In vitro Screening and Identification of P-Solubilizing Rhizobacteria Associated with Sorghum bicolor L.
Befekadu Teshome*, Endeshaw Abatneh and Enidegena Ayinalem
Ethiopian Biodiversity Institute, Microbial Biodiversity Directorate, Ethiopia
Submission: February 16, 2019, Published: February 28, 2019
*Corresponding author: Befekadu Teshome, Ethiopian Biodiversity Institute, Microbial Biodiversity Directorate, Addis Ababa, Ethiopia
How to cite this article: Befekadu Teshome, Endeshaw Abatneh, Enidegena Ayinalem. In vitro Screening and Identification of P-Solubilizing Rhizobacteria Associated with Sorghum bicolor L. Agri Res& Tech: Open Access J. 2019; 20(2): 556122. DOI: 10.19080/ARTOAJ.2019.20.556122
Abstract
In the present study, P-solubilizing rhizobacteria were screened and identified from Sorghum bicolor L root adhering soil and root which were collected from sorghum growing zones of Tigray, Ethiopia. A total of 94 bacteria were isolated from root washing solutions and surface sterilized roots. These isolates were evaluated for their ability to solubilize phosphates on Pikovskaya’s agar plates. The P-solubilizing bacterial isolates were identified by GEN III Biolog bacterial identification system. Fifty four of the 94 (57.5%) rhizobacterial isolates showed clearly visible haloes (>0.50cm) around their colonies on Pikovskaya’s agar after seven days of incubation. The solubilization index (SI) of the potential P-solubilizing rhizobacterial isolates differed significantly (p<0.05) and ranged from 0.5 to 4.83. Gram negative rhizobacteria dominated the identified P-solubilizing Rhizobacteria isolates and produced larger solubilization indices when compared with the Gram-positive isolates. Members of the phosphobacteria were dominated by the genus Pseudomonas (35.71%). Some of the isolates lost their capacity for phosphate solubilization on repeated sub-culturing. Overall, this finding indicated that there is a great number of rhizobacterial potential associated with Sorghum bicolor L which can be utilized for development of P-solubilizing bio-fertilizers.
Keywords: P-solubilizing rhizobacteria; Sorghum bicolor L.; Biolog bacterial identification
Abbreviations: PGPR: Plant Growth Promoting Rhizobacteria; PSM: Phosphate Solubilizing Microbes; PSB: Phosphate Solubilizing Bacteria; CD: Colony Diameter; SI: Solubilization Index; BUG: Biolog Universal Growth
Introduction
Plant growth promoting rhizobacteria (PGPR) flourish in the rhizosphere of plant, which may grow in, on, or around plant tissues and exert beneficial effects on plant development [1,2]. They possess the capacity to stimulate plant growth either directly or indirectly [3]. PGPR can affect plant growth by a wide range of mechanisms such as solubilization of inorganic phosphate, production of phyto-hormones, siderophores and organic acids, lowering of plant ethylene levels, N2 fixation and bio-control of plant diseases [4,5]. The use of such beneficial bacteria as bio-fertilisers and bio-control agents has currently attracted increased interest world-wide in attempts to achieve sustainability, particularly in agriculture, forestry and horticulture [5].
The number of PGPR that have been identified has seen a great increase in the last few years, mainly because of the role of the rhizosphere as an ecosystem has gained importance in the functioning of the biosphere. Various species of bacteria like Pseudomonas, Azospirillum, Azotobacter, Klebsiella, Enterobacter, Alcaligenes, Arthrobacter, Burkholderia, Bacillus and Serratia have been reported to enhance plant growth. There are several PGPR inoculants currently commercialized that seem to promote growth through at least one mechanism; suppression of plant disease (termed Bio-protectants), improved nutrient acquisition (Bio-fertilizers), or phyto-hormone production (Bio-stimulants) [2].
The use of PGPR offers an attractive way to replace chemical fertilizer, pesticides, and supplements; most of the isolates result in a significant increase in plant height, root length, and dry matter production of shoot and root of plants. The economic and ecological problems of today have re-invigorated the idea of using bio-fertilizers and bio-control agents in order to reduce the application of costly and environmentally-polluting agrochemicals to a minimum [6,7]. Agrochemicals (namely fertilizers and pesticides) have greatly influenced natural rhizosphere microbes in agro-systems [8]. Plant beneficial microbial bio-resources promise to replace or supplement many such destructive, high intensity practices and support ecofriendly crop production [6,7]. In particular, plant growth promoting rhizobacteria (PGPR) for the benefits of agriculture and ecosystem functions is gaining worldwide importance and acceptance [6,7,9,10].
