Investigating Incidence and Distribution of Alfalfa-Infecting Viruses in China
Xin Wang#1 2, Jiantai Zhang#1 2^, Chenchen Liu1, Rongqun Wang1, Zhaoyan Tan1, Yuanhong Wang2, Beilei Wu1* and Xiliang Jiang1*
1Institute of Plant Protection, Chinese Academy of Agricultural Sciences, China
2Tianjin Agricultural University, China
Submission: March 08, 2021; Published: March 15, 2021
*Corresponding author: Beilei Wu, Xiliang Jiang, Institute of Plant Protection, Chinese Academy of Agricultural Sciences, Beijing, China
How to cite this article: Wang X, Zhang J, Liu C, Wang R, Tan Z, Wang Y, et al. Investigating Incidence and Distribution of Alfalfa- Infecting Viruses in China. Agri Res & Tech: Open Access J. 2021; 25 (4): 556314. DOI: 10.19080/ARTOAJ.2021.25.556275
Abstract
Alfalfa is a primary forage crop in China that occupies a wide planting area and exhibits a variety of beneficial traits such as being nutrient rich and producing high yields. As the planting area continues to expand, alfalfa pests and pathogens are becoming more prolific, especially viral pathogens which negatively impact alfalfa quality and yield significantly. From 2016 to 2019, 516 alfalfa samples exhibiting potential symptoms of viral infection were collected from several major production regions in China, including 13 provinces and 21 collection sites. RT-PCR using special conserved primers designed to detect several common alfalfa-infecting viruses was used to determine the incidence and distribution of these viruses throughout China. AMV, TMV, ToMV, CPMV, BYMV, CMV, YMV, and SMV were detected in alfalfa crops throughout China, with AMV being most prevalent with an incidence rate of 57.6%, while this is the first time that CMV and TMV have been found infecting alfalfa crops in China. This is also the first investigation of the distribution of various alfalfa viruses throughout China, providing theoretical and technical support for early warning and control of viral disease of alfalfa in China.
Keywords:Alfalfa; Alfalfa virus disease; Alfalfa mosaic virus (AMV)
Introduction
Alfalfa, the king of forage, being a high yielding and nutrient rich crop, is among the most important legume forage crops worldwide [1-2]. At present, the planting area of Alfalfa in China ranks first among all other forage crops [3-4], reaching over 10.3 million acres by the end of 2017, ranking as the second largest alfalfa planting in the world [5,6]. Alfalfa plays important ecological roles in improving the environment, optimizing the structure of agriculture and animal husbandry, and improving land resources and agricultural economic benefits. Alfalfa production areas in China are mainly found in northwestern, northeastern, southwestern and northern China, with some smaller plantings also occurring in eastern China, mainly in the Anhui and Jiangsu provinces. In recent years, with the continuous expansion of alfalfa planting areas, continuous cropping, and increasing prevalence of insect vectors, outbreaks of alfalfa viral diseases have developed rapidly. Viral diseases have become one of the most serious diseases presently affecting alfalfa, significantly impacting the yield and quality of this forage crop. More than 130 virus species have been reported to infect legumes, of which 31 can infect alfalfa [7]; these include Alfalfa mosaic virus (AMV) [8], Bean Leafroll Virus (BLRV) [9], Al falfa Dwarf Virus (ADV), Alfalfa Leaf Roll Virus (ALRV) [10], Cucumber Mosaic Virus (CMV), red clover vein mosaic virus (RCVMV), white clover mosaic virus (WCMV) [11], Alfalfa-associated nucleorhabdovirus (AaNV) [12], alfalfa virus S (AVS)[13-15], alfalfa virus F [16], Of these, AMV is the most prevalent virus causing disease in alfalfa, worldwide [8].
