Leaf Protein Patterns: a Windows for Assessment of Genetic Diversity of Melissa officinalis Accessions
Daryush Talei1* and Amir Mohammad Naji2
1Medicinal Plants Research Center, Shahed University, Tehran, Iran
2Department of Biotechnology, Agriculture Faculty,Shahed University, Tehran, Iran
Submission: February 08, 2018; Published: March 29, 2018
*Corresponding author: Daryush Talei, Medicinal Plants Research Center, Shahed University, Tehran 3319118651, Iran, Tel: +98-2121514093 Email: d.talei1348@gmail.com
How to cite this article: Daryush T, Amir M N. Leaf Protein Patterns: a Windows for Assessment of Genetic Diversity of Melissa officinalis Accessions. Agri Res & Tech: Open Access J. 2018; 15(1): 555940. DOI:10.19080/ARTOAJ.2018.15.555940.
Abstract
Lemon balm (Melissa officinalis L.) is an important medicinal plant in the family of Lamiaceae. The purpose of the current study was to test the applicability of leaf protein patterns for assessment of genetic diversity of 25 accessions of Melissa officinalis. The leaf protein of 45 days seedling was extracted using HEPES/KOH buffer and separated by SDS-PAGE technique. The cluster analysis was performed based on protein bands using weighted pair group method arithmetic averages (WPGMA) of DARwin 5 software and PCoA. The results indicated a total of 18 bands ranged from 10 to 150kDa of which 59.09 percentage were polymorphic. The maximum number of the bands (16) was observed in Germany accession, while Italy and Mashhad accessions had the minimum number of bands (10). The cluster analysis of 25 accessions based on the protein data produced five clusters. The first cluster involved 4 accessions, the second cluster contained six accessions, the third cluster contained three accessions, the fourth cluster contained five accessions and fifth cluster contained seven accessions. Overall, the outcomes of the present study indicated a high genetic diversity among the M. officinalis accessions. As a result of this investigation, protein markers would be a useful tool for assessment of genetic diversity among Lemon balm genotypes, and the findings suggest that the plants belonging to different clusters can be used for intraspecific hybridization to generate useful recombinants in the segregating generations for breeding of M. officinalis.
Keywords: Genetic diversity; Melissa officinalis; Protein pattern; SDS-PAGE
Introduction
It has been suggested that the lemon balm (Melissa officinalis L.) is one of the most important medicinal herbs in the family Lamiaceae. Numerous studies have reported this species includes three subspecies: M. officinalis ssp. altissima, M. officinalis ssp. inodora and M. officinalis ssp. officinalis, however, only ssp. officinalis has commercial and medicinal value [1]. Reviewed from previous literatures the plant mainly grown in the Mediterranean regions such as Turkey, Southern Europe and Northern Africa, and Northern Iran [2,3]. In Iran, the plant has a broad distribution in Northern, Northeast (Golestan forest, Mazandaran, Haraz Valley), and West region of Kermanshah and Rijab, Tehran (Tochal altitudes) and between Qazvin and Karaj [4].
It is a well-documented fact the plant essential oil contains three main compounds namely, protocatechuic acid, caffeic acid and rosmarinic acid [5]. Among these, phytochemicals rosmarinic acid possesses the highest bioactivity in treating the hardly curable diseases. The herb exhibited a wide scope of pharmaceutical properties such as anti-oxidant, anti-allergic, sedative, carminative, anti-spasmodic, anti-microbial, antifungus, anti-inflammatory, hypolipidemic, and anti-oxidant [5,6]. Many authors have suggested that for the total content of oil in M. officinalis is about 0.02-0.3% per dry weight, which is relatively low as compared with other members of the Lamiaceae [7]. For this reason, the production cost of essential oil is too high in the market [8].
Several studies have argued the success of any breeding or genetic conservation program is dependent on an understanding of the amount and distribution of the genetic diversity available in the gene pool. Traditionally, a combination of morphological and agronomic traits has been used to measure genetic diversity. Genetic diversity is a fundamental issue estimated using different methods such as morphological traits, protein and DNA markers [9,10]. Morphological traits have a number of limitations, including low polymorphism, low heritability, late expression, and may be controlled by epistasis and pleiotropic gene effects Nakamura [11], while molecular markers such as protein markers reflect the genotype more directly, independent of environmental influences [12,13]. Protein markers are useful tools to identify cultivar, registration of new varieties and classification of crop species to study genetic diversity, thereby improving the efficiency of plant breeding programs [14-16]. Among biochemical techniques, Sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) due to its simplicity, minimum cost in time and labor, and effectiveness is commonly used in the studies of plants genetic diversity [17-20].
