Nuclear SSR-based Genetic Diversity and Structure Analysis of Vitis Rootstocks
Bibi Androniki C1, Lefort François2 and Roubelakis-Angelakis Kalliopi A3
1Institute of Molecular Biology and Biotechnology, Foundation for Research and Technology, Vassilika Voutes, Greece
2Plants and Pathogens group, Institute Land Nature and Environment, HEPIA, HES-SO University of Applied Sciences and Arts Western Switzerland,Geneva, Switzerland
3Laboratory of Plant Physiology and Biotechnology, Department of Biology, University of Crete, Greece
Submission: November 05, 2022; Published: November 29, 2022
*Corresponding author: Androniki Bibi, Institute of Molecular Biology and Biotechnology, Foundation for Research and Technology, Vassilika Voutes, Greece
How to cite this article: Bibi Androniki C, Lefort François, Roubelakis-Angelakis Kalliopi A. Nuclear SSR-based Genetic Diversity and Structure Analysis of Vitis Rootstocks. Agri Res & Tech: Open Access J. 2022; 27 (2): 556365. DOI: 10.19080/ARTOAJ.2022.27.556365
Abstract
We have genetically identified 43 plants representing 28 rootstocks from three different ampelographic collections in Greece with the use of seven nuclear single sequence repeats (SSR) previously characterized. Analysis of SSR alleles and a similarity dendrogram construction was followed by a STRUCTURE and a hierarchical STRUCTURE analysis. We also compared the obtained genetic data with literature data about the origins of these rootstocks. In most cases known pedigrees of the studied rootstocks were confirmed but in some other cases microsatellite profiling did not support the pedigrees data, casting a doubt about the identity of a particular rootstock or on pedigree given by the literature. The genetic profiling of these plants enabled to reassign wrong identities in the collections and proved to be very efficient in assessing rootstock identities in a commercial dispute. The identity of an unknown rootstock was also determined. This work provides the molecular fingerprinting of 28 rootstocks and adds to their ancestry characterization.
Keywords: Vitis; Grape rootstock; Simple sequence repeats (SSR); Molecular identification; Structure analysis; Genetic diversity
Introduction
Historical evidence indicates that European grape (Vitis vinifera L.) was introduced from Europe to the American continent in the 16th century (Mullins et al. 1992). However, it failed to thrive due to pests and deceases, mainly phylloxera (Phylloxera vastatrix) and nematodes. The ability of north American Vitis species to resist pests and diseases was soon recognized and they were used first as phylloxera resistant hybrids with Vitis vinifera, then as rootstocks for the sensitive European cultivars [1]. In commercial viticultural practice, rootstocks have been in use since the middle of the 19th century to protect grapevines form soil borne pests but also to ameliorate the effect of unfavourable soil and environmental conditions, such as lime, salinity, and drought. In addition, rootstocks can affect the grapevine growth, fruit yield, and other economically important vine attributes. From the late 1850’s, grafting on various Vitis species or interspecific Vitis hybrids became a common practice in California as a consequence of the introduction of phylloxera [2]. Later, this practice was spread in virtually all viticultural regions over the world. Numerous Vitis species and hybrids were tested for their resistance to phylloxera Makee et al. [3] of which the most efficient have been extensively used [4-6]. In addition to phylloxera, nematodes represent another pest threatening vineyards [7]. Thus, nematode resistant rootstocks were developed from the 1930’s [8-17].
Although the commercially used rootstocks are much fewer compared to grapevine cultivars, the importance of efficient and accurate genotype identification is of equal importance, if not more. There is a necessity for the grower to have an absolute certitude about the rootstocks on which his selected cultivars are grafted due to high cost for establishing a vineyard and to the long time required for full production [6]. Traditionally, the identification of Vitis genotypes, both cultivars and rootstocks, was based on ampelographic characters and biochemical markers, such as isoenzymic profile, until the progress in molecular biology allowed the development of numerous molecular markers for genotype identification. Among them are variable number tandem repeat (VNTR), short tandem repeat (STR), restriction fragment length polymorphism (RFLP), randomly amplified polymorphic DNA (RAPD), amplified fragment length polymorphism (AFLP), inter simple sequence repeat (ISSR), sequence characterized amplified region (SCAR), expressed sequence tag (EST), single nucleotide polymorphism (SNP), and microsatellites, single sequence repeats (SSR).
In early 1990’s, five RFLP probes designed from Vitis vinifera Chardonnay DNA were developed, which enabled to distinguish 16 rootstocks [18,19]. Concomitantly, Boursiquot and Para [20] assayed one isoenzyme system, previously characterized on grapevine cultivars Parfitt and Aruselkar [21] for fingerprinting 30 rootstocks authorized for trade in France and were able to distinguish uniquely 5 rootstocks while others clustered in 4 phenotypic groups. Walker and Liu [22] by using 5 isoenzyme systems were able to obtain a unique isoenzyme profile for each rootstock. Research for easier and more reproducible markers went on with the assessment of RAPD markers [23-25]. All the assayed markers, even though successful enough for providing a single identity profile for a rootstock, were dominant markers, which do not allow to differentiate between homozygotes and heterozygotes and thus to rebuild pedigrees, and consequently to carry out a follow up of selection processes, for creating and selecting new rootstocks for the future. Additionally, the methods are costly, cumbersome, affected by environmental conditions (isoenzymes) Royo et al. [26]. or subject to low reproducibility between laboratories. These disadvantages were overcome by the availability of specific microsatellite markers developed and characterized in a large number of grapevine cultivars [27].
