Machine Learning based Sorting of Somatic Embryos for In-Line processing in Automated SE Fluidics System
Punnag Chatterjee1,2, Aruna Weerasekara2,5, Ulrika Egertsdotter3,4 and Cyrus Aidun2,3,5
1Indian Institute of Technology - Dharwad
2G.W. Woodruff School of Mechanical Engineering, Georgia Institute of Technology, Atlanta, GA, USA
3Renewable Bioproducts Institute, Georgia Institute of Technology, Atlanta, GA, USA
4Department of Forest Genetics and Plant Physiology, Umea˚ Plant Science Centre (UPSC), Swedish University of Agricultural Science (SLU), Umea˚, Sweden
5Bioautomaton Systems Inc., Atlanta, GA, USA
Submission: January 11, 2024; Published: January 22, 2024
*Corresponding author: Punnag Chatterjee, 2G.W. Woodruff School of Mechanical Engineering, Georgia Institute of Technology, Atlanta, GA, USA
How to cite this article: P Chatterjee, A Weerasekara, U Egertsdotter, C Aidun. Machine Learning based Sorting of Somatic Embryos for In-Line processing in Automated SE Fluidics System. Agri Res& Tech: Open Access J. 2024; 28(1): 556399. DOI: 10.19080/ARTOAJ.2023.27.556399
Abstract
Somatic embryogenesis (SE) can be a viable method for the clonal propagation of many economically significant forest trees, particularly coniferous trees like pines and spruces. However, large-scale production of SE plants requires automation to reduce manual labor and attain cost-efficiency. The most labor-intensive step of the SE process for SE plant production is selecting and harvesting mature embryos. Embryo maturation is not a synchronized process; selecting the most developed embryos capable of continuous development is necessary. However, there needs to be more research conducted on mapping morphological features to germination-competent mature somatic embryos. This paper lays down the preliminary work of employing machine learning techniques for classifying large volumes of images of mature somatic embryos processed using an automated SE processing system based on fluidics processing referred to as SE Fluidics system. The results show that machine learning could be an alternative classification methodology instead of the traditional manual morphology-based classification process based on image analysis. The paper discusses two popular image classification techniques, namely Convolution Neural Network (CNN) and Support Vector Machine (SVM), applying them to both binary (black and white) and grayscale images. It is observed that grayscale images provide better accuracy with the SVM technique and outperform morphology-based classification in terms of processing speed (17.6% faster) across the test envelope. On the other hand, CNN-based classification shows better processing speeds only at a lower number of convolution layers. Hence, the data scientist can optimally select the number of convolution layers to get the desired accuracy-processing speed combination.
Keywords: SE Fluidics system; Image analysis; Somatic Embryogenesis; Machine learning; Embryo morphology
Abbreviations: SE: Somatic Embryogenesis; CNN: Convolution Neural Network; SVM: Support Vector Machine; ML: Machine Learning; PEM: Pro Embryogenic Mass; BW: Black and White; SS: Stainless Steel; ReLU: Rectified Linear Unit; HOG: Histogram of Oriented Gradients
Introduction
As the world moves toward a carbon-neutral future, active forest management and planting of genetically improved material will play an essential part in the realization of this goal. Although deforestation contributes to net atmospheric emissions, the remaining forests are a net carbon sink. Between 2011-2015 forests stored, on average, some 2.1 GT of CO2 annually, of which half was estimated to be due to net growth in planted forests [1]. Furthermore, biomass from wood has the advantage of possessing higher energy density when compared with non-woody biomass [2] Globally, conifers have long dominated production forestry in the northern hemisphere and are increasingly planted in the southern hemisphere. Norway spruce (Picea abies (L.) Karst) is northern and central Europe's commercially most important species. It is also the most planted species in northern Europe [3]. The feasibility of automating Norway spruce's SE plant production process has been demonstrated for scale up to industrial production [4] A vital step of the automation process is to characterize morphological parameters and link them to germination competent, mature somatic embryos [5]. There have been many publications in the agricultural domain involving the use of machine learning (ML) algorithms; in tissue culture, data-driven approach has been adopted by numerous researchers to simulate and predict further growth and developmental processes under in-vitro conditions. A review article by Hesami et al. [6] documents the latest application of ML models used in plant cell and tissue culture.
