Biological Active Rakicidins A, B and E produced by the Marine Micromonospora sp. Isolate Guam1582
Wiebke Landwehr1,2, Sabrina Karwehl2,3, Peter J Schupp4, Peter Schumann5 and Joachim Wink1,2*
1Department of Microbial Strain Collection, Helmholtz Centre for Infection Research, Germany
2German Centre for Infection Research (DZIF), Partner Site Hannover-Braunschweig, Germany
3Department of Microbial Drugs, Helmholtz Centre for Infection Research, Germany
4Department of Environmental Biochemistry, Institute for Chemistry and Biology of the Marine Environment, Germany
5Leibnitz-Institut DSMZ-Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH, Germany
Submission: July 28, 2016; Published: August 5, 2016
*Corresponding author: Joachim Wink, Department of Microbial Strain Collection, Helmholtz Centre for Infection Research, Inhoffenstraße 7, 38124, Braunschweig and German Centre for Infection Research (DZIF), Partner Site Hannover-Braunschweig, 38124, Braunschweig, Germany.
How to cite this article: Wiebke L, Sabrina K, Peter JS, Peter S, Joachim W. Biological Active Rakicidins A, B and E produced by the Marine Micromonospora sp. Isolate Guam1582. Adv Biotech & Micro. 2016; 1(2): 555558. DOI: 10.19080/AIBM.2016.01.555558
Abstract
The bioactive substances rakicidin A (1), B (2) and E (3), were isolated from Micromonospora sp. cultivated after insertion of a diffusion growth chamber into the sponge Rhabdastrella globostellata. The three rakicidins were tested in various antibiotic bioassays revealing their selective activity against Gram-positive bacteria including resistant strains of Staphylococcus aureus and Enterococcus faecium. With the help of MALDI-TOF MS and RiboPrinter® analysis the relationship between the producer strain and 13 Micromonospora type strains (99% identity in 16S rRNA sequence) could be clarified.
Keywords: Actinobacteria; Micromonospora; Rakicidin; Secondary metabolites; Bioactivity; MALDI-TOF, RiboPrinter®
Abbreviations: MRSA: Methicillin Resistant Staphylococcus aureus; VREF: Vancomycin Resistant Enterococcus faecium; MIC: Minimal Inhibition Concentration Test; TetR: Tetracycline Resistance; ESBL: Extended–Spectrum Beta-Lactamase; JSRM: Jump Start Ready Mix; MALDI-TOF MS: Matrix-Assisted Laser-Desorption-Ionization Time-of-Flight Mass Spectrometry
Introduction
Actinobacteria are one of the richest sources of secondary metabolites [1] and more than 50% of clinical useful antibiotics of microbial origin were produced by these microbes, especially by the soil inhabiting genera Streptomyces and Micromonospora.[2].
However, while the rate of newly discovered antibiotics from soil inhibiting actinobacteria decreased the rate of re-isolation increased [3,4]. To further explore this well studied source, new strains have to be isolated with alternative methods or from unexplored environments.[4] Oceans are the world’s biggest habitat. They cover 70% of the earth surface and have a microbial abundance of 106 bacteria per mL in sea water and 109 bacteria per mL in ocean bottom sediments.[5] Not only sea water and sediments are microbial rich environments but also marine sponges harbour diverse and abundant microbial communities, which comprise up to 35% of sponge-biomass.[6-8] These sponge associated bacteria already have been shown to be prolific producers of bioactive natural products.[9-11] Strain Guam1582, the producer of rakicidins A [12], B [12] and E [13] was isolated from the shallow water sponge Rhabdastrellaglobostellata in Guam using a recently developed diffusion growth chamber.
It was classified as Micromonospora sp. on the basis of 16S rRNA sequence. To date, rakicidins A (1) and B (2) have been reported to inhibit tumour cell lines [14-16] without showing any antibiotic activity. However, in our ongoing screening for antibiotics strain Guam1582 stood out with activities against methicillin resistant Staphylococcus aureus (MRSA) and vancomycin resistant Enterococcus faecium (VREF). These substances are described to be produced by different Micromonospora spp. [16,12] as well as Micromonospora chalcea strains (Rakicidin B) [14,17]. Because of the close relationship of the marine isolate Guam1582 to M. chalcea (DSM 43026T) and further Micromonospora type strains in the 16S rRNA sequence (99% identity), the differentiation from the other strains was done using MALDI-TOF and RiboPrinter analysis.