The use of nuclear SSR markers for genotype identification of cultivars and hybrids in Vitis vinifera and Vitis species has been developing fast and assessed for a variety of us [28,29]. The nuclear data were complemented by the use of chloroplast microsatellite [30,31]. Microsatellite profiling also opened the path for gene pools studies Sefc et al. [32,33], molecular assisted management of genetic resources [34-37]. Furthermore a neural network algorithm and a combination of RAPD and SSR molecular markers were used for the genetic characterization of different rootstock varieties [29]. Riaz et al [38] developed nuclear and chloroplast SSR fingerprint data from rootstocks in germplasm collections, compared them to develop a reference dataset, and carried out parentage analysis to resolve previously reported, and determine new breeding records. They refined and updated the parentage of 26 rootstocks based on 21 nuclear and 14 chloroplast markers. Results indicated that 39% of the genetic background of analyzed rootstocks originated from only three accessions of three grape species: Vitis berlandieri cv. Rességuier 2, V. rupestris cv. du Lot and V. riparia cv. Gloire de Montpellier. Results determined that Rességuier 2 is the maternal parent for 14 commercial rootstocks, 9 of which are full-sibs with Gloire de Montpellier as the paternal parent. Similarly, du Lot is the paternal parent of nine rootstocks. Here we present, microsatellite profiling data of 43 plants representing 28 rootstocks from 3 ampelographic collections in Greece, in an effort to support the practical application of microsatellite profiling for chasing management mistakes, solving commercial dispute or recovering lost information. In addition, the possible pedigrees of this rootstock gene pool were assessed. The previously reported pedigree information was correct for only eight of the 28 rootstocks [39]. Additionally, misnaming of rootstocks originating from a commercial orchard was solved by applying microsatellite profiling to plants sampled in the vineyard.
Materials and Methods
Plant material
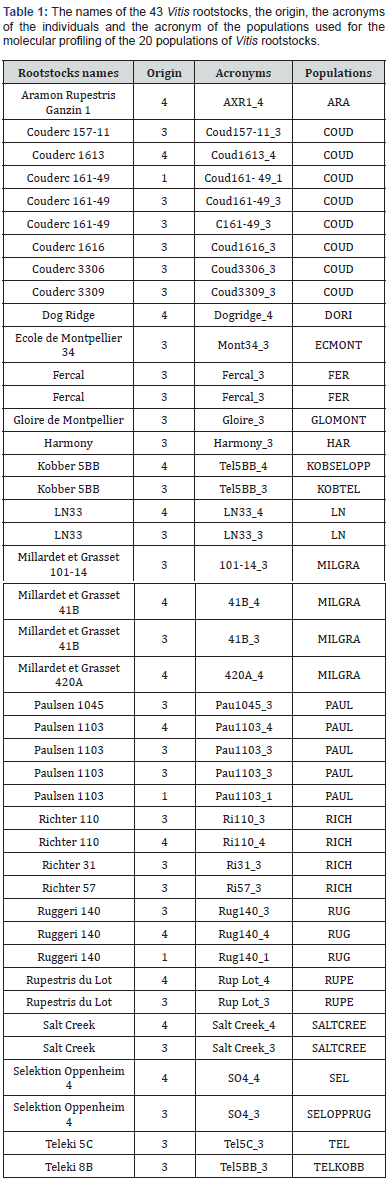
Leaves from 43 Vitis spp. and hybrids used as rootstocks (Table 1) were collected from the Laboratory of Plant Physiology and Biotechnology at the University of Crete, Heraklion, the Institute of Floriculture, Horticulture and Viticulture, National Agricultural Research Foundation (NAGREF), Katsabas, Heraklion and the Institute of Vine and Wine, Lykovrisi (NAGREF), Athens. Leaves were kept frozen at -80°C until used. Additionally, leaves from 12 plants in a private nursery and one from a private vineyard were sampled and stored in the same way.
DNA extraction
DNA was extracted from 100 to 150 mg FW of leaf tissue according to a micro-method of DNA purification described elsewhere [28].
Microsatellite PCR and microsatellite profile analysis
Amplification primer sequences for 7 nuclear microsatellite loci from Vitis vinifera, UCH2, UCH11, UCH12, UCH19, UCH29, UCH35 and UCH40 Lefort et al. [30], were used for DNA amplification. PCR amplifications were carried out as described previously [30]. PCR products sizing and SSR profiles analysis were performed as described elsewhere [28].