With respect to genetic diversity, several polymorphic proteins have been reported in the genus Mentha Hassan et al. [21], Ocimum species Mustafa et al. [22], legume Boulter et al. [23], wheat El-Bakatoushi [24] and Brassicaceae [25]. An enormous lack of information related to the genetic diversity of M. officinalis is tangible. Therefore, the objective of present study was investigation on genetic diversity of M. officinalis accessions from different countries using protein analysis, which might increase efficiency of conservation of germplasm in order to utilize in breeding programs.
Material and Methods
Plant materials
A total of 25 accessions of M. officinalis were collected from different countries (England, Germany, Iran, Italy, Japan and Majarestan) (Table 1). The seeds were germinated into the growth chamber at 28 °C. After 48h, the germinated seeds were transferred into the pots with mixture of top soil and sand (2:1 v/v) in the greenhouse conditions with a temperature of 25 °C and irrigation every 48h at Medicinal Plants Research Center, Shahed University, Tehran. The leaves were harvested from the 45 days seedling to extract the total protein.
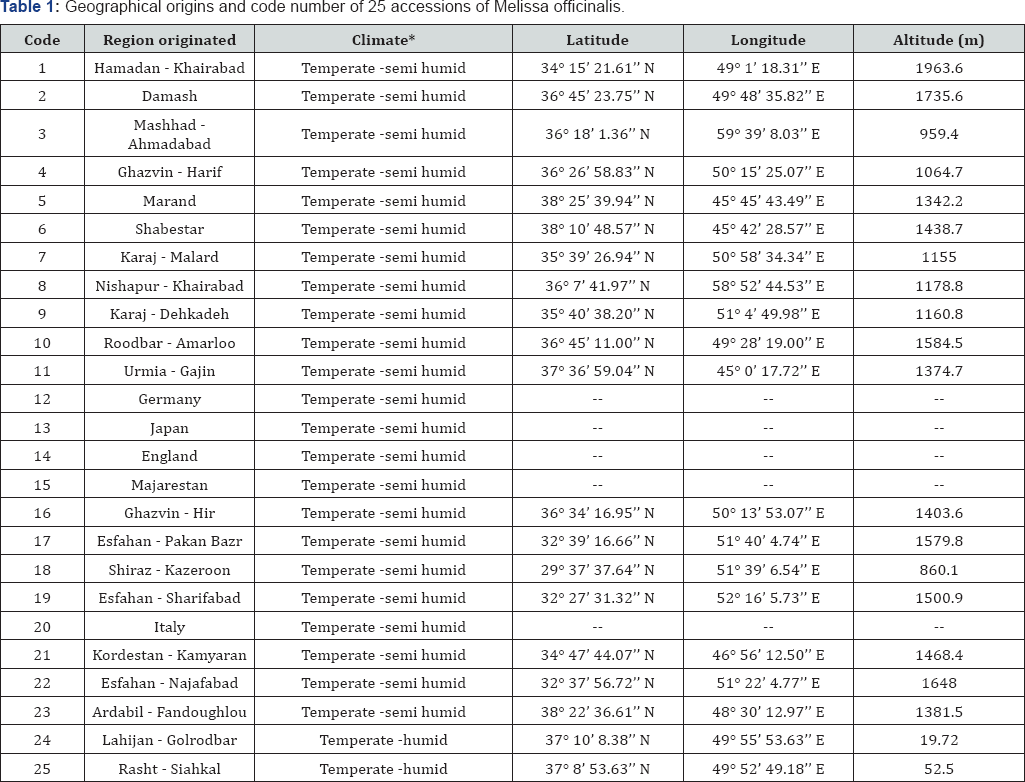
*Annual mean temperature in temperate climate is 10 °C - 15 °C. Annual mean rainfall in humid and semi humid climates is 600-1400mm and 300-600mm, respectively.
Leaf protein extraction
One gram of fresh leaf tissue from three samples of each accession was grounded in liquid nitrogen using pre-chilled mortar and pestle to obtain a fine powder. One g of the powder was homogenized and mixed with 2.5mL of the HEPES/KOH buffers according to the method of Talei et al. [26].