In 1997, Shang first began investigating virus diseases of alfalfa in China and found the incidence rate of AMV disease in the Ningxia province was over 50%, and that AMV was primarily distributed throughout northwestern and northern China [12]. In 2015, Wen and Nan detected AMV and tomato mosaic virus (ToMV) in alfalfa samples from the Zhangye region of Gansu province in China, which was the first report of ToMV infecting alfalfa [17]; In 2013, Xiang conducted virus pathogen detection on clover samples from Lanzhou, in the Gansu Province, and found the associated pathogens to be AMV and WCMV [18]. In 2016, Zhou et al. detected four viral pathogens in alfalfa in Lanzhou and Baiyin City of Gansu Province, which included AMV, WCMV, Bean Yellow Mo- saic Virus (BYMV) and Cowpea Mosaic Virus (CPMV) [19]. In 1990, Zhang et al. identified BYMV and CPMV in alfalfa from the Xinjiang province [20]. Later, Li identified Bean Leaf Roll Virus (BLRV) as a pathogen of Alfalfa in Urumqi, Changji and in Altay, Xinjiang province in 2019 [21]. Altogether, viruses documented to cause disease in alfalfa are AMV, WCMV, BYMV, CPMV, ToMV, and Soybean Mosaic Virus (SMV) in China. This area of research is mainly concentrated in Gansu and Ningxia in northwestern China. However, with the rapid expansion of the alfalfa industry in recent years, the various species of alfalfa-infecting viruses and their distribution in China are incompletely determined. To further characterize the types and distribution of these alfalfa viral diseases in China, we collected 516 samples exhibiting possible symptoms of virus infection from 21 collection sites in 13 major alfalfa growing provinces between 2016-2019, then identified and analyzed the predominant virus pathogens present.
Methods
Data collection and isolates get
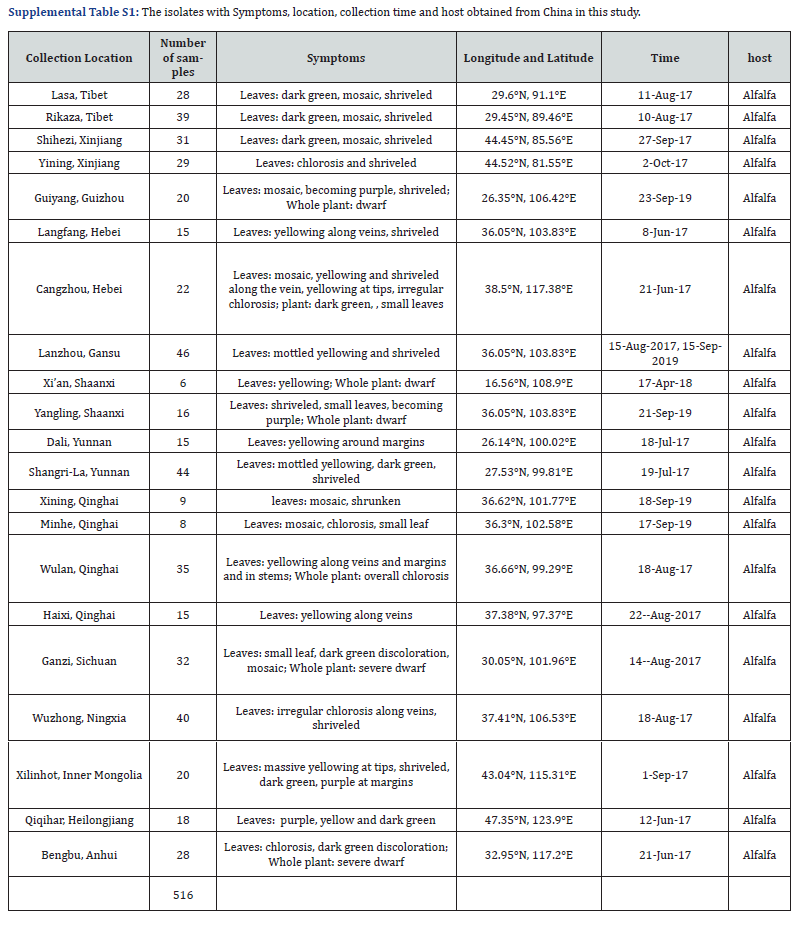
Virus isolates. The 516 alfalfa samples were collected from 21 locations of thirteen main alfalfa producing provinces and five climates regions in China from 2016-2019, the isolate names, their hosts, time and sites of collection and climates regions are (Supplemental Table S1).