Genetic Analysis and Neighbor‑Joining Tree Construction
The allele sizing per locus was based on published repeat patterns. The data matrixes were produced and the genetic diversity measures were determined for each employed locus across all fingerprinted genotypes. These methods included the individual locus polymorphic information content PIC; Botstein et al. [40], the observed heterozygosity (HO), and the expected heterozygosity (HE). PIC, HO, HE, estimated frequency of null alleles and probability of identity (PI) were calculated with the software CERVUS ver. 3.0.3 [41]. A matrix was produced employing Nei’s distance matrix within GenAlEx [42]. Furthermore, a neighbor-joining tree was produced using MEGA 7 Tamura et al. [43] Kumar et al. [44] starting with the Nei’s distance matrix. Twenty (20) populations were formed from 43 Vitis rootstocks. In order to determine the divergence between the different population, pairwise Fst measurements were calculated according to Weir and Cockerham [45] using GenAlEx 6 [42]. Analysis of molecular variance (AMOVA) was also performed to assess the genetic structure of the 20 population, using GenAlEx 6.
Population Structure
The STRUCTURE 2.3.4 software was used to analyse the germplasm genetic structure. This software utilize a Bayesian clustering algorithm to identify subpopulations, assign individuals to them, and estimate population allele frequencies [46]. The analysis was carried out using a burning period of 10,000 iterations and a run length of 200,000 MCMC replications. We tested a continuous series of K, from 1 to 10, in 10 independent runs. We did not introduce any prior knowledge about the origin of the population, and assumed correlated allele frequencies and admixture [47]. For selecting the optimal value of K, ΔK values Evanno et al. [48] were calculated using STRUCTURE harvester [49]. POPHELPER, proposed by Francis [50], was used to analyse and visualize population structure. Furthermore, a “hierarchical STRUCTURE analysis” was applied in this study by running STRUCTURE on subsequently partitioned data, as suggested by Pritchard and Falush [47].
Results and Discussion
Genetic diversity analysis of Vitis rootstocks
Genetic diversity is caused by multitude factors. Many evolutionary forces can change genetic frequencies of crop species, which in turn determine the genetic diversity of a population. Gene flow, genetic drift, and mutations are among the evolutionary forces, which affect gene pool of a given population [51]. Riaz et al. [39] provided valuable historical information on the diachronic efforts and the contributors worldwide in the production of the existing and extensively used grape rootstocks. In the last three decades, biochemical markers (isoenzymes), DNA-based markers, such as RAPD, RFLP, AFLP and SSR have been used to fingerprint grape rootstocks at different geographical ampelographic collections [18,22,36,37]. In addition, SSR markers were used in Spain to confirm identification (De Andris et al.), and elucidate the parentage of selected rootstocks (Laucou et al., Crespan et al., Garris et al.). Some other attempts exists in order to assess the clonal variation and possible parental relations, focusing primarily on the diversity of Teleki lines and comparing them with other rootstocks [36]. The results revealed that the existing information in the database of many historical rootstocks is not always accurate and should be corrected (De Andris et al.). This work aims to add to this effort, using SSR because of their polymorphism, reproducibility, and codominant nature.
Herein, a total of 43 individuals from 20 Vitis rootstocks were genotyped employing 7 SSR loci (Table 1). Polymorphic fragments were reproducibly amplified at all 7 SSR loci and microsatellites profiles of these 43 plants are given in Table S1. Microsatellite profiles of 13 plants from a private nursery and a private vineyard are given in Table S2. The genetic diversity measures determined for each employed locus across all fingerprinted genotypes are shown in Table 2. The number of amplified alleles (k) by each SSR primer pair varied from eleven for UCH19 to twenty-three for UCH12 with an average number of alleles per locus of 15.857. Different amplified alleles are shown in Table S4. Observed heterozygosity (Ho) ranged from 0.721 in UCH2 and UCH35 to 0.930 in UCH19, with an average value of 0.827, while the expected heterozygosity (He) ranged from 0.783 in UCH73 to 0.941 in UCH12 with an average value of 0.8774. Bianchi et al. (2020) found rootstock heterozygosity values of Ho = 0.099 and Ho = 0.734 for SNPs and SSRs, respectively, whereas SSR He value of core collection was higher than the values detected in other studies Emanuelli et al., Dzhambazova et al. [35] Sefc et al., 1998; Upadhyay et al., 2007) confirming the uniqueness and preciousness of the analyzed germplasm collection. In the study by Emanuelli et al. (2013) using SSR, the subset of rootstocks revealed the highest number of alleles (405) and the highest heterozygosity (0.86), in spite of their relatively small sample size.