Protein separation
One dimensional SDS-PAGE was performed according to the method of Laemmli [27]. Twenty |ig of protein from each sample were solubilised with 4x SDS/sample buffer (0.12 5M Tris base (pH 6.8), 80% (w/v) glycerol, 10% (w/v) SDS, 4% (v/v) 2-mercaptoethanol and 0.05% (w/v) bromophenol blue) and heated at 100 °C for 3 min. The protein samples were run on 12% separating gel at 90 V for 90 min using a Bio-Rad, Mini Protein Tetra Cell electrophoresis system. The electrophoresis gels were stained with Coommassie Blue (0.25% Coommassie Brilliant Blue G-250, 40%(v/v) methanol and 7%(v/v) acetic acid) for one h and destained with 40% (v/v) methanol and 7% (v/v) acetic acid. The gels scanning were done using a Gel Documentation (Bio-Rad, XR-S Plus).
Statistical analysis
The SPSS software version 22 was used for variance analyses and the NTSYS-pc version 2.1 software was used to calculate the Jaccard coefficient and cophenetic coefficient [28]. The cluster analysis was performed based on protein bands using weighted pair group method arithmetic averages (WPGMA) of DARwin 5 software and PCoA [29].
Results
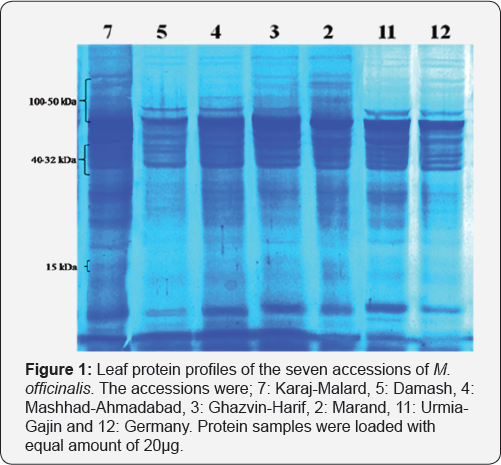
In practice, the seed protein analysis of the 25 accessions of Melissa officinalis unveiled the presence of 16 different types of proteins ranged from 10 to 100kDa, yet, only six of them with low molecular weight were found polymorphic (Figure 1). The results indicated that most of the bands were similar in all accessions. Potentially, these protein bands can also serve as useful markers to hybridization and breeding programs in future studies.
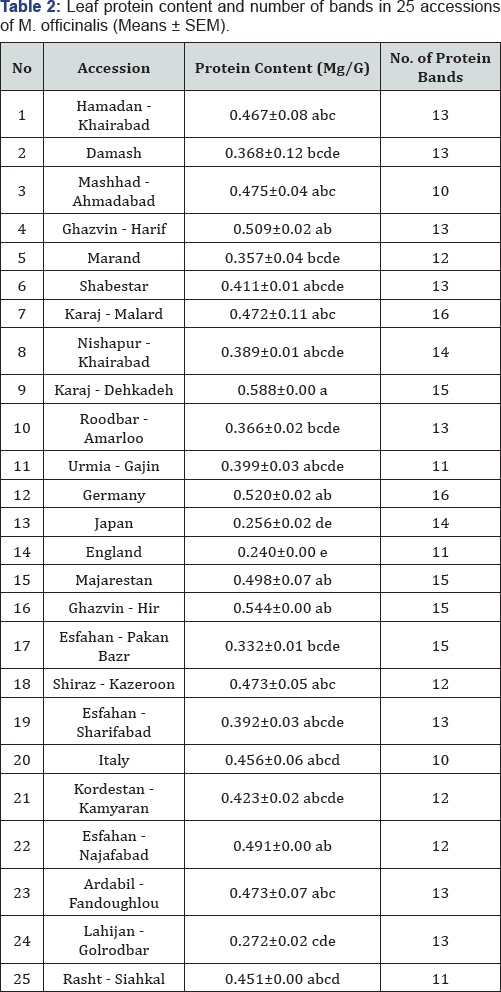
Different letters indicate significant difference between the values of pairs of accession in the same column at P<0.01 according Duncan's multiple comparisons.
The analysis of variance indicated that there were significant differences among the accessions in terms of protein content (P<0.01). The highest protein content (0.588mg/g) and the lowest protein content (0.240mg/g) were observed in Karaj - Dehkadeh and England accessions, respectively (Table 2).