Material and Methods
Cultivation of strain Guam1582
The pre-culture of strain Guam1582 was grown in 100 mL of GYM medium (0.4% glucose, 0.4% yeast extract, 1% malt extract, 0.2% CaCO3; pH 7.2; sterilized for 20 min at 121°C) in a 250 mL flask for 5 days at 30 °C and 160 rpm on a rotary shaker. The production culture was transferred 1:10 in 50 250 mL flasks, filled with 100 mL of medium 5294 (1% soluble starch, 0.2% yeast extract, 1% glucose, 1% glycerol, 0.25% corn steep liquor, 0.2% peptone, 0.1% NaCl, 0.3% CaCO3: pH 7.2) laced with artificial sea water (“Coral Ocean”, ATI) and sterilized for 20 min at 121 °C. The cultures were grown for 7 days at 30 °C and 160 rpm on a rotary shaker.
Isolation and purification of rakicidins A, B and E
The cells were separated from the culture broth by centrifugation. The culture broth was extracted in portions of 1 L with ethyl acetate (three times 500 mL). The combined ethyl acetate extract was dried with Na2SO4 and evaporated in vacuo to give a crude extract, which was dissolved in 100 mL methanol supplemented with 3% H2O and 50 mL heptane. The methanol layer was extracted three times with 50 mL heptane and the methanol was evaporated in vacuo to give 129 mg crude extract.
The cells were kept in 1 L acetone for 48 h. The acetone extract was evaporated to an aqueous mixture (300 mL) and extracted with ethyl acetate (three times with 150 mL). The combined ethyl acetate extract was dried with Na2SO4. It was evaporated in vacuo to give 88 mg of crude extract, which was dissolved in 50 mL methanol supplemented with 3% H2O and 50 mL heptane. The methanol layer was extracted three times with 50 mL heptane and the methanol was evaporated in vacuo to give 28 mg crude extract.
The crude extract was separated by preparative RP-MPLC [column: Kronlab ODS-AQ C18, 480×30 mm, 15 μm; solvent A: H2O/methanol (8/2); solvent B: methanol; gradient system: holding 50% B for 5 min, increasing to 65% B in 5 s, holding at 65% B for 5 min, increasing to 100% B in 130 min and holding at 100% B for 30 min; flow rate 30 mL/min, UV detection at 270 nm] to give 4.3 mg of rakicidin A (1), 6.5 mg of rakicidin B (2) and 1.4 mg of rakicidin E (3) (Figure 1).
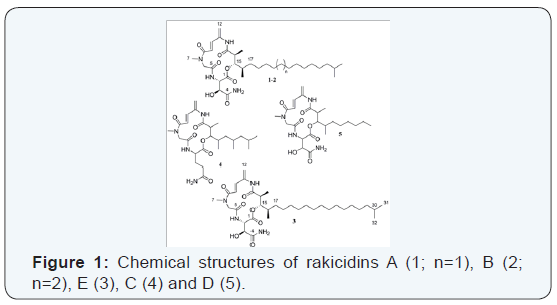
Antimicrobial testing
The biological activity was tested using a minimal inhibition concentration test (MIC). The assay was done in a 96 well plate. Rakicidins A, B and C were solved in DMSO in a 1 mg/ mL concentrated stock solution. This solution was mixed with the test strain (OD600: 0.01 (bacteria)/0.05 (yeast)/OD548: 0.1 (Schizosaccharomyces pombe; Rhodosporidium toruloides ) to a final concentration of 66.6 μg/mL in the first lane and was diluted in 1:2 steps with the inoculated test strain medium.