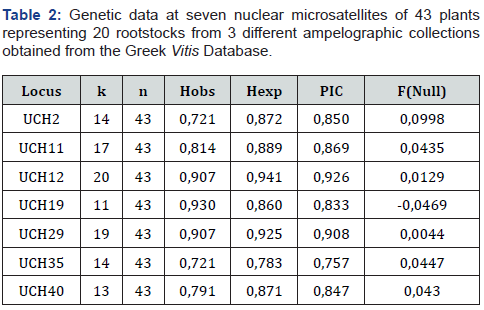
The genotype level of polymorphism was assessed by calculating polymorphic information content (PIC) values for each of the 7 SSR loci. The average PIC was 0.8556 with a minimum value of 0.757 in UCH35 and a maximum value of 0.926 in UCH12. One of the seven loci, the UCH19 exhibited probability of null alleles, F (null), greater than 0.05. Propability Identity (PI) is the probability with which two randomly taken genotypes display the same SSR profile. The probability of obtaining an identical genotype, the cumulative probability of identity, was calculated with a value of 8.9 × 10e−12, using the 7 SSR markers in combination. This number corresponds to a statistical potential of distinguishing a large number of unrelated grapevine genotypes. AMOVA was conducted to determine the variation explained by populations. The results indicated that 30% of the genetic variation (p < 0.0001) resided among populations and 0% (p < 0.0001) resided among individuals. The remaining 70% of the total variation (p < 0.0001) was explained within individuals. The RST value was 0.342 (p < 0.0001). The genetic variation was tested using the FST statistic estimated from pairwise comparisons as a measure for genetic distance between populations and individuals. The results indicated that 24% of the genetic variation (p < 0.0001) resided among populations and 76% resided within individuals (p < 0.0001). The FST value was 0.287 (p < 0.001).
The phylogeny and genetic relations among grapevine cultivars is of great importance in genetic improvement of Vitis germplasm (Emanuelli et al., Marsa et al., Bibi et al.). Emanuelli et al. tested the structure analysis of 111 Vitis rootstocks. The genetic structure of cultivated grapevine has been influenced by human selection and it can be largely understood as a complex pedigree, due to the vast number of higher order pedigree relationships Myles et al. Αn unknown relevance among the grape genotypes affect the study of the genetic structure, resulting in the overestimation of the probable subpopulation number (K) MINE using standard methods Pritchard and Falush, Vos and Didelot [52].
Structure Analysis
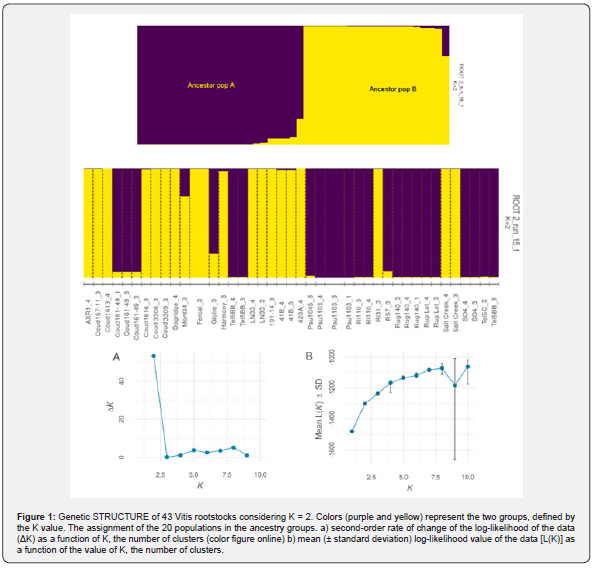
The genetic structure of the germplasm collection was evaluated using STRUCTURE software. The 20 populations of the rootstocks were assigned in its ancestor population as shown in Figure 2. The analysis provided evidence for a significant population structure in this set of rootstocks. A maximum value of the rate of change in the log probability of the data was revealed at K = 2, using Evanno’s method (Figure 1). The highest Delta K value was observed at K = 2 (Figure 2b). The estimated logarithm of probability of the data [L(K)] increased linearly from K = 2 up to K = 10 showing a clear point of inflection (Figure 2a). The estimated population structure inferred from the analysis identifies two genetic groups, Ancestor population (pop) A and Ancestor population (pop) B (Figure 2).