RNA Extraction and RT-PCR
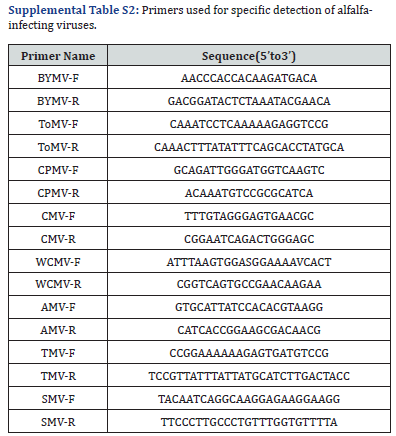
For the species of the eight viruses belong to the RNA virus, so RNA extracting of the samples were used as a template for reverse transcription in a reaction containing total RNA (50-5000 ng), gDNA Remover 1 μL, cp-F 1 μL cp-R 1 μL, 2×Reaction Mix 10 (1 μL). Enzyme Mix (1 μL) with 5 mM dNTP (each), RNase-free water to 20 μL, then incubated at 25 ℃ for 10 min, 42 ℃ for 45 min, and 85 ℃ for 5 ses. The resulting cDNA was used as template for PCR; the reaction mixture contained 2×Es Taq Mix with 5 mM dNTP (each) and 0.3 mL (5 U/mL) Ampli Taq RNA polymerase (Applied Biosystems, Foster City, CA, USA), add double distilled water to 20 μL. The thermocycling conditions are as follows: 95℃ 1.5min; 94 ℃ for 3 min; 35 cycles of 94 ℃ for 1 min, 57 ℃ for 1 min, 72 ℃ for 3 min; and 72 ℃ for 5 min. The expected RT-PCR products for each virus were obtained using specific primers targeting each of the eight viruses under detection (AMV, TMV, SMV, ToMV, WCMV, BYMV, CPMV and cucumber mosaic virus (CMV)) (supplemental Table S2) [17,20,22-23].
DNA sequencing
The PCR product segments were electrophoresed in 1.0% agarose gels and purified by the BioTeq PCR quick Gel Extraction Kit (BioTeq, USA). The purified fragments were cloned into the pMD18-T vector (Takara, Dalian, China) and used to transform Escherichia coli strain JM110. Insert sequences were determined for three clones at least for each fragment using either the ABI (ABI BigDye 3.1, Applied Biosystems) or Beckman Systems (GeXP with Genome Lab DTCS sequencing kit). Sequence data were assembled using BIOEDIT version 5.0.9 (Hall, 1999) [24]. The sequences obtained should be blasted in the NCBI (https://blast. ncbi.nlm.nih.gov/Blast.cgi) to identify the species of plant virus.