In addition to a panel consisting of Schizosaccharomyces pombe (DSM 70572), Pichia anomala (DSM 6766), Candida albicans (DSM 1665), Mucor hiemalis (DSM 2656), Rhodosporidium toruloides (DSM 10134), Micrococcus luteus (DSM 1790), Bacillus subtilis (DSM 10), Staphylococcus aureus Newman [18], Enterococcus faecium (DSM 20477), Mycobacterium sp. (DSM 43270), Escherichia coli (DSM 1116), Chromobacterium violaceum (DSM 30191), Pseudomonas aeruginosa (PA 14), the substances were also tested on some multi resistant bacteria: Escherichia coli WT-3 (#644; gyrA[S83L,D87G]) (quinolone resistant), Escherichia coli XL-1 blue (tetracycline resistance, Tn10 (TetR) on F plasmid), Escherichia coli ESBL (DSM 22664) (Extended–Spectrum Beta-Lactamase; produces CTX-M15 beta-lactamase), Staphylococcus aureus N315 (DSM 11822) (MRSA) and Enterococcus faecium (DSM 17050) (VREF (vanA); microaerophilic).
The strains were cultivated in Müller-Hinton Broth (Müller-Hinton Bouillon (Carl Roth, X927)), MYC medium (0.4 % phyto peptone, 1% glucose, 50 mM HEPES; pH; 7.0; sterilized at 121 °C for 20 min) (Candida albicans, Mucor hiemalis, Schizosaccharomyces pombe) or Tryptic Soy Broth (BactoTMTriptic Soy Broth, Soybean-Casein digest medium; Hach; sterilized at 121°C for 20 min) (Enterococcus faecium). After inoculation, the plates were incubated at 30/37 °C and 160 rpm for one day.
Cytotoxicity
To determine the cytotoxicity of rakicidins A, B and E, the murine fibroblast cell line L929 was used as target in a minimal inhibition concentration (MIC) assay in 96 well plates. The cells were cultured in tissue culture flasks in culture medium consisting of DULBCCO’s modified EAGLE’s medium (DMEM, Bio Whittaker, Walkersville, MD) supplemented with 10% fetal bovine serum (FBS, JRH Bioscience, Lenexa, KS). For the cytotoxicity assay the cells were removed from the flask using 0.25% of a trypsin solution (Gibco). After removing the trypsin, the cells were incubated for 5 min at 37 °C and under an atmosphere with 5% CO2.
The detached cells were resuspended in 10 ml of the culture medium DMEM + 10% FBS and the concentration of 50,000 cells/mL were adjusted. 87 μL of culture medium was filled in the first wells of the 96 well plate. After the addition of 3 μL of the compounds, the substances were diluted in 1:6 steps with the medium stock solution. Subsequently, 120 μL of the previously prepared L292 cell line stock solution (50,000 cells/ mL) was added to every well. The cells were incubated for 5 days at 37 °C under an atmosphere with 5% CO2. For the analysis the cells were examined under a light microscope.
Taxonomy
The DNA was isolated via Invisorb Spin Plant Mini Kit (stratec molecular, Germany). For preparation 1.5 mL of a well grown cell suspension was centrifuged for 2 min at 13000 rpm. The cell pellet was mixed with 100 μL of lysation buffer and incubated at 95 °C for 5 min. 300 μL lysation buffer and 20 μL proteinase K were added, mixed up and incubated for 30 min at 65 °C. The rest of the DNA isolation was done according to the manufacturer´s description.
For analysis of the 16S rRNA gene sequence, two primers were used matching most of the known eubacterial orders on the positions 27 (forward) and 1525 (reverse). For the conduction of the 16S PCR a special mastermix was created, containing water, primers and “Jump Start Ready Mix” (JSRM). The JSRM is a mixture including JumpStart Taq DNA polymerase, 99 % pure deoxynucleotides and buffer in an optimized reaction concentration. The PCR mixture had a final volume of 50 μL and consisted of the following components: 25 μL JSRM, 22 μL H2O, 1 μL forward and reverse primer dilution as well as 1 μL of the template DNA. The PCR within the Thermocycler started with an activation temperature of 95 °C for 5 min. Followed by 34 cycles of denaturation (94 °C, 0.5 min), a primer annealing at 52 °C for 0.5 min and 2 min of elongation at 72 °C. The final elongation had a temperature of 72 °C for 10 min. The samples were sequenced und compared to bacterial type strain sequences using the NCBI BLAST function [19].