In the agriculture industry, image processing was introduced almost a decade ago to aid the sorting and classification of seeds for crop production [7,8]. Researchers have recently started implementing machine learning (ML) algorithms in somatic embryogenesis. (Mohsen et al.) [9] captured images in-vitro to measure the physical properties of embryogenic callus using artificial neural networks (ANN). Researchers have also used genetic algorithms ANN to predict and optimize the constituents in the culture medium used in tissue culture studies [10,11]. However, there needs to be more research exploring the viability of using ML algorithms as an alternative to morphological feature-based sorting algorithms. The closest research data publicly available pertains to the classification of plant embryos via absorption, transmittance, and reflectance data processed from digital images [12] and is different from the techniques explored in this paper. Our group has previously demonstrated a method for distinguishing between mature embryos having a range of morphological properties such as length, width, and cotyledon count. This distinction could be used to track the potential of these embryos for plant formation [5]. This exercise has been possible because of the in-house developed SE Fluidics system [4], which can preprocess, image, and sort a high volume of somatic embryos in an automated fashion. In this paper, we use the available information on germination-competent embryos to investigate an alternative way of sorting the good embryos from bad and detecting the embryos' orientation, using ML. We have used convolution neural network (CNN) and support vector machine (SVM) based ML techniques for classifying mature somatic embryos. This model will give the user another tool to segregate the mature somatic embryos and, when appropriately tuned (discussed later), achieve improved image processing speeds over morphology-based segregation.
Methods
The SE Fluidic system [4,13] provides an integrated and automated approach for processing the mature somatic embryos that are produced on a large scale in bioreactors or on solid media in a petri dish. This is achieved through a network of flow loops that serve as the conduit for the fluidic transportation medium carrying the mixture of mature embryos embedded in proembryogenic mass (PEM). The SE Fluidics system can be divided into three main inline subsections based on their significant functionalities, as shown in the schematic in Figure 1. The major subsystems are the extraction section, separation section and the deposition section. The extraction section is designed to allow the embryos-PEM mixture to be introduced into the system within a sterile environment. The separator section is designed to segregate the embryos from (PEM) and sort the embryos according to a predefined morphology-based acceptance criterion. The good embryos are deposited on the deposition tray in the deposition section and the unwanted embryos are collected separately to be periodically removed from the flow loop. The deposition rate from a single deposition tube depends on the quality of the embryos and could vary from 1 to 0.25 Hz. Through multiplexing the deposition rate increases linearly.
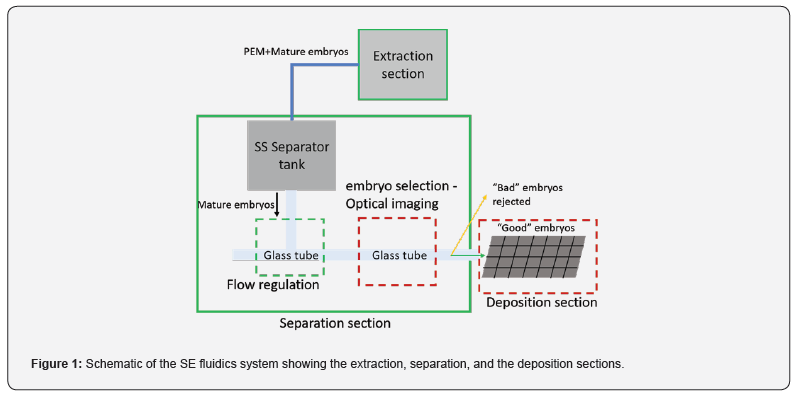
A schematic of the SE Fluidics system is provided in Figure 1, showing the various subsystems in a continuous sterile flow loop. The imaging takes place within a glass tube at the red dashed rectangular region. The unwanted PEM is separated from the mature somatic embryos within the stainless steel (SS) separator tank and the aqueous-suspended embryos descending from the separator tank are regulated to sequentially (one-by-one) enter the horizontal portion of the glass tube and are pushed towards the imaging section where they are imaged. The captured image is processed in-line using morphology-based image processing operations, to be classified as accepted or rejected along with orientation detection. We see that the SE Fluidics system provides an automated platform for imaging embryos at an industrial scale that allows the creation of an image library of all the embryos.