Hierarchical STRUCTURE Analysis
A subsequent round (second round) of STRUCTURE revealed that the two populations A and B (K =2) were further separated as follows (Figure 2):
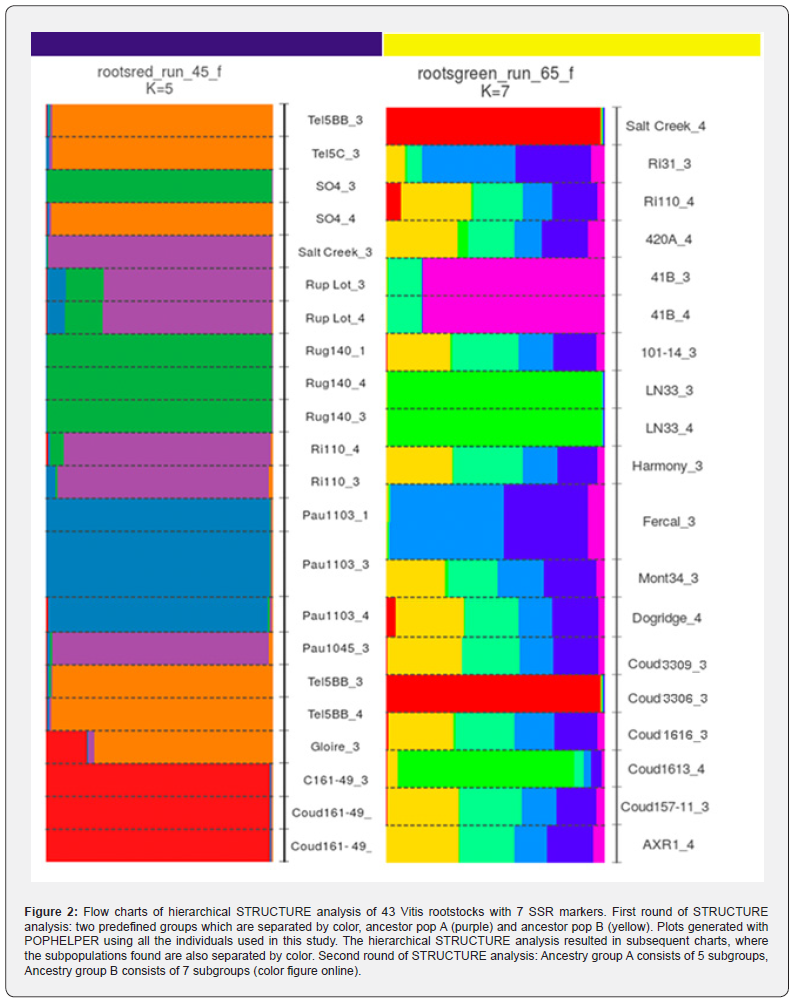
Ancestry group A is subdivided into five distinct subgroups (K=5) (Figure 3). Twenty-three Vitis rootstocks are presented in the ancestry group A (Figure 2). These are Tel5C_3, Tel5BB_3, SO4_3, SO4_4, Salt Creek_3, Rup Lot_4, Rup Lot_3, Rug140_3, Rug140_4, Rug140_1, RI57_3, RI110_3, Pau1103_1, Pau1103_3, Pau1103_4, Pau1045_3, Tel5BB_3, Tel5BB_4, Gloire_3, Coud161- 49_3, Coud161- 49_1. Interestingly some populations appear identical a) Tel5BB_3 (KOBTEL population) and Tel5BB_3 (TELKOBB population) (subgroup-orange), b) Tel5BB_4 with SO4_4 and (subgroup-orange), c) Rug140_1,3,4 with SO4_3 (subgroup-green). Further research is needed to verify these data. Three rootstock populations appear to have individuals in both ancestor populations (1st round) these are Richter (subgrouppurple), Couderc (subgroup-red) and Salt Creek (subgrouppurple). The remaining individuals mingle with each other according to dendrogram and their distances are higher than 0.2.
Ancestry group B is subdivided into seven distinct subgroups (K =7) (Figure 2). Twenty Vitis rootstocks are presented in ancestry group B (Figure 2). These are Salt Creek_4 (subgroupred), RI31_3 (subgroup-light blue), RI110_4 (subgroup-yellow), 420A_4 (subgroup- yellow), 41B_3 (subgroup-magenta), 41B_4 (subgroup- magenta), 101-14_3 (subgroup-light green), LN33_4 (subgroup-green), LN33_3 (subgroup-green), Harmony_3 (subgroup-light green), Fercal_3 (subgroup-light blue), Mont34_3 (subgroup-yellow), Dogridge_4 (subgroup- yellow), Coud3309_3 (subgroup- yellow), C1613_4 (subgroup-green), Coud3306_3 (subgroup-red), Coud157-11_3 (subgroup-yellow), and AXR1_4 (subgroup-yellow). Again interestingly, we can see that none of the populations are close to each other. Their distances are higher than 0.2 (Figure 3). According to Emanuelli et al. (2013), the hierarchical approach needs to be applied to delve deeper into the complex relationships of the Vitis germplasm. Considering the relevant role of rootstocks on environmental stress tolerance, their low genetic diversity reduces the ability of grapevine cultivars to adapt to several environmental constraints. This issue can be overcome by increasing the genetic and phenotypic diversity of the breeding material, including non-conventional material in the further breeding programs.