Phosphorus is the second most important nutrient for plants, after nitrogen. It exists in soil as mineral salts or incorporated into organic compounds. Despite these phosphorus compounds being abundant in agricultural soils, the majority of them occur in an insoluble form. Plants require approximately 30μmol l-1 of phosphorus for maximum productivity, but only about 1μmol l-1 is available in many soils. Therefore, the unavailability of phosphorus in many soils has been recognized as a major growth limiting factor in agricultural and horticultural systems. This necessitates the application of soluble forms of phosphorus in the form of phosphate fertilizers, which in itself has constraints in that it too is rapidly immobilized (fixed) to insoluble forms upon its application in the soil due to its reaction with aluminum and iron minerals. The efficiency of applied phosphorus rarely exceeds 30% due to fixation in soil. It is also lost as a result of run-off and leaching, leaving as little as 10-20% available for plant utilization. Phosphate fertilizers are dependent on phosphorus derived from phosphate rock, which is a non-renewable resource and current global reserves may be depleted in 50-100 years. Therefore, exploring alternative forms of agriculture, where nutrient conservation is key, is of vital importance [11].
Several reports have indicated that different bacterial species, particularly rhizosphere colonizing bacteria, have the ability to liberate organic phosphates or to solubilize insoluble inorganic phosphate compounds such as tri-calcium phosphate, di-calcium phosphate, hydroxyapatite, and rock phosphate. These bacteria make available the soluble phosphates to the plants, and in return gain root borne carbon compounds, mainly sugars and organic acids, necessary for bacterial growth [12]. Current research suggests that the inoculation of crops with Phosphate Solubilizing Microbes (PSM) has the potential to reduce application rates of phosphate fertilizer by 50% without significantly reducing crop yield [13,14]. Phosphate Solubilizing Bacteria (PSB) may also be useful in the phyto-remediation of heavy metal impacted soil [15,16] or for bioleaching of rare Earth elements for mined ores [17].
Most soils in tropical and subtropical areas are predominantly acidic and extremely P-deficient due to their strong fixation of P as insoluble phosphates of iron and aluminum [9,12,18]. This leads to wide P deficiency which is particularly the case for the large parts of Ethiopian soils [19,20]. To alleviate P deficiency, chemical phosphate fertilizers are widely used. However, a large proportion of the soluble forms of P fertilizers is precipitated in insoluble form soon after application and becomes unavailable to plants [21]. This in turn leads to a need for excessive and repeated application of soluble P fertilizers, which in addition to the economic constraint can pose a serious threat to groundwater. These have been the major stresses that constrain the production of crops in the country.
Thus, in relation to this fact, P-solubilizing Rhizobacteria associated with cultivated Sorghum plant roots that displayed bio-fertilizer characteristics and have potential applications as native P-solubilizing bacterial bio-fertilizers were screened and identified in this study.
Materials and Methods
Description of sample collection areas
Sample collection was carried out in two major sorghum producing zones of Tigray region in Ethiopia. The sample collection site is shown in Figure 1. It comprises Central Tigray and South Tigray zones which are found in the northern part of Ethiopia. Based on the GPS data recorded during sample collection, the sample collection sites are located between 12O28.0988’- 13O19.9522’N and 38O53.1815’- 39O40.9870’E with an altitude range of 1342-1822m a.s.l.
Sample collection
A total of 93 sorghum roots with adhering soil samples were collected in sterile plastic bags. Samples were collected based on altitude differences of sorghum plant growing areas, cultivar types and plant age group. At each sampling site, plant roots with adhering soil (approximately 50g) were uprooted and placed into a sterile plastic bag. Care was taken to keep rhizosphere soil intact around the root. The collected samples were kept in ice-box and transported to Ethiopian Biodiversity Institute Microbiology Laboratory. All samples were kept at 4 ˚C until use [22-24].