This paper utilizes the images from this library to explore two different machine learning algorithms and the results have been compared with morphologically driven image analysis. A database of labelled images has been carefully assembled from the image library generated by the SE Fluidics system. The images have been captured using two Prosilica GC1290 cameras fitted with 25mm fixed focal length lens from Edmund Optics Inc. The images were saved in a monochrome format with an image size of 501 X 400 pixels having a resolution of approximately 70 pixels/mm. The glass tube is backed with black colored paper and a LED light source illuminates the scene from the front. The captured images are processed on an Optiplex 990 computer running on core i7 processor, 16 GB of RAM, solid state hard drive and a Gigabyte GeForce GT 1030 2GB Low Profile graphics card.
Firstly, an image of the scene is taken with the camera without the embryos and is used as a reference to subtract the light intensity from the images containing the embryos. A couple of sample images are shown for reference in Figure 2. The images are then processed by the morphological image processing algorithm and are classified into two categories and labelled into two folders as accepted and rejected. The accepted embryo images are further classified into right or left orientations. The orientation is determined based on the orientation of the cotyledon side with respect to the flow direction (in this case, flow is from left to right of the image) of the embryo-suspended medium within the glass tube. For example, if the suspended embryo is moving from left to right of the image and the cotyledon is on the left side of the image, the orientation is considered left (Figure 2a).
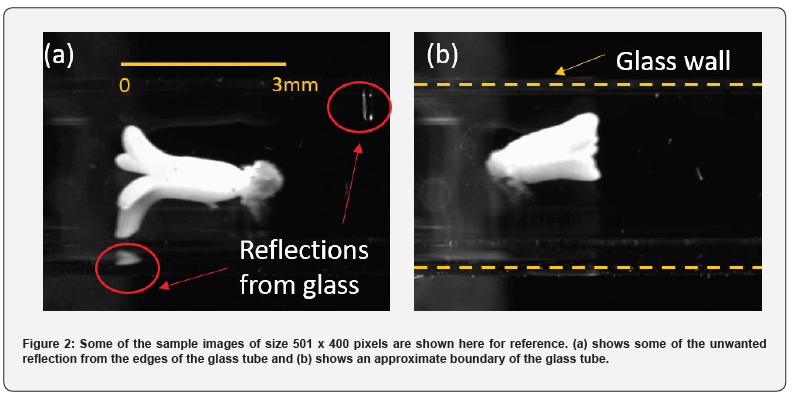
Image pre-processing
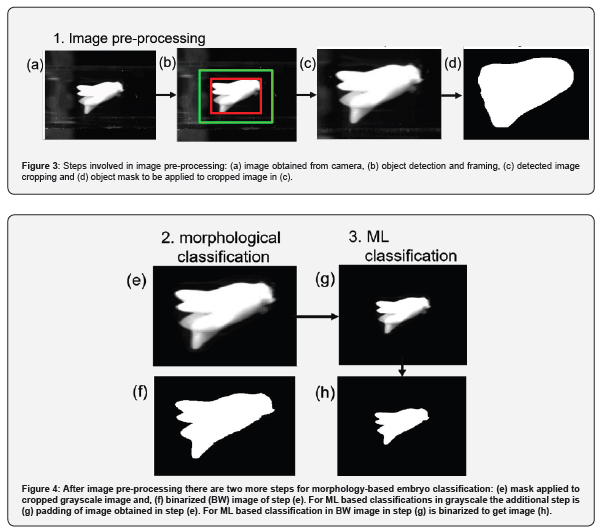
The captured images are processed before being evaluated by the morphological evaluation program. Some unwanted artifacts, such as bright spots emanating from the reflection against the glass walls, as illustrated in Figure 2, and air bubbles are removed. Figure 3 shows the different pre-processing steps involved. First, the embryo is detected by finding the most significant area above a given threshold intensity. The red rectangular bounding box illustrated in Figure 3(b) shows the detected embryo in the image. The detected embryo is cropped along the green-colored box, ensuring that the region of the embryo near its envelope, closely encapsulated by the red box, is not discarded by the subsequent cropping operation. The cropping operation results in differently sized images and is shown in Figure 3(c). As shown in Figure 3(d), an image mask is created, slightly larger in shape compared to the detected embryo. This mask is applied to the grayscale image such that everything outside the mask region is forced to be completely black (i.e., a pixel value of 0). This step removes all unwanted background noise, such as air bubbles, glass reflections, and unwanted floating particulate matter. The masked grayscale image shown in Figure 3(e), is binarized