Jaccard's similarity coefficients were computed for all Germany and Malard accessions with a value of 1.0 and the lowest possible pairs of electropherograms among the 25 accessions one's was obtained between Italy and N. Khorasan accessions (Table 3). The highest genetic similarity was observed between with a value of 0.5.

1: Hamadan - Khairabad; 2: Damash; 3: Mashhad - Ahmadabad; 4: Ghazvin - Harif; 5: Marand; 6: Shabestar; 7: Karaj - Malard; 8: Nishapur - Khairabad; 9: Karaj - Dehkadeh; 10: Roodbar - Amarloo; 11: Urmia - Gajin; 12: Germany; 13: Japan; 14: England; 15: Majarestan; 16: Ghazvin - Hir; 17: Esfahan - Pakan Bazr; 18: Shiraz - Kazeroon; 19: Esfahan - Sharifabad; 20: Italy,; 21: Kordestan - Kamyaran; 22: Esfahan - Najafabad; 23: Ardabil - Fandoughlou; 24: Lahijan - Golrodbar; 25: Rasht - Siahkal
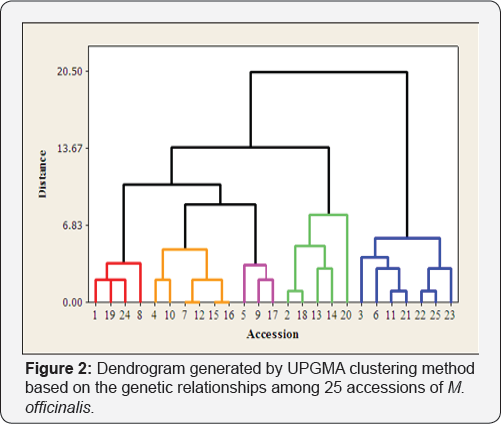
The UPGMA cluster analysis of 25 accessions of M. officinalis based on the protein profiles produced five clusters (Figure 2). The first cluster (red cluster) involved 4 accessions including, Hamadan-Khairabad, Nishapur-Khairabad, Esfahan-Sharifabad and Lahijan-Golrodbar. The second cluster (orange cluster) contained six accessions (Ghazvin-Harif, Karaj-Malard, Roodbar- Amarloo, Germany, Majarestan and Ghazvin-Hir). Interestingly, Germany, Majarestan and some Iranian accessions with high geographical distance were happened in the first cluster. The third cluster (pink cluster) contained three accessions (Marand, Karaj-Dehkadeh and Esfahan-Pakan Bazr). The fourth cluster contained five accessions (Damash, Japan, England, Shiraz- Kazeroon and Italy) and fifth cluster contained seven accessions (Mashhad-Ahmadabad, Shabestar, Urmia-Gajin, Kordestan- Kamyaran, Esfahan-Najafabad, Ardabil-Fandoughlou and Rasht- Siahkal) (Figure 2). The lowest distance (0.00) was happened between Malard and Germany accessions, while the highest distance (6.49) was happened between Karaj-1 and Shiraz accessions (Blue cluster).
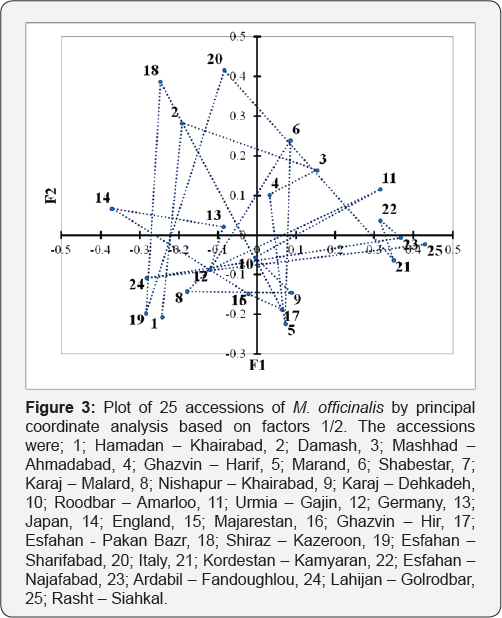
A principle coordinate's analysis (PCoA) of 25 accessions of M. officinalis were done for establishing the relationship among the accessions (Figure 3). The distribution pattern of accessions in PCoA was principally similar to the result of cluster analysis. The eigenvectors and eigenvalues of the PCoA obtained from data of the total proteins revealed that the accessions were separated on the five principal factors and influencing 53.7 percentages of the variation accumulated up to the first two factors.