Isolation of Rhizobacteria
Sorghum roots with adhering soils were merged into 17 composite samples separately based on similarity of cultivar type, plant and age group. The root adhering soils were dislodged from the roots using sterile distilled water by shaking at 250rpm for 20 minute and the root washing solutions were used for the isolation of rhizoplane bacteria [25]. For the isolation of bacterial endophytes, merged and washed roots were surface sterilized in 99% ethanol for 1min, 3% NaOCl for 6 minutes, and 99% ethanol for 30 seconds and followed by rinsing with sterile distilled water for 6 times [23]. Before homogenization, a root fragment was imprinted on nutrient agar to serve as a sterility check. Roots were homogenized and macerated with a sterile mortar and pestle [26]. The root washing solutions and homogenized roots were serially diluted (10-2 to 10-4) aseptically for inoculation. 0.1ml inoculums of the prepared samples were spread onto Nutrient agar plates and incubated at 30+2 ˚C for 48h [27,28]. Bacterial colonies with distinct and peculiar morphologies were selected and re-streaked to obtain pure colonies [24].
In vitro screening of bacteria for P-solubilization potential
Phosphate solubilization ability of the isolated bacteria was determined on Pikovskaya’s agar. The isolates were spotted onto Pikovskaya’s agar and incubated for 7 days at 30 ± 2 ˚C. The presence of halo zone around the bacterial colony was considered as indicator for positive phosphate solubilization. Further, the solubilization index (SI) of the isolates was determined by measuring the halo zone of clearance (HD) in the Pikovskaya’s agar plates and the colony diameter (CD) [29]. SI was calculated with the formula: SI = (CD+HD)/CD. Three replicate plates were used for each isolate [30].
Identification of P-solubilizing rhizobacteria
Preliminary identification of P-solubilizing Rhizobacteria isolates were performed by examination for cell morphology using optical microscopy, Gram staining, and colony morphology [27,24]. Biochemical identification including the carbohydrate fermentation patterns and chemical sensitivity tests were determined using GEN III Biolog bacterial identification system kit. The Biolog GEN III Micro Plate analyzes a microorganism in 94 phenotypic tests: 71 carbon source utilization assays and 23 chemical sensitivity assays. The test panel provides a “Phenotypic Finger print” of the microorganism that can be used to identify it at the species level. The plates contained 96 wells, with a dehydrated panel of necessary nutrient medium (a carbon source), biochemical and tetrazolium violet. Tetrazolium violet is a purple formazan, a redox dye that turns purple when reduced, indicating use of the carbon source provided or resistance to inhibitory chemicals. Each plate contained a positive and negative control well. Pure culture of bacteria isolates was grown on Biolog BUG agar plates at 30 ± 2 ˚C for 20-24 hours. Single colonies were swabbed and suspended in inoculating fluid A. Cell suspensions (100μl) adjusted at 90-98% transmittance was pipetted into 96 well Biolog Micro-plates for carbon utilization and chemical test. Panels were incubated at 30 ± 2 ˚C for 20-24 hours. The microplates were inserted into the Omnilog automatic system and the identification process was carried out using GEN III Biolog- Omnilog identification system software [31].