Results
Detection of alfalfa-infecting viruses
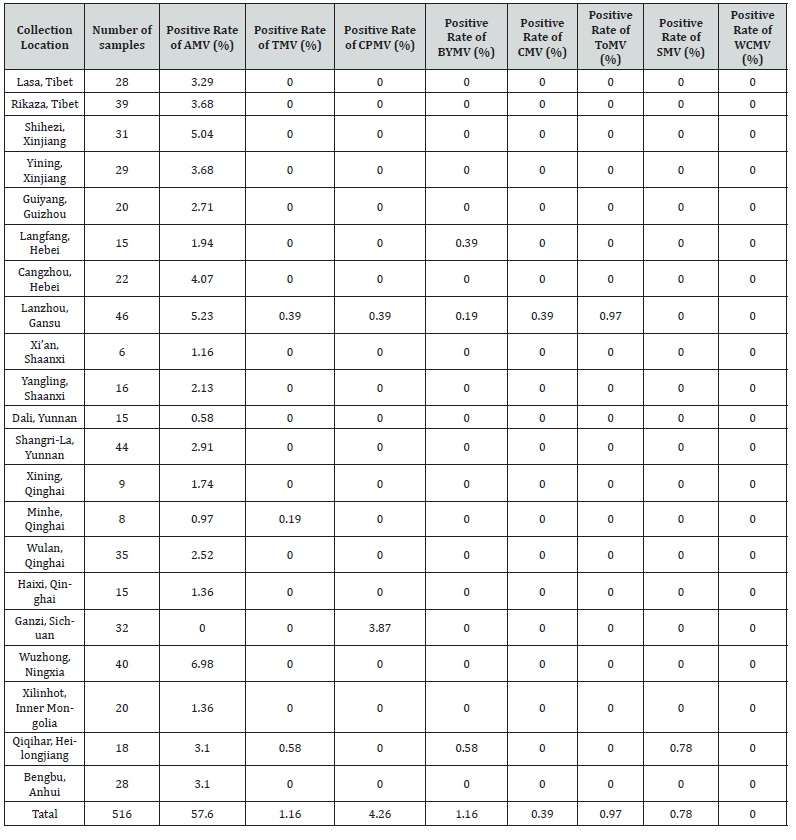
From 2016-2019, our research team collected alfalfa samples exhibiting typical symptoms of possible virus infection such as leaf tufted, dwarfing, dark green, mosaic, and Shrunken, from 21 collection sites in 13 provinces and autonomous regions (Heilongjiang, Inner Mongolia, Xinjiang, Qinghai, Tibet, Ningxia, Gansu, Yunnan, Hebei, Anhui, Shaanxi and Guizhou), from northwestern, southwestern, northeastern, northern and eastern China. Total 516 isolates were taken back to the laboratory and stored at -80℃. Information on collection and symptoms are shown in Supplemental Table S1. The results of RT-PCR detection revealed from each of the potentially infected samples showed that the viral pathogens associated with alfalfa mosaic disease in China were AMV (57.6%), TMV (1.16%), CPMV (4.26%), BYMV (0.39%), CMV (0.39%), ToMV (0.97%) and SMV (0.78%) (Tables 1 and 2; Supplemental Table S2). WCMV was not detected in any of the samples, and the dominant viral species detected was AMV.
Distribution of Alfalfa-Infecting Viruses in China
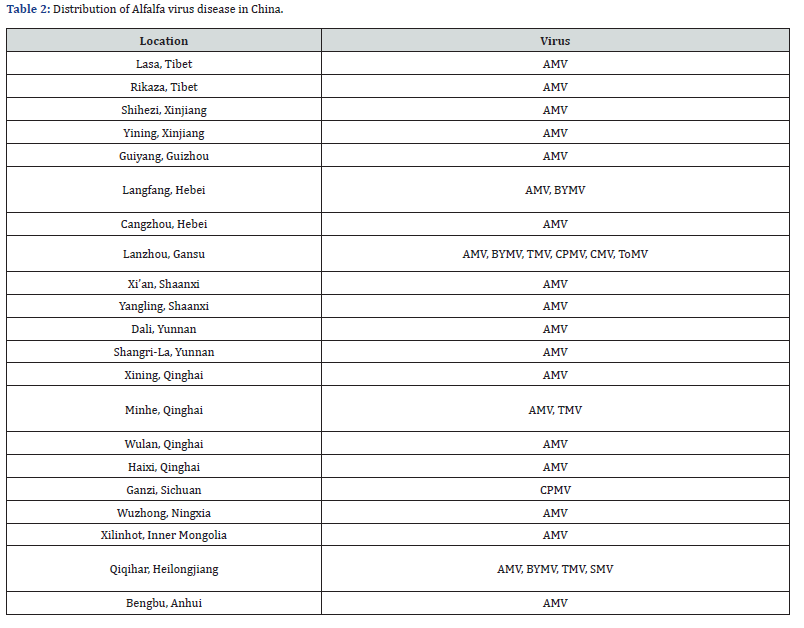
The results of testing the 516 symptomatic alfalfa revealed that the eight target viruses (AMV, TMV, CPMV, BYMV, CMV, ToMV, SMV and WCMV) were unevenly distributed across China; of these viruses AMV was the most widely distributed, being detected in 57.6% of the entire sample population. AMV was detected in all locations except Ganzi, Sichuan, however the incidence of rates of AMV in each location were variable (Table 1 & Table 2). Among the 8 viruses, AMV was the only virus detected in Lasa and Rikaza,
Tibet; Shihezi and Yining, Xinjiang; Guiyang, Guizhou; Xi’an, and Yangling, Shannxi; Dali and Xianggelila, Yunnan; Xining, Qinghai; Wuzhong, Ningxia; and Xilinhaote, Inner Mongulia. CPMV was only detected in Ganzi, Sichuan; none of the other seven viruses were detected in this location. The non-dominant viral species, such as TMV, CPMV, BYMV, CMV, ToMV, SMV and WCMV were sporadically distributed TMV was detected in Lanzhou from Gansu, Minhe from Qinghai and Qiqihar from Heilongjiang, with