Another marker for the phylogenetic description based on the ribosomes of the bacteria is the pattern of the ribosomal proteins as analysed by matrix-assisted laser-desorption/ ionization time-of-flight spectroscopy (MALDI-TOF MS). The sample preparation was done according to the ethanol/formic acid extraction protocol described by Schumann [20]: about 10 mg of biomass from a liquid culture (cultivated in complex medium DSMZ medium 65 at 30°C for 7 days) was suspended in 300 μl H2O and homogenised carefully. Afterwards, 900 μl ethanol was added to the suspension. The cells were collected after a centrifugation step and resuspended in 50 μl of 70% formic acid. After the addition of 50 μl of acetonitrile, the suspension was mixed and centrifuged.
The supernatant was removed immediately and aliquots of 1.5 μl were placed on each spot of a stainless-steel target plate. After air drying, 1.5 μl of matrix solution (saturated solution of α-cyano-hydroxy-cinnaminic acid in 50% aqueous acetonitrile containing 2.5% trifluoro acetic acid) per spot was added. MALDI-TOF mass spectra were recorded using a Microflex L20 mass spectrometer (Bruker Daltonics) with a N2 laser unit. The spectra were measured in linear positive mode and the acceleration voltage was 20 kV. In sum a spectra was collected out of 250 shots across a spot. To analyse the data, a mass range of 2000-20,000 m/z was observed. For internal calibration the Bacterial Test Standard #255343 (Bruker Daltonics) was used.
Using the Flexanalysis software (version 3.3, Bruker Daltonics), the MALDI-TOF MS spectra were smoothed, baseline corrected and re-calibrated. A score-orientated dendrogram was calculated by using the BioTyper software (version 3.1, Bruker Daltonics).
The RiboPrinter® analysis was carried out as described by Schumann & Pukall [20]. For the sample preparation, the strain Guam1582 and the type strain M. chalcea DSM 43026T were cultivated for 10 days on GYM medium agar plates (0.4% glucose, 0.4% yeast extract, 1% malt extract, 0.2% CaCO3, 1.5% agar; pH 7.2). A small amount of cells was processed by the fully automated procedure of the RiboPrinter® system (Du Pont).
Results
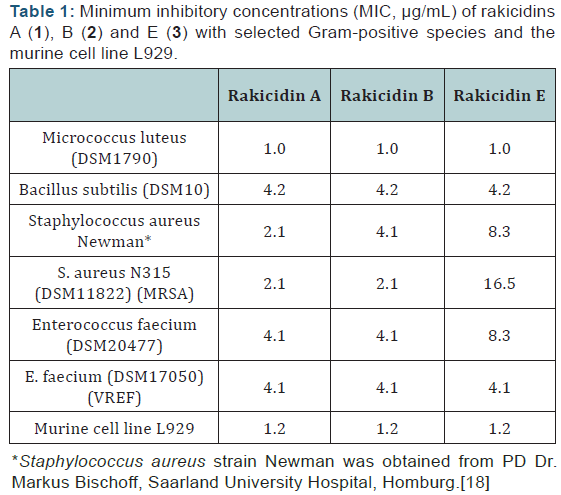
Rakicidins A (1), B (2) and E (3) (Figure 1) showed selective antibiotic activity against Gram-positive bacteria, including the methicillin resistant Staphylococcus aureus N315 (MRSA) and the vancomycin resistant Enterococcus faeciumDSM17050 (VREF) as well as inhibition of the murine fibroblast cell line L929. The biological activity against Micrococcus luteus (1.0 μg/mL), Bacillus subtilis (4.2 μg/mL) and the VREF strain (4.2 μg/mL) could be detected for all tested rakicidins at similar concentration. In contrast, against S. aureus rakicidin A (1) showed the lowest MIC value of 2.1 μg/mL and rakicidins B (2) and E (3) higher values of 4.1 μg/mL and 8.3 μg/mL, respectively. 1–3 had the same MIC values (4.1 μg/mL) against the MRSA strain but against the VREF strain rakicidin E (3) showed considerably decreased activity with an MIC of 16.5 μg/mL compared to 2.1 μg/mL for 1 and 2 (Table 1).