Data analysis
Data were analyzed using SPSS software version 20 (SPSS Inc., Chicago, IL, USA). Coefficient of variation was calculated for the significances of differences within samples and ANOVA was employed for significances of differences between mean counts of microbial groups. DIVA_GIS 7.5.0 was used for mapping study areas.
Results and Discussion
In vitro screening of P-solubilizing rhizobacteria
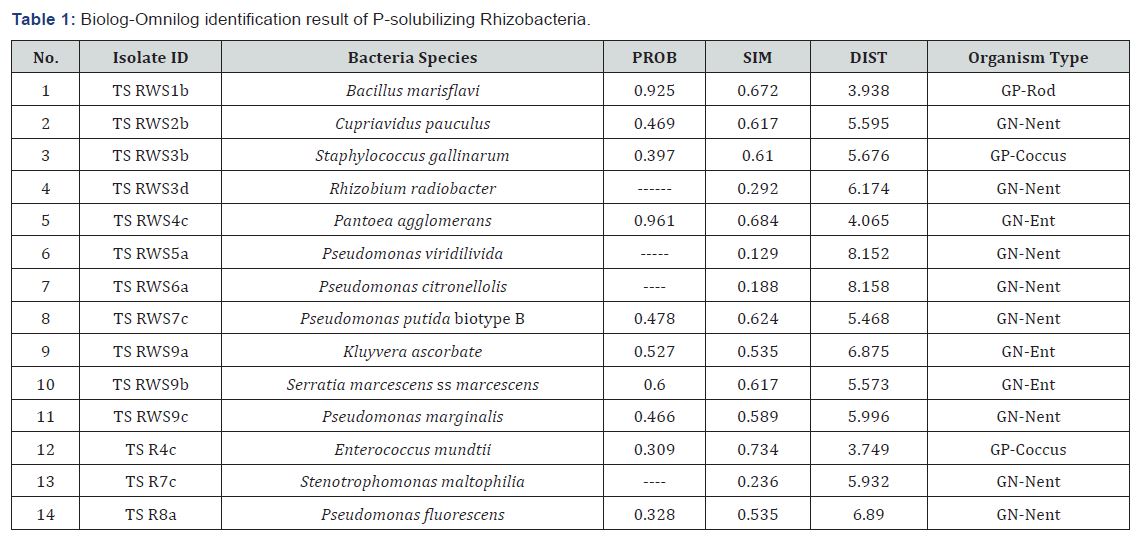
Ninety-four bacteria were isolated from root washing solutions and surface sterilized roots on nutrient agar. Fifty-one bacteria were isolated from sorghum root washing solutions which were prepared from the root adhering soils and the rest 43 were endophyte bacteria isolated from sorghum roots. These 94 bacterial isolates were evaluated for their ability to solubilize phosphates on Pikovskaya’s agar plates (Table 1). Fifty four of the 94 (57.5%) rhizobacterial isolates showed clearly visible haloes (>0.50cm) around their colonies on Pikovskaya’s agar after seven days of incubation. The solubilization index (SI) of the potential P-solubilising rhizobacterial isolates differed significantly (p<0.05) and ranged from 0.5 to 4.83. Bacterial strain TS RWS7b produced the largest zone of solubilisation, followed by TS RWS 1b.
Identification of P-solubilizing rhizobacteria
Based on colony morphology shown on nutrient agar and Biolog Universal Growth (BUG) agar, and Gram staining similarity, the 54 P-solubilizing Rhizobacteria screened from root washing solutions and sorghum roots were clustered into 17 representative isolate morphological groups. Inoculums of the 17 clustered representative isolates were prepared and transferred into GEN III Micro-plates. After 24 hours of incubation at 30+2 ˚C, the microplates were subjected to Biolog-Omnilog bacterial identification system test. Fourteen of the 17 clustered representative P-solubilizing Rhizobacteria isolates were identified (Table 1).
Eleven of the 14 identified P-solubilizing Rhizobacteria were isolated from root washing solution and the rest 3 were isolated from sorghum root. Gram negative rhizobacteria dominated the system accounting for 78.57% (11/14) of the identified P-solubilizing Rhizobacteria isolates (Table 1,2). Previous observation showed that the rhizosphere of many agriculturally important plants favors more Gram negative rhizobacteria than the Gram positives [4,32]. The largest solubilization index was also produced by Gram negative isolate when compared with Gram-positive isolate. Some of the isolates lost their capacity for phosphate solubilization on repeated sub-culturing as previously reported in many other studies [33,34].
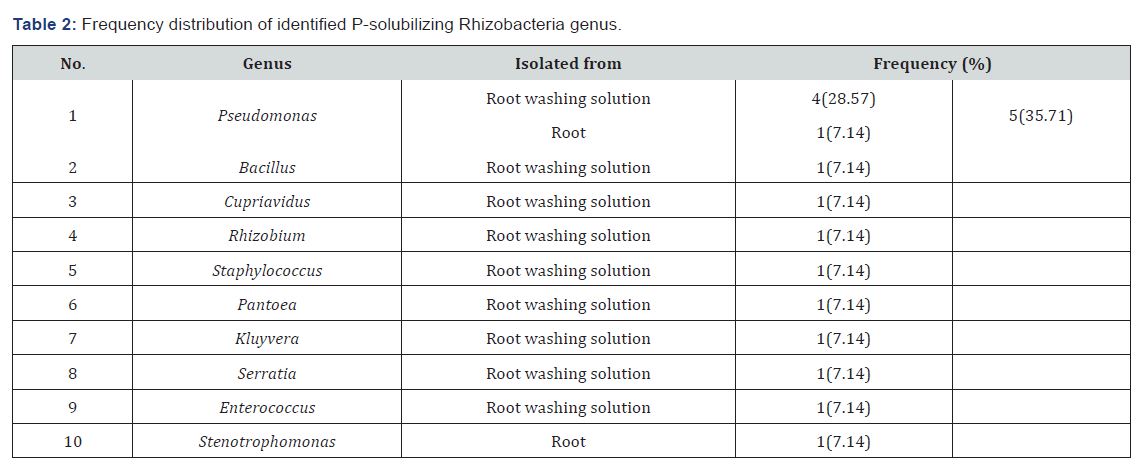
Ten different genera of Rhizobacteria were identified. Most of them were isolated from root washing solutions. Eight of the 10 identified Rhizobacteria genera were isolated only from sorghum root washing solutions. But, only Stenotrophomonas species was isolated from root. Meanwhile, Pseudomonas species was isolated from both root washing solutions and root. Members of the phosphobacteria were dominated by the genus Pseudomonas (35.71%) (Table 2). Pseudomonas are the most dominant genera commonly reported in many plant studies [35].