to obtain a black and white (BW) image. These BW images provide information about the embryo envelope, shown in Figure 3(f), and are used for all morphology-based evaluations. Two additional processing steps are required for ML-based classification, as discussed next. As discussed before, different-sized embryos result in differently sized cropped images that are not usable for training classifiers. The first step is to ensure image uniformity in pixel width and height by padding the edges of smaller images with zeros. The grayscale image obtained from Figure 3(e) is first padded with zeros along its edges to ensure image pixel height and width consistency across all images. A resizing operation follows this to obtain the grayscale image shown in Figure 3(g). Resizing the padded photos to 50% of their original pixel dimensions reduces computational time while training the networks and during real-time image classification. The additional steps required from the morphology-based evaluation to ML classification result in increased processing times, which have been captured in this analysis and presented in the next section.

Morphology-based classification
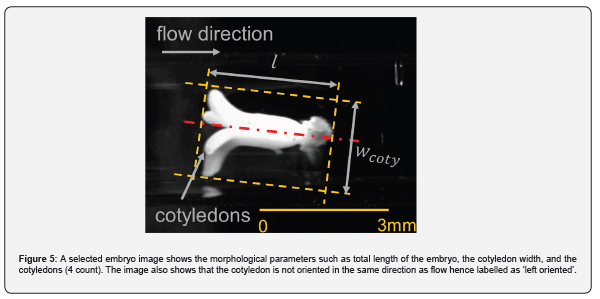
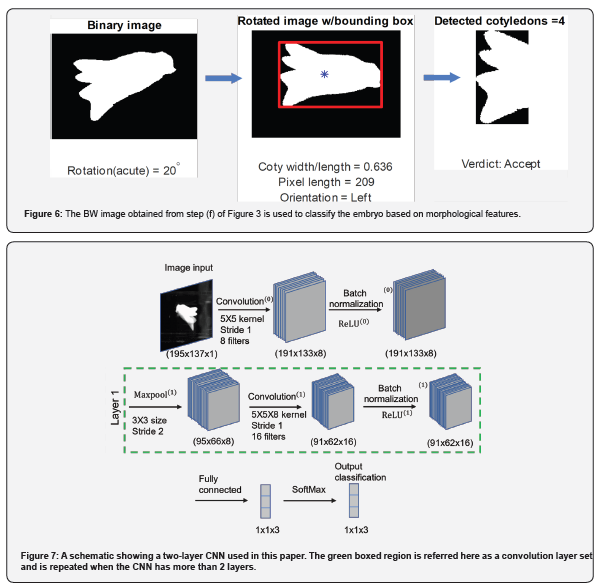
A MATLAB code is developed that determines the morphological features of the embryos such as, cotyledon width (𝑤𝑐𝑜𝑡𝑦), embryo length (𝑙), number of cotyledons (𝑛𝑐𝑜𝑡𝑦), and the ratio of embryo width to length (𝑤𝑐𝑜𝑡𝑦/𝑙). These critical dimensions are shown in Figure 5. Additionally, the algorithm determines the cotyledon's orientation, which could be a useful parameter if the user is concerned about the orientation with which the cotyledon is deposited on the deposition tray. The image is rotated to align it with the flow, and then morphological measurements are carried out. For the batch of photos processed here, some of the selection criterion is based on the publication of [11] where the cotyledon width to length ratio, 0.5 < 𝑤𝑐𝑜𝑡𝑦/𝑙 < 0.7, 𝑛𝑐𝑜𝑡𝑦 ≥ 2, and 𝑙 > 1.43 𝑚𝑚 have been chosen to be the criterion for accepting the embryos. The algorithm is flexible to adapt to the user's morphological selection criteria. A series of three images is illustrated in Figure 6, where the morphological classification is shown in action starting with the images in Figure 3(f). The time elapsed between opening the image for processing and the last classification step, where the image is classified either in 'Accept Left', 'Accept Right' or 'Reject' categories, is recorded for each processed image. For BW images, the processing time is the sum of the time taken from step (a) of Figure 3 to step (f) of Figure 3 (or step (e) of Figure 3 for grayscale images). This time includes the entirety of pre-processing steps (𝑡𝑝𝑟𝑒) plus the processing time of applying mask to cropped image Figure 3(c) (𝑡𝑚𝑎𝑠𝑘), followed by the morphological algorithm that classifies the image (𝑡𝑚𝑐). Mathematically, the total processing time required for morphology-based classification, 𝑡𝑡𝑜𝑡, 𝑚 = 𝑡𝑝𝑟𝑒 + 𝑡𝑚𝑎𝑠𝑘 + 𝑡𝑚𝑐.
ML-based classification
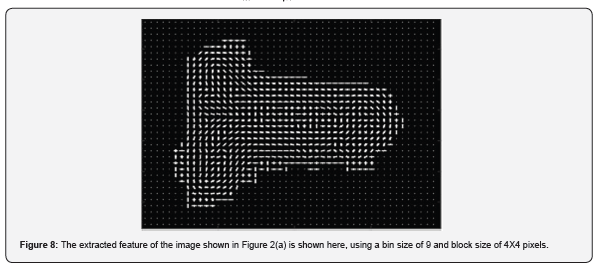