positive infection rates of 0.39%, 0.19% and 0.58%, respectively. CPMV was detected in Lanzhou, Gansu and Ganzi, Sichuan, with positive rates of 0.39% and 3.87%, respectively. TMV was detected in Langfang, Gansu and Qiqihar, Heilongjiang, with positive rates of 0.39% and 3.87%, respectively The positive rates of BYMV were 0.39% in Langfang, Hebei, 0.19% in Lanzhou, Gansu and 0.58% in Qiqihar, Heilongjiang respectively; CMV and ToMV were detected in Lanzhou, Gansu, with positive rates of 0.39% and 0.97% respectively. This is the first report of CMV being detected in alfalfa production regions in China. SMV was detected in Qiqihaer, Heilongjiang, with a positive rate of 0.78%; WCMV was not detected in any of the samples (Table 1 & Table 2).
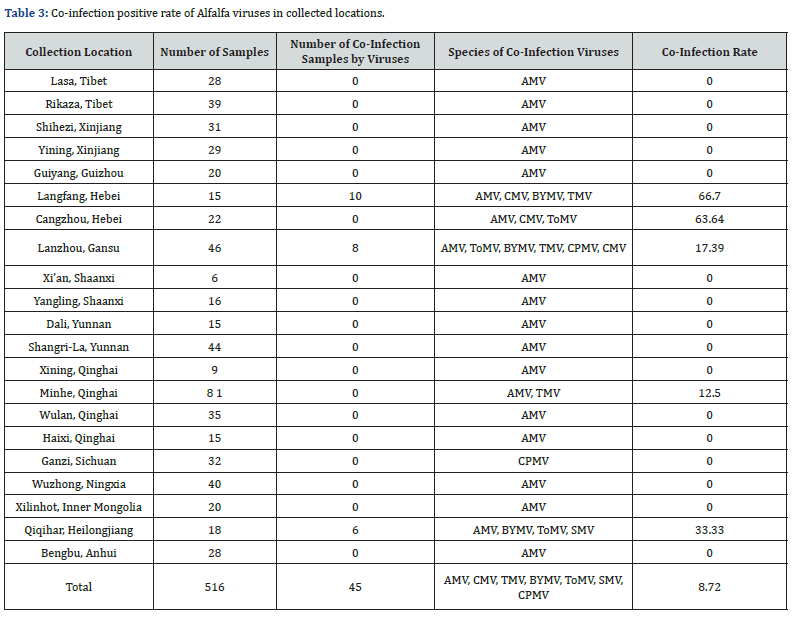
Incidence of co-infection by viruses
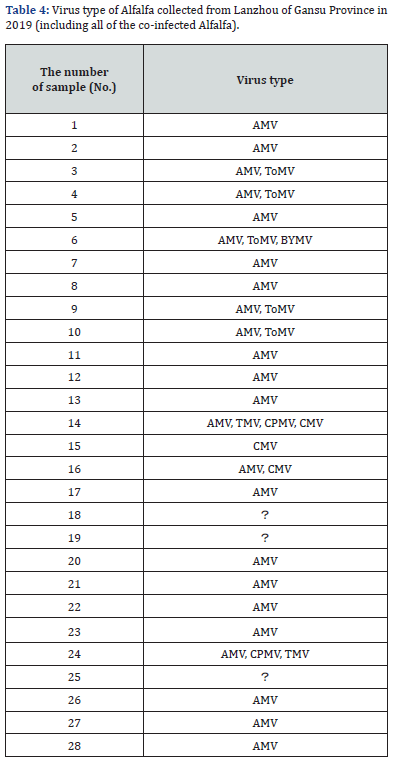
Co-infections by multiple viruses were detected several samples. For example, in the samples from Langfang, Hebei, the co-infection rate of AMV, CMV, BYMV and TMV was about 66.7%, while in Qiqihar, Heilongjiang, the co-infection rate of AMV, BYMV, ToMV and SMV was 33.33%. In Lanzhou, Gansu, co-infection by AMV, ToMV, BYMV, TMV, CPMV and CMV occurred at a rate of 60.87%. In Minhe, Qinghai, the co-infection rate was 12.5% for AMV and TMV. Lanzhou, Gansu exhibited the highest diversity of virus species, and co-infection was common (Table 2 & Table 3); 7 of 28 samples were found to be co-infected. Typical symptoms observed include plant being dwarf, leaves being tufted, dark green, yellow and/or purple discoloration, shriveled , and mosaic and so on.
The richness of symptoms of Alfalfa mosaic virus disease
Corresponding with the positive detection results, a diverse range of symptoms was also observed in the samples collected, including green, yellow, or purple discoloration at leaf margin, greening and yellowing along veins, veins yellowing, yellowing at leaf tips, shriveled leaves, dwarfing, deep green discoloration in leaves and stems, branches being erect, and incompletely expanded leaves (Figure 1).

Discussion