Conclusion
This study showed that there are a large proportion of P-solubilizing rhizoplane and endophytes rhizobacteria associated with Sorghum bicolor L. Pseudomonas is the most dominant rhizobacteria both in the root adhering soil and roots of sorghum. In general, Gram negative bacteria were not only more predominant than Gram positive bacteria but also, they produced the largest solubilization index. This finding indicated that there is a great number of rhizobacterial potential associated with Sorghum bicolor L. which can be utilized for development of P-solubilizing bio-fertilizers.
>
Acknowledgement
We would like to extend our gratitude to agricultural experts working at Central Tigray and Southern Tigray zones and district (woreda) offices, and extension workers at each administration units (kebeles) where sample collection was undertaken in Northern Ethiopia.
Conflicts of interest
Authors did not declare any conflict of interest.
References
- Akhtar A, Hisamuddin, Robab MI, Sharf A, Sharf R (2012) Plant growth promoting Rhizobacteria: An overview. J Nat Prod Plant Resour 2(1): 19-31.
- Saharan BS, Nehra V (2011) Plant Growth Promoting Rhizobacteria: A Critical Review. Life Sciences and Medicine Research 2011: LSMR-21.
- Kloepper JW, Schroth MN (1980) Plant Growth Promoting and Plant Growth Under Gnotobiotic Conditions. Ecology and Epidemiology 71(6): 642-644.
- Muleta D (2009) Phosphate solubilizing microbes: potentials and success in greenhouse and field applications. In: Khan MS, Zaidi A (eds.), Phosphate Solubilizing Microbes for Crop Improvement. Nova Science Publishers Inc., New York, USA, pp: 281-308.
- Datta M, Palit R, Sengupta C, Kumar M, Banerjee S (2011) Plant growth promoting rhizobacteria enhance growth and yield of Chilli (Capsicum annuum L.) under field conditions. Australian Journal of Crop Science 5(5): 531-536.
- Hart MM, Trevors JT (2005) Microbe management: Application of mycorrhyzal fungi in sustainable agriculture. Frontiers in Ecology & the Environment 3(10): 533-539.
- Rodríguez H, Fraga R, Gonzalez T, Bashan Y (2006) Genetics of phosphate solubilization and its potential applications for improving plant growth-promoting bacteria. Plant & Soil 287: 15-21.
- Matson PA, Parton WJ, Power AG, Swift MJ (1997) Agricultural intensification and ecosystem properties. Science 277(5325): 504-509.
- Vessey JK (2003) Plant growth promoting rhizobacteria as biofertilizers. Plant & Soil 255(2): 571-586.
- Lucy M, Reed E, Glick BR (2004) Applications of free-living plant growth-promoting rhizobacteria. Antonie van Leeuwenhoek 86(1): 1-25.
- Oteino N, Lally RD, Kiwanuka S, Lioyd Aryan D, Germaine KJ, et al. (2015) Plant growth promotion induced by phosphate solubilizing endophytic Pseudomonas isolates. Front Microbiol 6: 745.
-
- Khan RU, Durrani FR, Chand N, Anwar H (2010) Influence of feed supplementation with Cannabis sativa on quality of broilers carcass. Pakistan Vet J 30(1): 34-38.
- Ahmad R, Jilani G, Arshad M, Zahir ZA, Khalid A (2007) Bio-conversion of organic wastes for their recycling in agriculture: an overview of perspectives and prospects. Annals of Microbiology 57(4): 471-479.
- Yazdani M, Ali Bahmanyar M, Pirdashti H, Ali Esmaili M (2009) Effect of Phosphate Solubilization Microorganisms (PSM) and Plant Growth Promoting Rhizobacteria (PGPR) on Yield and Yield Components of Corn (Zea mays L.). International Scholarly and Scientific Research & Innovation 3(1): 50-52.
- Ahmad N, Ahmad Zada, Muhammad Junaid, Akhtar Ali (2015) Bridging the Yield Gap in Rice Production by Using Leaf Color Chart for Nitrogen Management. Journal of Botany 2016: 6.