Typically for image classification, CNN primarily involves convolution layers for feature extraction, pooling layers for feature compression and fully connected layers for image classification. Multiple bundles of such layers can be used in succession to learn features with increasing levels of complexity. In this study, a multi-layer CNN is designed with each layer containing a convolution, batch normalization, rectified linear unit (ReLU) and pooling layers. For every convolution layer we have used a fixed kernel size of [5, 5] with a gradually incrementing number of filters for each successive layer (8, 16, 32 and 64 channels). The pooling layers are used to reduce the spatial dimensions of the feature maps, thus reducing the computational effort of the neural network. In our numerical experiments, we have used a fixed maxpool matrix size of [3,3] across all the layers with a step size of vertical and horizontal traversal (stride) of 2 pixels. The convolution layer sets are followed by a fully connected layer and a SoftMax layer [14] which categorizes the images into three categories – 'accept left', 'accept right' and 'reject'.
For the SVM algorithm, the feature extraction has been performed by extracting histogram of oriented gradients (HOG) [14] using MATLAB's inbuilt function 'extractHOGFeatures'. The gradient of the image intensity is first calculated both in x and y directions, followed by image discretization into small blocks of size 4X4 pixels. For each block a 9-point (if no. of bins is = 9) histogram is calculated using 9 bins. The orientation angle of 0-180 degrees is equally spaced for each bin and the gradient magnitude corresponding to its gradient angle is stored in the appropriate bin – creating a histogram for the block. This procedure is repeated for all the blocks within the given image. In our simulations, we have varied the no. of bins from 9 to 36 (9, 18, 27, 36), to capture the finer gradient variations occurring within the images. An extracted feature on a sample image using HOG is shown in Figure 8.
Both the CNN and the SVM networks were trained with 240 images in each of the three labeled image categories, namely 'Accept Left', 'Accept Right' and 'Reject'. To understand the effect of increasing hidden layers, on the accuracy and the processing times of the images, networks were trained with the following different - 2, 3, 4, 5, 6 and 7 layers. In each trained network, 70% of the images were reserved for training and 30% for validation. The images were randomly selected into the training and validation pools. Hence, every time a network is trained with the same number of layers, a slightly different accuracy is achieved because of the randomization of the selected images. To minimize the effect of random image selection affecting the final accuracy of the trained network, ten networks were trained for each of the CNN layers to fully capture the variation in its accuracy. A similar treatment was performed for the SVM network training too. Here, instead of layers, each selected 'Bin No.' was iterated ten times, to obtain ten different networks for each 'Bin No.'.