In this study, 516 symptomatic alfalfa samples collected from 21 sites from 13 provinces and autonomous regions of China between 2016 to 2019 were tested for 8 different species of viral pathogens [12, 17-20]. The results showed the associated viruses to be AMV, TMV, CPMV, BYMV, CMV, ToMV and SMV, among them, the positive detection rate of AMV was the highest, indicating AMV is the dominant virus species found in symptomatic alfalfa samples and is widely distributed. Previous studies focused primarily on regions of northwestern and northern China [10-12]. In this study we expanded the collection areas to include regions of northeastern, southwestern and eastern China, providing a more comprehensive understanding of the types and distribution of alfalfa-infecting viruses in alfalfa production regions throughout China. We found that AMV was distributed throughout northwestern, northeastern, southwestern and eastern China.
In this investigation, several nondominant virus species were detected, including BYMV in Langfang, Hebei, CPMV in Ganzi, Sichuan, CMV in Lanzhou, Gansu Province, TMV in Minhe, Qinghai, ToMV and SMV in Qiqihar, Heilongjiang. In 2016, it was found that CMV could infect alfalfa in Saudi Arabia [8], but in this study, CMV and TMV were each found infecting alfalfa alone and associated with mosaic symptoms, which was first report of these viruses infecting alfalfa in China. While the presence of WCMV was found in a study by Xiangdong’s study [11], this virus was not detected in our study. Trucco et al. [9] in 2016 and Li Kemei et al. [25] in 2019 both reported infection of Bean Leaf Roll Virus (BLRV) in alfalfa, however this pathogen was not detected in our study. Moreover, in the checking, there were some isolates not checked out identified 8 virus pathogens, which indicated perhaps the other plant virus should be existed in China, in 2020, AVS was check on the seed in China, AVS maybe far more widespread than previously thought [15]. These results suggest that in the future more collection sites and more virus pathogens should be chosen to be sampled and checked for avoiding missing detection and to achieve a more comprehensive understanding of incidence and distribution of viruses in alfalfa in China.