Discussion
In this study we investigated the pattern and variability of M. officinalis is an important medicinal plant and characterization of genetic diversity in plants is susceptible to ontogeny and environmental condition. We hypothesized protein markers were used to distinguish genetic diversity among 25 accessions of M. officinalis. The gel analysis resulted in the detection of 18 protein bands with 59.09 percentage polymorphism. The results indicated that the main bands (common proteins) were same in all the accessions and the fundamental differences were for minor bands.
The results provided some interesting findings regarding to Jaccard's similarity coefficient results, the highest genetic distance was between Italy and N. Khorasan accessions. The results matched up well with the findings of Radwan et al. [19] that show crosses between accessions with high polymorphism could create more genetic diversity. Genetic diversity among the accessions was independent in terms of the collecting region. Some accessions with near geographically relationships were really unlike in their leaf protein bands, while some distant geographically accessions were too similar in their leaf proteins.
Dendrograms based on the Jaccard index and the WPGMA method are used to evaluate the degree of plants genetic diversity [30]. In the current study, the WPGMA cluster analysis of 25 accessions of M. officinalis based on the protein profiles generated three clusters. In summary, this study indicated that the ability to select leaf protein pattern for study of variability of officinalis accessions and showed that some Iranian accessions belonged to the first cluster were geographically close intervals, while Germany and Japan accessions with high geographical distance were geographically more distant from each other. The sources of the Pakan Bazr Company may be being collected at the close or the same original population of accessions segregation in the first group. The accessions of the second groups were collected from different region far apart from each other, which may be diverging from the same original population. These accessions could be mixed from different genotypes, or random mating populations with the same alleles but differing allele frequencies and probability due to open pollination of M. officinalis. Our results matched up well with the findings of Aharizad et al. [31], who reported that the morphological and essential oil content data of different M. officinalis accessions with high geographical distances were placed in the same group. In addition, Ghaffariyan et al. [32] in a study on genetic diversity of the 12 ecotypes of lemon balm using IRAP markers, based on long terminal repeats (LTRs) of barley retrotransposons showed that the average PIC and the average marker index between ecotypes were 0.27 and 14.36, respectively. Molecular analysis of variance indicated that the variance within population is greater than the variance between the populations, and finally, the populations were grouped into three clusters. Mustafa et al. [22] reported that the cluster analysis based on morphological trait, isozyme and seed protein data demonstrated genetic diversity among and within population of two species of Ocimum that may be due to environmental modification and natural hybridization. Sharma & Krishna [16] in a study on 52 accessions of cowpea (Vigna unguiculata) reported that SDS - PAGE approach is a powerful tool to identify diverse genotypes of cowpea. SDS-PAGE analysis of eleven samples of leaf protein in Ocimum showed that the method was adequate to determine genetic diversity Bompalli & Nallabilli [33].
Protein diversity is not random, since they associated with the genome expression and supply additional opportunities for polymorphism that noticed the presence of important genes for breeding purposes. Improvement in plant breeding program through selection is possible, especially if we extend the genetic basis from various habitats to comprise most of the genetic determinants of a desirable trait, such as; yield, a biotic stress tolerance, and quality Ghafoor & Arshad [34]. This result is reasonable given that reasonable genetic variation in the leaf protein pattern in 25 accessions of M. officinalis. This variation might depend on environmental conditions such as geographic regions, season of culturing, altitude, annual rainfall, temperature, land fertility and genotype variation [35].
Conclusion
This present study demonstrated the protein variation among the M. officinalis populations can be seriously taken into account in molecular studies. Furthermore, the electrophoresis of leaf proteins can be utilized as an effective strategy in the programs involved with M. officinalis conservation [15]. The observed differences among the accessions would be of immediate importance for development of the M. officinalis gene bank and may be used in hybridization and breeding programs to identify diverse parental combinations and creating segregating progeny with high genetic diversity [36]. Such a strategy in its turn could potentially lead to improving the phytochemical content of the plant [37]. However, future investigations into the limitations research and some other research questions to improve our knowledge are necessary
Acknowledgments
We thank to Medicinal Plants Research, Shahed University for supporting the present study
References
- Marongiu B, Porcedda S, Piras A, Rosa A, Deiana M, et al. (2004) Antioxidant activity of supercritical extract of Melissa officinalis subsp. officinalis and Melissa officinalis subsp inodora. Phytotherapy Research 18(10): 789-792.