Moreover, to understand the effect of grayscale vs binary images, the entire process described above was repeated for training and validating images in (1) grayscale format and (2) binary format. This exercise is performed to ascertain if the additional image information extracted by the machine learning algorithms via the grayscale images results in improvement in the classification accuracy. For grayscale images, the total processing time taken is the summation of 𝑡𝑝𝑟𝑒, 𝑡𝑚𝑎𝑠𝑘, padding and resizing time (𝑡𝑝𝑎𝑑), and the ML processing time (𝑡𝑀𝐿). Therefore, the total processing time required for ML-based classification, 𝑡𝑡𝑜𝑡, 𝑀𝐿 = 𝑡𝑝𝑟𝑒 + 𝑡𝑚𝑎𝑠𝑘 + 𝑡𝑝𝑎𝑑 + 𝑡𝑀𝐿. The total processing time of the 𝑖𝑡ℎ image for the ML based classification has been made non-dimensional, i.e., 𝜏𝑖 = (𝑡𝑡𝑜𝑡, 𝑀𝐿⁄𝑡𝑡𝑜𝑡, 𝑚) 𝑖. This allows us to easily understand the time benefit or penalty associated with ML based classification process.

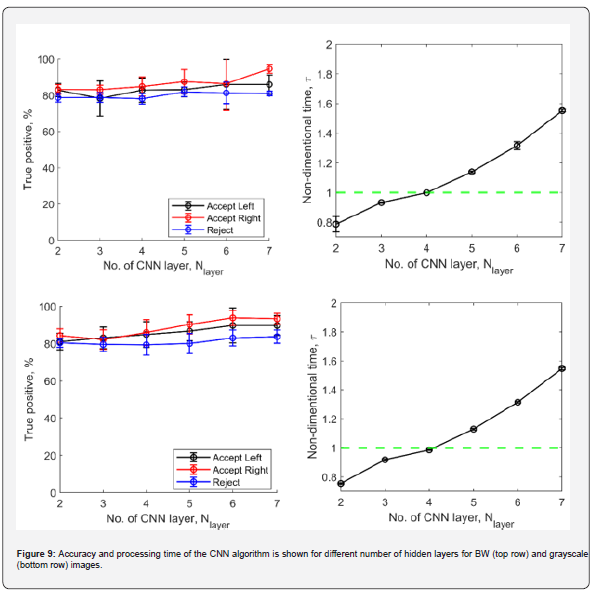
The plots generated in the next section have been made using a total of 556 fresh images (39 in 'Accept Right', 79 in 'Accept left' and 438 in 'Reject') never 'seen' by the machine learning algorithm. For a given layer or bin number, all 556 images are classified, and a confusion matrix [15] is generated. The true positives for each classification category are the major diagonal elements of the confusion matrix. To obtain the true positive percentage (TPP) corresponding to each category, the major diagonal elements are divided with the total images i.e. 556 and multiplied by 100. The process is repeated for each of the ten such trained networks with a given layer or bin number. The mean TPP and standard deviation is evaluated for all ten iterations and presented in the accuracy plots in Figure 9 and Figure 10. For a given iteration of a layer or bin number, mean processing time for ML is divided by the mean morphology-based classification to obtain the processing time plots, i.e. 𝜏𝑖𝑡𝑒𝑟𝑎𝑡𝑖𝑜𝑛 = 〈𝑡𝑡𝑜𝑡, 𝑀𝐿〉⁄〈𝑡𝑡𝑜𝑡, 𝑚〉. The non-dimensional time 𝜏 used in the plots below is the mean value of the ten iterations for a single layer/bin number, plotted along with error bar showing one standard deviation from the mean value.