Genetic Distance Analysis
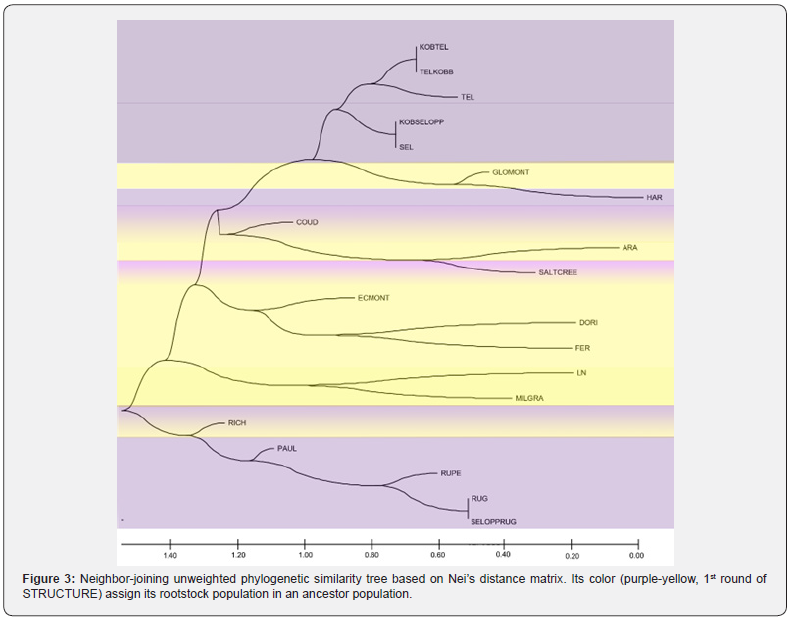
A neighbor-joining tree was built based on Nei’s distance matrix (Figure 3). Based on this phylogenetic tree the 43 Vitis rootstocks form 20 populations belonging to six major clusters. 1) KOBTEL-TELKOBB mingled with TEL and all of them mingled with KOBSELOPP-SEL, all assigned in the same ancestor population A and the same subgroup orange. 2) GLOMONT is mingled with HAR but their big distance becomes obvious with the Structure Analysis. GLOMONT belongs to the ancestor population A and HAR to the ancestor population B. 3) ARA, SALTCREE and COUD appear in the same cluster however many individuals belong to different ancestor populations. 4) ECMONT, DORI and FER belong to the same ancestor population B, with ECMONT and DORI in subgroupyellow and FER in subgroup-light blue. 5) LN and MILGRA form a cluster and belong to the same ancestor population A, LN has individuals in subgroup-green and MILGRA has some in subgroups magenta and yellow. And 6) RICH, PAUL, RUPE, RUG, SELOPPRUG form another cluster.
Solving identity ambiguities
Microsatellite profiles of 13 plants from a private nursery and a private vineyard are given in Table S2, along with profiles of reference rootstocks and cultivars (Paulsen 1103, Ruggeri 140, Couderc 3309 and Soultanina). Phenotypic discrepancies based the need for such microsatellite profiling. From the results it appeared that the allegated Paulsen 1103 was the cultivar Soultanina, that the allegated Ruggeri 140 was an unknown profile not present in the Greek Vitis Database and that an unknown plant from a private vineyard was in fact the rootstock Couderc 3309. This example shows the resolution of microsatellite profiling of rootstocks and cultivars to solve possible commercial disputes between nurseries and their customers [54].
Conclusion
The significance of rootstocks to reduce the impact of biotic and abiotic stresses on V. vinifera cultivars requires the increase of the genetic and phenotypic diversity of the breeding rootstock material. Most of the currently used rootstocks globally were bred nearly a century ago; however, the changing climate has prompted a renewed interest in breeding rootstocks to address Riaz S, et al. current and future vineyard issues. If the lineage of classic grape rootstocks is uncovered, and their inheritance is established and understood, this information can be used to broaden the genetic basis of new rootstocks Riaz S, et al. Therefore, the existing germplasm collections are valuable resources for exploring the genetic and phenotypic diversity and providing new genetic resources to support plant breeding efforts.
Acknowledgement
This work was supported by the INTERREG II programme of the European Union.
References
- Pavloušek P (2016) New Rootstock Hybrids with Vitis Cinerea and Their Resistance to Phylloxera and Callus Formation. In Acta Hortic 1136: 265-268.
- Gastón GG, Encarna GP, Ana BO, Teresa GC, Yerko MM, et al. (2019) Rootstock Effects on Grape Anthocyanins, Skin and Seed Proanthocyanidins and Wine Color and Phenolic Compounds from Vitis Vinifera L. Merlot Grapevines. Journal of the Science of Food and Agriculture 99(6).
- Makee H, Charbaji T, Ayyoubi Z, Idris I (2004) Evaluating Resistance of Some Rootstocks to Grape Phylloxera with in Vitro and Excised Root Testing Systems. In Vitro Cellular and Developmental Biology - Plant 40(2): 225-229.
- Gale G (2003) Saving the Vine from Phylloxera: A Never-Ending Battle. In: Sandler M, Pinder R (Eds), Wine - a Scientific Exploration. Taylor & Francis, London, United Kingdom, p. 70-91.
- Granett J, Walker MA, Kocsis L, Omer AD (2001) Biology and Management of Grape Phylloxera. Annu Rev Entomol 46: 387-412.
- Yin, Lu, Matthew DC, Eric CB, William DH (2019) Grape Phylloxera (Hemiptera: Phylloxeridae), on Cold-Hardy Hybrid Wine Grapes (Vitis): A Review of Pest Biology, Damage, and Management Practices. Journal of Integrated Pest Management 10(1): 1-9.