- Monica Sen, Harshada J (2017) Characterization of Phosphate Solubilizing Bacteria Isolated from Mine Tailings of Zawar Mines, Udaipur, India. Int J Curr Microbiol App Sci 6(8): 1-9.
- Shin K, Fujii K, Yumoto S, Ishimoto M, Shiraiwa T, et al. (2015) Seed yield and its components of indeterminate and determinate lines in recombinant inbred lines of soybean. Breed Sci 65(2): 154-160.
- Gyaneshwar P, James EK, Pallavolu MR, Ladha JK (2002) Herbaspirillum colonization increases growth and nitrogen accumulation in aluminium‐ tolerant rice varieties. New Physiologist 154(1): 131-145.
- Beyene D (1988) Biological Nitrogen Fixation Research on Grain Legumés in Ethiopia-An Overview. Nitrogen Fixation by Legumes in Mediterranean Agriculture, p. 73-78.
- Mamo M, Rosen CJ, Halbach TR, Moncrief JF (1988) Corn Yield and Nitrogen Uptake in Sandy Soils Amended with Municipal Solid Waste Compost. Journal of Production Agriculture 11(4): 469-475.
- Mahantesh P, Patil CS (2011) Isolation and Biochemical Characterization of Phosphate Solubilizing Microbes. International Journal of Microbiology Research 3(1): 67-70.
- Thanh DTN, Diep CN (2014) Isolation and identification of rhizospheric bacteria in Acrisols of maize (Zea mays L.) in the eastern of South Vietnam. American Journal of Life Sciences 2(2): 82-89.
- Monk J, Gerard E, Young S, Widdup K, O’Callaghan M (2009) Isolation and identification of plant growth-promoting bacteria. Proceedings of the New Zealand Grassland Association 7: 211-216.
- Gupta S, Meena MK and Datta S (2014) Isolation, characterization of plant growth promoting bacteria from the plant Chlorophytum borivilianum and in-vitro screening for activity of nitrogen fixation, phospthate solubilization and IAA production. Int J Curr Microbiol App Sci 3(7): 1082-1090.
- Costa R, Götz M, Mrotzek N, Lottmann J, Berg G, et al. (2005) Effects of site and plant species on rhizosphere community structure as revealed by molecular analysis of microbial guilds. FEMS Microbiol Ecol 56(2): 236-249.
- Jimenez-Salgado T, Fuentes-Ramirez LE, Tapia-Hernandez A, Mascarua- Esparza MA, Martinez-Romero E, et al. (1997) Coffea arabica L., a new host plant for Acetobacter diazotrophicus, and isolation of nitrogen-fixing acetobacteria. Appl Environ Microbiol 63(9): 3676- 3683.
- Yamaoka-Yano DM, Amorim Epr, Valarini PJ, Melo IS, Kosako Y (1988) Characterization of rhizobacteria from Citrus rhizosphere to control Phytophthora nicotinae var. parasitica and P. citrophthora. Nature Biotechnology 6(3): 282-286.
- Raj DP, Cherian N (2013) Isolation and characterization of rhizobacteria from forest soils for in vitro study on crop growth enhancement. INT J CURR SCIE 9: E19-E23.
- Pikovskaya RI (1948) Mobilization of phosphorus in soil connection with the vital activity of some microbial species. Microbiologiya 17: 362-370.
- Edi-Premono M, Moawad MA, Vleck PLG (1996) Effect of phosphate solubilizing Pseudomonas putida on the growth of maize and its survival in the rhizosphere. Indonesian Journal of Crop Science 11(1): 13- 23.
- Biolog’s Microbial Identification Systems users guide (2008) GEN III Micro Plate Instructions for Use. Hayward, USA, p. 1-8.
- https://portal.findresearcher.sdu.dk/
- Rashid M, Khalil S, Ayub N, Alam S, Latif F (2004) Organic acid production and phosphate solubilization by phosphate solubilizing microorganisms (PSM) under in vitro conditions. Pakistan Journal of Biological Sciences 7(2): 187-196.
- Muleta D, Assefa F, Borjesson E, Granhall U (2013) Phosphate-solubilising rhizobacteria associated with Coffea arabica L. in natural coffee forests of southwestern Ethiopia. Journal of the Saudi Society of Agricultural Sciences 12(1): 73-84.
- Hallmann J, Berg G (2006) Microbial root endophytes: Endophytic bacteria. Springer, p: 9.






