Results and Discussion
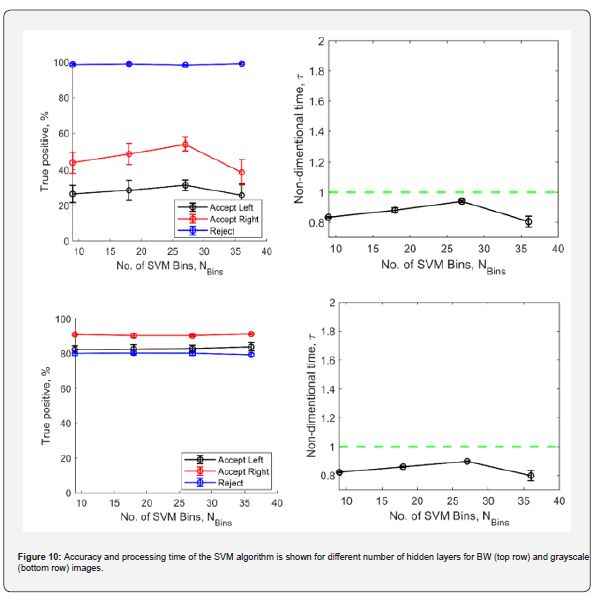
Figure 9 shows the accuracy and processing time plots for CNN based classification for BW and grayscale images. The plots show that with increasing number of convolution layers, the accuracy increases along with increase in processing times for both BW and grayscale images. This is expected as increasing the number of hidden layers increases the total amount of useful features extracted by the trained network, resulting in an increased network complexity. In BW images, the ML algorithm is just presented with the outline of the embryos. Even though the embryo envelope remains the same, the shadow of the foreground cotyledons on the background ones provides more information for the ML algorithm for the grayscale images. This results in a slightly improved accuracy of the grayscale images. In both BW and grayscale images, if the number of layers is ≤ 4, we can take advantage of a slightly improved processing speed (~1.4% - 4.3% faster) when compared with morphology-based classification. The processing time is observed to be monotonically increasing, within the test envelope of two to seven convolution layers, Figure 10 shows the accuracy, and the processing time plots for SVM based classification for BW and grayscale images. For all the test cases, SVM based classification consistently resulted in total processing times that are always observed to be lesser than morphology-based evaluation. However, for BW cases the accuracy is below par when compared with CNN based classification process. The algorithm underperforms when classifying between accepted left and accepted right images in BW. This once again is assumed to be because of the lack of information available in the BW images which just shows an outline of the embryo without providing any pixel information of the cotyledons and their associated shadows.
It is observed that the processing time monotonically increases to 27 bins, beyond which it reduces at 36 bins. This result was consistently obtained across multiple runs of the simulation on different computing systems over different days, leading the authors to believe the reason to be mathematical origin. The number of bins can affect the feature space and the number of support vectors, which can indirectly affect the processing time of the SVM algorithm's implementation in MATLAB. It is possible that increasing the number of bins led to a sparser feature representation [16], which can be faster to process. For example, if most of the bins have low values for a given image (which is the case for both BW and grayscale images here), then many of the features in the high-dimensional feature space will be zero, leading to faster processing times. This information is useful to optimally select the number of bins to avoid data overfitting. Overall, the relationship between the number of bins and processing time in SVM image classification is complex and can depend on many factors, including the specific implementation of the algorithm and the characteristics of the image data.
Conclusion
In this paper, we have explored using machine learning based sorting as opposed to traditional morphology-based sorting algorithm. We have provided an outline of the SE Fluidics setup that provides the capability of collecting thousands of images of somatic embryos in an automated fashion. Through these images, the mature embryos have the potential to be tracked from the deposition-germination planting stages and hence can be mapped with the genetic gains expected from these somatic embryos. The images used in this investigation are obtained from the SE Fluidics system used for segregation, singulation and sorting of mature somatic embryos of Norway spruce. Out of the two tested ML methodologies, namely CNN and SVM, we have observed that SVM based sorting when applied to grayscale images can produce faster processing times than morphology-based sorting along with a very good accuracy level. Moreover, increasing the bin size does not provide any appreciable improvement in the accuracy of classifying the embryos. Using a typical bin size of 9, results in accuracy across all three classifications namely, 'reject', 'accept right' and 'accept left' of 80.1%, 82.3% and 90.9% respectively along with a 17.6% faster processing speed.