- Ferris H, Zheng L, Walker MA (2012) Resistance of Grape Rootstocks to Plant-Parasitic Nematodes. Journal of Nematology 44(4): 377-386.
- Kasimatis AN, Lider LA (1967) California Agriculture Extension Service 47.
- Lider LA (1954) Inheritance of Resistance to a Root-Knot Nematode Meloidogyne Incognita Var. Acrita Chitwood) in Vitis Proceedings of the Helminthological Society of Washington 21: 53-60.
- Lider LA (1959a) The Performance of Vigorous Nematode Resistant Rootstocks in Field Trials in the San Joaquin Valley. Am J Enol Vitic 10: 147-151.
- Lider LA (1959b) Nematode Resistant Rootstocks for California Vineyards. California Agricultural Experiments Station Extension Service Leaflet.
- Snyder E (1936) Susceptibility of Grape Rootstocks to Root-Knot Nematodes. US Department of Agriculture Circular 405: 16.
- Bloodworth PJ, Nesbitt WB, Barker KR (1980) Resistance to Root-Knot Nematodes in Euvitis 3 Muscadinia Grape Hybrids.” 3rd International Symposium on Grape Breeding, Davis, California, USA, 275-292.
- Nesbitt WB (1974) Breeding Resistant Grape Rootstocks. HortScience 9: 359-361.
- Harris AR (1983) Resistance of Some Vitis Rootstocks to Xiphinema Index. J Nematol 15(3): 405-409.
- Walker MA, Ferris H, Eyre M (1994) Resistance in Vitis and Muscadinia Species to Meloidogyne Incognita. 78: 1055-1058.
- Cousins PS, Reisch BI, Owens CL (2003) Grape. In: Badenes ML, Byrne DH (eds.) Fruit Breeding, Handbook of Plant Breeding 8, Springer Science+Business Media, LLC.: 225-262.
- Bourquin JC, Otten L, Walter B (1991) Identification of Grapevine Root-Stocks by RFLP. CR Acad Sci Paris Sér III 312: 593-598.
- Bourquin JC, Tournier P, Otten L, Walter B (1992) Identification of sixteen grapevine rootstocks by RFLP and RFLP analysis of nuclear DNA extracted from the wood 31: 157-162.
- Boursiquot JM, Parra P (1992) Application d’une Méthode d’électrophorèse Pour La Caractérisation et La Reconnaissance Des Porte-Greffe. Vitis 31(4): 189-194.
- Parfitt DE, Arulsekar S (1989) Inheritance and Isozyme Diversity for GPI and PGM among Grape Cultivars. J Am Soc Hortic Sci 114(3): 486-491.
- Walker L, liu L (1995) The Use of Isozymes to Identify 60 Grapevine Rootstock (Vitis ). Am. J. Enol. Vitic 46(3): 299-305.
- Xu H, Bakalinsky AT (1996) Identification of Grape (Vitis) Rootstocks Using Sequence Characterized Amplified Region DNA Markers. Hort Science 31(2): 267-68.
- Azab A (2017) CAROB (Ceratonia Siliqua): Health, Medicine, Chemistry. ECB 6(10): 456-469.
- Wolf T, Eimert K, Ries R (1999) Reliable Identification of Grapevine Rootstock Varieties Using RAPD PCR on Woody Samples. Austr J Grape Wine Res 5(2): 34-38.
- Royo J, Cabello F, Miranda S, Moreno S, Itoiz R, et al. (1997) The Use of Isoenzymes in Characterization of Grapevines (Vitis Vinifera L.). Influence of the Environment and Time of Sampling. Scientia Horticulturae 69(3-4): 145-155.
- This P, Jung A, Boccacci P, Borrego J, Botta R, Costantini L, et al. (2004) Development of a Standard Set of Microsatellite Reference Alleles for Identification of Grape Cultivars. Theor Appl Genet 109(7): 1448-1458.
- Sefc KM, Lefort F, Grando MS, Scott KJ, Steinkellner H, et al. (2001) Microsatellite Markers for Grapevine: A State of the Art. In Molecular Biology and Biotechnology of Grapevine, Kluwer Publishers, Amsterdam, pp. 433-463.
- Costa MO, Capel LS, Mora CM, Maria FS (2019) High Genetic Differentiation of Grapevine Rootstock Varieties Determined by Molecular Markers and Artificial Neural Networks. Acta Scientiarum - Agronomy 42: 1-10.
- Arroyo GR, Lefort F, De Andrés MT, Ibañez J, Borrego J, et al. (2002) Chloroplast Microsatellite Polymorphisms in Vitis Species. Genome 45(6): 1142-1149.
- Pervaiz T, Cheng Z, Faheem M, Mu Q, Fang J (2017) Chloroplast Based Genetic Diversity among Chinese Grapes Genotypes. Mitochondrial DNA Part A: DNA Mapping, Sequencing, and Analysis 28(4): 565-569.