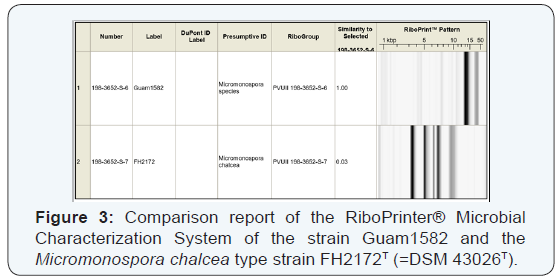
For the taxonomic description of strain Guam1582, 16S rRNA sequence analysis was performed. The sequence showed 99% similarity to 13 Micromonospora type strains (M. echinospora DSM 43816T, M. Citrea DSM 43903T, M. endolithica DSM 43813T, M. sagamiensis subsp. Flava DSM 44887T, M.endolithicaDSM 44398T, M. Rosaria DSM 803T, M.sagamiensis DSM 43912T, M. Echinofusca DSM 43913T, M.chalcea DSM 43026T, M. Inyonensis DSM 46123T, M.chersina DSM 44151T, M. pallida DSM 43817T, M.halophytica DSM 43171T. MALDI-TOF analysis suggested that the closest relative of strain Guam1582 was the type strain M. Chalcea DSM 43026T (Figure 2). This strain is also known to be the producer of rakicidin B (2). [17,14] However according to the different PvuII RiboPrinter patterns (Figure 3) strain Guam1582 and M. Chalcea are not identical at strains level. Studies on the metabolic profile of the M. Chalcea type strain DSM 43026T merely verified the production of rakicidin A (1) and B (2) but not ofrakicidin E (3).

Discussion
To date, rakicidins A (1) and B (2) have only been reported to inhibit tumour cell lines [14-16] In 1995 McBrien et al. [12] provided IC50 values for 1 and 2 against the murine cell line M109 (MADISON lung carcinoma 109) of 40 ng/mL and 200 ng/mL, respectively, but stated simultaneously that 1 and 2 did not show any antibacterial or antifungal activities. However, an antimicrobial activity of a raw extract has been described for 1in the study of Songsumanus et al. [17] against the Gram-positive bacterium Kocuria rhizophila. Of the other two rakicidins known in this family, rakicidin C (4, Figure 1) did not show any activity [21,22] against Gram-positive or -negative bacteria as well as against various cell lines whereas rakicidin D (5, Figure 1) inhibited the invasion of murine carcinoma colon 26-L5 cells with an IC50 of 3.3 μg/mL.[22] In contrast, we observed in liquid cultures that rakicidins A (1) and B (2) as well as the variant E (3) were selectively active against Gram-positive bacteria including VREF and MRSA strains.
With the help of the MADI-TOF MS and RiboPrinter® analysis we were able to define the close relationship between the Isolate Guam1582 (producer strain of rakicidin A, B and E) and the closest relatives detected by the analysis of the 16S rRNA sequence. It became clear that in most cases, especially in such huge Actinomycetes families like Micromonospora, it is necessary to use phylogenetic analysis methods of different taxonomic resolution [21] to distinguish between the strains which are very closely related in the 16S rRNA sequence. We could demonstrate that our isolate is on the one hand a very close relative of the type strain M. chalcea, which was described in literature to be the producer of rakicidin B, but with the help of the RiboPrinter® analysis we were able to show that they are not identical at the strains´ level.
Conclusion
In this study the antibiotic activity of the rakicidins A, B and E against Gram-positive bacteria could be shown. Especially the activity against the methicillin resistant Staphylococcus aureus N315 (MRSA) and the vancomycin resistant Enterococcus faecium DSM 17050 (VREF) were interesting new insights. However, the activity against the murine cell line was between 2 and 16 times stronger than against tested multi resistant bacteria and therefore, further studies will probably deal with the cytotoxic instead of the antibiotic activity of these substances.
With the help of different taxonomic approaches, we could show that the producer strain Guam1582 is very close related to 13 Micromonospora type strains (especially to M. chaleca), however with the help of MALDI-TOF and RiboPrinter® approaches, it could be demonstrated that it is an individual and probably new strain.
Acknowledgment
We thank W. Collisi, B. Trunkwalter and K. Schober for excellent technical assistance, C. Kakoschke for recording NMR spectra and A. Gollasch for recording the HRESIMS data. We also thank Susanna Whitfield for assistance with the initial isolation of producer strain Guam1582.
References
- Manivasagan P, Venkatesan J, Sivakumar K, Kim SK (2013) Marine actinobacterial metabolites: Current status and future perspectives. Microb Research 168(6): 311-332.
