et al. (2002) Outbreak of Infections Caused by Enterobacter cloacae Producing the Integron-Associated Beta Lactamase IBC-1 in a Neonatal Intensive Care Unit of a Greek Hospital. Antimicrob Agents Chemother 46(5): 1577- 1580.
- Vourli S, Tzouvelekis SL, Tzelepi E, Lebessi NJ, et al. (2002) Characterization of in In111 a Class 1 Integron That Carries the Extended Spectrum Beta Lactamase Gene bla IBC-1. FEMS Microbiology Lett 225(1): 149-153.
- Tzelepi E, Managa C, Platsouka E, Sofianou D, Paniara O, et al. (2003) Extended Spectrum Beta Lactamase Types in Klebsiella pneumoniae and E. coli in Two Greek Hospitals. Int J Antimicrob Agents 21(3): 285- 288.
- Taylor PD, Vinn O, Kudryavtsev A, Schopf JW (2010) Raman spectroscopic study of the mineral composition of cirratulid tubes (Annelida, Polychaeta). Journal of Structural Biology 171(3): 402-405.
- Choo-Smith LP, Edwards HGM, Endtz HP, Kros JM, Heule F, et al. (2002) Medical applications of Raman spectroscopy: from proof of principal to clinical implementation. Biopolymers 67(1): 1-9.
- Kneipp K, Haka AS, Kneipp H, Badizadegan K, Yoshizawa N, et al. (2002) Surface-enhanced Raman spectroscopy in single living cells using gold nanoparticles. Applied Spectroscopy 56(2): 150-154.
- Campion A, Kambhampati P (1998) Surface-enhanced Raman scattering. Chemical Society Reviews 27: 241-250.
- Almarashi JFM, Kapel N, Wilkinson TS,Telle HH (2012) Raman Spectroscopy of Bacterial Species and Strains Cultivated under Reproducible Conditions. Spectroscopy: An International Journal 27(5-6): 361-365.
- Kirschner C, Maquelin K, Pina P, Ngo Thi NA, Choo-Smith LP (2001) Classification and identification of enterococci: a comparative phenotypic, genotypic, and vibrational spectroscopic study. J Clin Microbiol 39(5): 1763-1770.
- Maquelin K, Choo-Smith LP, Endtz HP, Bruining HA, Puppels GJ Rapid identification of Candida species by confocal raman microspectroscopy. Journal of Clinical Microbiology 40(2): 594-600.
- Samek O, Almarashi JFM, Telle HH (2010) The potential of Raman spectroscopy for the identification of biofilm formation by Staphylococcus epidermidis. Laser Physics Letters 7(5): 378-383.
- Clinical and Laboratory Standards Institute (2012) Performance standards for antimicrobial susceptibility testing. Twenty second informational supplement update. CLSI document M100-S22 U Clinical and Laboratory Standards Institute, Wayne, PA, USA.
- Maniatis SF (1989) Molecular Cloning: A Laboratory Manual. 2nd Edition, Vol. 3, pages E3 - E4; Cold Spring Harbor Laboratory Press, AAP 3/31/991.89.
- Bali EB, Açık L, Sultan N (2010) Phenotypic and molecular characterization of SHV, TEM, CTX-M and extended-spectrum -lactamase produced by Escherichia coli, Acinobacter baumannii and Klebsiella isolates in a Turkish hospital. African Journal of Microbiology Research 4: 650-654.






