Additionally, it was found that virus co-infection was very common, as demonstrated in samples collected from Lanzhou, Gansu, Langfang, Hebei, Minhe, Qinghai and Qiqihar, Heilongjiang, indicating that there was a large diversity of alfalfa-infecting viruses in these locations. In previous studies, Xiang Dong found that the rate of co-infection by AMV and WCMV in alfalfa was 83.3% in Lanzhou, Gansu in 2013 [11]. In our study, co-infection by six pathogens in one sample was found in Lanzhou, Gansu, which reflected the kinds of pathogens in Lanzhou, Gansu were very rich. A wide variety of symptoms were observed in the collected alfalfa samples, which may relate to whether or not the varieties samples were resistant or susceptible, the soil in which they were grown, water stress and geographical environment. It is worth noting that the incidence of AMV has experienced an increasing trend across the years. From 2017 to 2018, 405 samples of AMV were collected, and 206 samples tested positive for AMV, giving a positive detection rate of 50.9%; in 2019, 111 alfalfa samples were collected, 91 of which were positive for AMV, giving a rate of 82.0%. This may suggest that alfalfa virus diseases are not being well and effective control measures may be lacking in China.
With the rapid development of animal husbandry, the production area of alfalfa will continue to expand [25] and the accumulation of virus vectors and alfalfa diseases, especially those caused by viruses, will likely increase in severity [26]. More alfalfa production sites should be samples and monitored for viral pathogens so as to provide theoretical and technical support for early warning and control of alfalfa mosaic diseases in China.
Declarations
Ethics approval and consent to participate
This article does not contain any studies with human participants performed by any of the authors.
Funding
This work was supported by the Natural Science Foundation of Beijing, China: Effect of environment on the evolution of fitness and virulence of Alfalfa mosaic virus (No. 31872414) granted to Beilei Wu; National key research and development program (2019YFD1002000-02) granted to Xiliang Jiang; Demonstration of comprehensive prevention and control technology of non-point source pollution in main vegetable producing areas of Huang Huai Hai (2018YFD0800401) granted to Beilei Wu.
Ethical approval
This article does not contain any studies with human participants performed by any of the authors.
Consent for publication
Informed consent was obtained from all participants included in the study.
Authors’ contributions
XW and JZ performed the experiments, CL, RW and ZT collected the samples, YW, BW and XJ designed the study. All authors read and approved the final manuscript.
References
- Liu YF, Wang ML, Shi ZZ (2014) Analysis of technical efficiency and technical progress contribution of alfalfa production. Pratacultural Science 31(10): 1990-1997.
- Lu XS (2013) Problems with the development of alfalfa industry in China. Chinese Journal of Grassland 35(5): 1.
- Zhang YF (2001) Alfalfa becoming a sunrise industry in the new century. China Flowers & Horticulture (4): 25-26.
- Han QF, Jia ZK, Wang JP (2005) The analysis of current situation and development prospect of alfalfa industry at home and abroad. Pratacultural Science 22(3): 22-25.
- Yang QC, Kang JM, Zhang TJ (2016) Distribution, breeding and utilization of alfalfa germplasm resources. Scientific Bulletin, 61(2): 261-270.
- (2017) National Animal Husbandry Station. Prataculture statistics in China. Beijing: China Agriculture Press. p.62-63.
- Stuteville DL, Erwin DC (1990) Compendium of Alfalfa Diseases, 2nd St Paul, MN, USA: APS Press. P. 51- 53.