- Sefc KM, Machado A, Lefort, F, Botta R, Roubelakis KA, et al. (2000) Microsatellite Variability in Grapevine Cultivars from Different European Regions and Evaluation of Assignment Testing to Specify the Geographic Origin of Cultivars. Theoretic and Applied Genetics 100(3/4): 498-505.
- Sefc KM, Steinkellner H, Lefort F, da Machado AC, Borrego JP, et al. (2003) Substantial Genetic Contribution of Local Wild Vines to European Grapevine Cultivars. Am J Enol Vitic 54(1).
- Lefort F, Roubelakis KA (2000) The Greek Vitis Database, a Multimedia Web-Backed Genetic Database for Germplasm Management of Vitis Resources in Greece. Journal of Wine research 11(3): 233-242.
- Dzhambazova, Hvarleva T, Hadjinicoli A, Tsvetkov I, Atanassov A, Atanassov I (2007) Characterization of Grapevine Rootstocks Using Microsatelite Markers.” Biotechnology & Biotechnological Equipment 21(1): 58-62.
- Jahnke G, Gitta KM, Janos M , Barna S, Erik T, et al. (2011) Analysis of Grape Rootstocks by SSR Markers. Journal International des Sciences de la Vigne et du Vin 45(4): 199-210.
- Laucou V, Launay A, Roberto B, Thierry L, Anne FA, et al. (2018) Extended Diversity Analysis of Cultivated Grapevine Vitis Vinifera with 10K Genome-Wide SNPs. PLoS ONE 13(2): 1-27.
- Bibi AC, Evangelos DG, Andreas GD (2020) Genetic Diversity and Structure Analysis Assessed by SSR Markers in a Large Collection of Vitis Cultivars from the Island of Crete, Greece. Biochemical Genetics 58(2): 294-321.
- Riaz S, Pap D, Uretsky J, Laucou V, Boursiquot JM, et al. (2019) Genetic Diversity and Parentage Analysis of Grape Rootstocks. Theor Appl Genet 132(6): 1847-1860.
- Botstein D, White RL, Skolnick M, Davis (1980) Construction of Genetic Linkage Map in Man Using Restriction Fragment Length Polymorphisms. Am J Hum Genet 32(3): 314-331.
- Kalinowski ST, Taper ML, Marshall TC (2007) Revising How the Computer Program CERVUS Accommodates Genotyping Error Increases Success in Paternity Assignment. Mol Eco 16(5): 1099-1106.
- Peakall R, Smouse PE (2006) GenAlEx 6: Genetic Analysis in Excel. Population Genetic Software for Teaching and Research. Mol Ecol Notes 6: 288-295.
- Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) Software Version 40. Mol Biol Evol 24(8): 1596-1599.
- Kumar S, Stecher G, Tamura K (2015) MEGA7: Molecular Evolutionary Genetics Analysis Version 70 for Bigger Datasets. Mol Biol Evol 33(7): 1870-1874.
- Weir BS, Cockerham C (1984) Estimating F-Statistics for the Analysis of Population Structure. Evolution 38: 1358-1370.
- Pritchard JK, Stephens M, Donnelly P (2000) Inference of Population Structure Using Multilocus Genotype Data. Genetics 155(2): 945-959.
- Falush D, Stephens M, Pritchard JK (2003) Inference of Population Structure Using Multilocus Genotype Data: Linked Loci and Correlated Allele Frequencies. Genetics 164(4): 1567-1587.
- Evanno G, Regnaut S, Goudet J (2005) Detecting the Number of Clusters of Individuals Using the Software STRUCTURE: A Simulation Study. Mol Ecol 14(8): 2611-2620.
- Earl DA, Von Holdt BM (2012) Structureharvester: A Website and Program for Visualizing STRUCTURE Output and Implementing the Evanno Method Conservation. Genet Resour 4(2): 336-359.
- Francis RM (2017) POPHELPER: An R Package and Web App to Analyze and Visualize Population Stru. Mol Ecol Resour 17(1): 27-32.
- Begna T (2021) Role and Economic Importance of Crop Genetic Diversity in Food Security. Role and Economic Importance of Crop Genetic Diversity in Food Security. J Agric Sc Food Technol 7(1): 164-169.
- Vos M, Xavier D (2009) A Comparison of Homologous Recombination Rates in Bacteria and Archaea. The ISME Journal 3(2): 199-208.
- Cuisset B (1997) Development of Stable RAPD Markers for the Identification of Grapevine Rootstocks and the Analysis of Genetic Relationships. Am J Enol Vitic 48: 492-501.
- (2015) Application of Isozymes and SSR Markers for the Analysis of the Genetic Background of Some Rootstocks Derived from Teleki’s Seedlings (Teleki 5C, Kober 5BB, SO4). Mitteilungen Klosterneuburg 65(4): 221-236.






