This work provides a preliminary study of exploring machine learning based sorting for somatic embryos with further scope of optimizing the ML pipeline and exploring other tuning parameters available in both CNN and SVM image sorting techniques. The preliminary results observed in this paper show that ML, especially SVM, could be effectively employed to sort images of somatic embryos in real-time as well as identify its orientation for in-line automated operation in a SE Fluidics system. However, in this work we have restricted the classification process by self-imposing the three categories of classification. In the future, we will allow the ML algorithm to determine the groupings, i.e., unsupervised learning.
Acknowledgement
We acknowledge helpful assistance from Dr. Cuong Le Kim and Ms. Shannon Johnson.
References
- FAO (2015) FAO assessment of forests and carbon stocks, 1990-2015. In FAO.
- Mckendry P (2002) Energy production from biomass (part 1): overview of biomass. Bioresour Technol 83(1): 37-46.
- Rytter L, Ingerslev M, Kilpeläinen A, Torssonen P, Lazdina D, et al. (2016) Increased Forest biomass production in the Nordic and Baltic countries - A review on current and future opportunities. In Silva Fennica. Finnish Society of Forest Sci 50(5).
- Aidun CK, Egertsdotter U (2018) SE Fluidics System. Springer pp. 211-227.
- Le KC, Weerasekara AB, Ranade SS, Egertsdotter EMU (2021) Evaluation of parameters to characterise germination-competent mature somatic embryos of Norway spruce (Picea abies). Biosys Engineer 203: 55-59.
- Hesami M, Maxwell A, Jones P (2020) Mini-Review Application of artificial intelligence models and optimization algorithms in plant cell and tissue culture. Appl Microbiol Biotechnol 104(22): 9449-9485.
- Jamuna KS, Karpagavalli S, Vijaya MS, Revathi P, Gokilavani S (2010) Classification of Seed Cotton Yield Based on the Growth Stages of Cotton Crop Using Machine Learning Techniques. 2010 Int Confer Adv Comput Engineer, pp. 312-315.
- Vlasov AV, Fadeev AS (2017) A machine learning approach for grain crop seed classification in purifying separation. J Physics: Confer Series 803(1).
- Niazian M, Sadat-Noori SA, Abdipour M, Tohidfar M, Mortazavian SMM (2018) Image Processing and Artificial Neural Network-Based Models to Measure and Predict Physical Properties of Embryogenic Callus and Number of Somatic Embryos in Ajowan (Trachyspermum ammi (L.) Sprague). In Vitro Cellular and Developmental Biology - Plant 54(1): 54-68.
- Arab MM, Yadollahi A, Shojaeiyan A, Ahmadi H (2016) Artificial neural network genetic algorithm as powerful tool to predict and optimize in vitro proliferation mineral medium for G × N15 rootstock. Front Plant Sci 7: 1526.
- Jamshidi S, Yadollahi A, Ahmadi H, Arab MM, Eftekhari M (2016) Predicting in vitro culture medium macro-nutrients composition for pear rootstocks using regression analysis and neural network models. Front Plant Sci 7: 274.
- Timmis R, Toland R, Ghermay M, Carlson WC, Grob JA (2015) Image classification of germination potential of somatic embryos.
- Egertsdotter U, Ahmad I, Clapham D (2019) Automation and scale up of somatic embryogenesis for commercial plant production, with emphasis on conifers. Front Plant Sci 10: 109.
- Bishop CM (2006) Pattern Recognition and Machine Learning (1st). Springer New York, NY.
- Shultz TR, Fahlman SE, Craw S, Andritsos P, Tsaparas P, et al. (2011) Confusion Matrix. In Encyclopedia of Machine Learning. Springer US pp. 209-209.
- Platt JC (1999) Probabilistic outputs for support vector machines and comparisons to regularized likelihood methods. Advances in Large Margin Classifiers 10(3): 61-74.






