- Massumi H, Maddahian M, Heydarnejad J, Hosseini Pour A, Farahmand A (2012) Incidence of viruses infecting alfalfa in the southeast and central regions of Iran. Agricultural Science and Technology 14(5): 1141-1148.
- Trucco V, Debrwuil S, Bejerman N, Lenardon S, Giolitti F (2016) Bean leafroll virus (BLRV) in Argentina: Molecular characterization and detection in alfalfa fields. European Journal of Plant Pathology146: 207-212.
- Bejerman N, Giolitti F, Trucco V, Soledad de Breuil, Claudia Nome, et al. (2015) Complete genome sequence and integrated protein localization and interaction map for alfalfa dwarf virus, which combines properties of both cytoplasmic and nuclear plant Rhabdoviruses. Virology 483: 275-283.
- AlShahwan IM, Abdalla OA, AlSaleh MA, Amer IM (2017) Detection of new viruses in alfalfa, weeds and cultivated plants growing adjacent to alfalfa fields in Saudi Arabia. Saudi J Biol Sci 24(6):1336-1343.
- Shang W J (1997) Alfalfa leaf disease survey and pathogen identification in Ningxia. Pratacultural Science 14 (1): 23-25
- Nemchinov LG, Grinstead SC, Mollov DS (2017) Alfalfa virus S, a new species in the family Alphaflexiviridae. PLoS ONE 12(5): e0178222.
- Nemchinov LG, Grinstead S, Irish B M, Shao J (2020) Identification and Complete Genome Sequencing of Alfalfa Virus S Diagnosed in Alfalfa Plants (Medicago sativa ) from Washington State, USA Plant Disease 104(12).
- Nemchinov LG, Franc ois S, Roumagnac P, Ogliastro M, Hammond RW, et al. (2018) Characterization of alfalfa virus F, a new member of the genus Marafivirus. PLoS ONE 13(9).
- Nemchinov LG, Grinstead S (2020) Identification of a Novel Isolate of Alfalfa virus S from China Suggests a Possible Role of Seed Contamination in the Distribution of the Virus. Plant Disease 104(12).
- Wen ZH, Nan ZB (2015) Detection of pathogenic organism in medicago sativain Zhangye. Acta Prataculturae Sinica 24(4): 121-126.
- Xiang D, Liang QL, Xu BL (2013) Symptoms and occurrence conditions of clover virus disease. Plant Protection 39(6): 130-136.
- Zhou QY, Liang QL, Han L (2016) Symptoms and pathogen detection of alfalfa virus disease. Pratacultural Science 10(7): 1297-1305.
- Zhang XL, Yin YQ, Li GY, Cui XM (1990) Bean yellow mosaic virus (BYMV)-identification of isolate of Xinjiang alfalfa, Virology Journal. 1990 (1): 88-96.
- Li KM, Aziguli Muhanmaiti, Ge RY (2019) Identification of a new virus on alfalfa in Xinjiang. Pratacultural Science, Pratacultural Science 36(9): 2319-2324.
- Jin LL, Zhang BH, Chen JS (2015) Complete nucleotide sequence and biological characteristics of a new Alfalfa mosaic virus isolate infecting Trifolium repens. Journal of Nuclear Agronomy 29(6): 1061-1067.
- Wen ZH, Yu L, Wang YQ (2018) Detection of viral pathogens in Medicago sativa seeds. Pratacultural Science 35(11): 2704- 2710.
- Hall TA (1999) Bio Edit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95-
- Liu YZ, Wang ML, Jiang NH (2018) Analysis of trade patterns and development trend of Chinese grass products. Pratacultural Science 35(11): 2765-2772.
- Jiang YZ, Zhang JB, Nan ZB (2016) Competitiveness analysis of alfalfa international trade in China. Pratacultural Science 33(2): 322-329.






