- Berdy J (2005) Bioactive microbial metabolites. J Antibiot (Tokyo) 58(1): 1-26.
- Fenical W, Baden D, Burg M, de Goyet CV, Grimes JD, et al. (1999) Marine-derived pharmaceuticals and related bioactive compounds. In:Fenical W, From Monosoons to Microbes: Understanding the Ocean´s Role in Human Health. National Academies Press, pp. 71-86.
- Lam KS (2006) Discovery of novel metabolites from marine actinomycetes. Curr Opin Microbiol 9(3): 245-251.
- Fenical W, Jensen PR (2006) Developing a new resource for drug discovery: marine actinomycete bacteria. Nat Chem Biol 2(12): 666- 673.
- Hentschel U, Piel J, Degnan SM, Taylor MW (2012) Genomic insights into the marine sponge microbiome. Nat Rev Microbiol 10(9): 641- 654.
- Webster NS, Taylor MW (2012) Marine sponges and their microbial symbionts: love and other relationships. Environ Microbiol 14(2): 335- 46.
- Steinert G, Whitfield S, Taylor MW, Thoms C, Schupp PJ (2014) Application of diffusion growth chambers for the cultivation of marine sponge-associated bacteria. Mar Biotechnol 16(5): 594-603.
- Blunt JW, Copp BR, Munro MHG, Northcote PT, Prinsep MR (2011) Marine natural products. Nat Prod Rep 28(2): 196-268.
- Blunt JW, Copp BR, Keyzers RA, Munro MHG, Prinsep MR (2012) Marine natural products. Nat Prod Rep 29(2): 144-222.
- Blunt JW, Copp BR, Keyzers RA, Munro MHG, Prinsep MR (2013) Marine natural products. Nat Prod Rep 30(2): 237-323.
- McBrien KD, Berry RL, Lowe SE, Neddermann KM, Bursuker I, et al. (1995) Rakicidins, new cytotoxic lipopeptides from Micromonospora sp. fermentation, isolation and characterization. J Antibiot 48(12): 1446-1452.
- Oku N, Matoba S, Yamazaki YM, Shimasaki R, Miyanaga S, et al. (2014) Complete stereochemistry and preliminary structure-activity relationship of rakicidin A, a hypoxia-selective cytotoxin from Micromonospora sp.J Nat Prod 77(11): 2561-2565.
- Xie JJ, Zhou F, Jiang H, Du ZP, Lin R, et al. (2011) FW523-3, a novel lipopeptide compound, induces apoptosis in cancer cells. Mol Med Rep 4(4): 759-763.
- Takeuchi M, Ashihara E, Yamazaki Y, Kimura S, Nakagawa Y (2011) Rakicidin A effectively induces apoptosis in hypoxia adapted Bcr‐Abl positive leukemic cells. Cancer Sci 102(3): 591-596.
- Yamazaki Y, Kunimoto S, Ikeda D (2007) Rakicidin A: a hypoxiaselective cytotoxin. Biol Pharm Bull 30(2): 261-265.
- Songsumanus A, Kudo T, Igarashi Y, Tanasupawat S (2013) Characterization and screening of antimicrobial activity of Micromonospora strains from Thai soils. Mal J Microbiol 9: 260-269.
- Duthie ES (1952) Variation in the antigenic composition of staphylococcal coagulase. J Gen Microbiol 7(3-4): 320-326.
- Schumann P, Maier T (2014) MALDI-TOF mass spectrometry applied to classification and identification of bacteria. Methods Microbiol 41: 275-306.
- Schumann P, Pukall R (2013) The discriminatory power of ribotyping as automatable technique for differentiation of bacteria. Syst Appl Microbiol 36(6): 369-375.
- Hu JF,Wunderlich D, Sattler I, Feng XZ, Grabley S, et al. (2000) Rakicidin C, A New Cyclic Depsipeptide from Streptomyces sp. Eur J Org 2000(19): 3353-3356.
- Igarashi Y, Shinasaki R, Miyanaga S, Oku N, Onaka, et al. (2010) Rakicidin D, an inhibitor of tumor cell invasion from marine-derived Streptomyces sp. W J Antibiot 63(9): 563-565.






