- Bagdat RB, Cosge B (2012) The essential oil of lemon balm (Melissa officinalis L.), its components and using fields. Anadolu journal of agricultural sciences 21: 116-121.
- Soltani-Maivan E, Radjabian T, Abrishamchi P, Talei D (2017) Physiological and biochemical responses of Melissa officinalis L. to nickel stress and the protective role of salicylic acid. Archives of Agronomy and Soil Science 63(3): 330-343.
- Zargari A (1991) Medicinal plants. Tehran University Publications, Tehran, Iran.
- Kim S, Yun EJ, Bak JS, Lee H, Lee SJ, et al. (2010) Response surface optimised extraction and chromatographic purification of rosmarinic acid from (Melissa officinalis) leaves. Food chemistry 121: 521-526.
- Stefanovic O, Comic L (2012) Synergistic antibacterial interaction between Melissa officinalis extracts and antibiotics. Journal of Applied Pharmaceutical Science 02(01): 01-05.
- Sanchez-Medina A, Etheridge CJ, Hawkes GE, Hylands PJ, Pendry BA, et al. (2007) Comparison of rosmarinic acid content in commercial tinctures produced from fresh and dried lemon balm (Melissa officinalis). Journal of Pharmacy & Pharmaceutical Sciences 10(4): 455-463.
- Moradkhani H, Sargsyan E, Bibak H, Naseri B, Sadat-Hosseini M, et al. (2010) Melissa officinalis L., a valuable medicine plant: A review Journal of Medicinal Plants Research 4(25): 2753-2759.
- Chahal G, Gosal S (2002) Principles and procedures of plant breeding: Biotechnological and conventional approaches. Alpha Science Int'l Ltd.
- Talei D, Valdiani A, Abdullah MP (2014) Impact of protein diversification on morphometric behavior of Andrographis paniculata Nees. Plant systematics and evolution 300(5): 1003-1010.
- Nakamura H (2001) Genetic diversity of high-molecular-weight glutenin subunit compositions in landraces of hexaploid wheat from Japan. Euphytica 120(2): 227-234.
- Brown AH, Weir BS (1983) Measuring genetic variability in plant populations. Developments in Plant Genetics and Breeding 1(Part A): 219-239.
- Ernst M, Putnova L, Stohl R, Matouskova J (2017) Comparison of Czech and Latvian Beaver Population by Microsatellite Analysis and Genetic Differences between Castor Fiber and Castor Canadensis. Transylvanian Review 1 XXV(21).
- Gianibelli M, Gupta R, Lafiandra D, Margiotta B, MacRitchie F (2001) Polymorphism of high M r glutenin subunits in Triticum tauschii: Characterisation by chromatography and electrophoretic methods. Journal of Cereal Science 33(1): 39-52.
- Kutka-Hlozakova T, Gregova E, Vivodik M, Galova Z (2016) Genetic diversity of European cultivars of common wheat (Triticum aestivum L.) based on RAPD and protein markers. Journal of Central European Agriculture 17(4): 957-969.
- Sharma DB, Krishna KR (2017) Genetic Diversity in Cowpea [Vigna unguiculata (L.) Walp.] Accessions using Protein Profiling. Int J Pure App Biosci 5(2): 491-496.
- Abou-Ellali M, Hattem ME, Mohamad AM, Usama LA, Hasan AG, et al. (2014) Appraisal of biochemical and genetic diversity of Mango cultivars using molecular makers. African journal of biotechnology 13: 2796-2806.
- Masoumi SM, Kahrizi D, Rostami-Ahmadvandi H, Soorni J, Kiani S, et al. (2012) Genetic diversity study of some medicinal plant accessions belong to Apiaceae family based on seed storage proteins patterns. Molecular biology reports 39: 10361-10365.
- Radwan SA, El-Koly AS, Sammour RH (2013) Genetic variation among accessions of (Lathyrus inconspicuous L.) as revealed by SDS Polyacrylamide Gel Electrophoresis. Acta agriculturae Slovenica 101(1): 21-30.
- Sadia M, Malik SA, Rabbani MA, Pearce S (2009) Electrophoretic characterization and the relationship between some Brassica species. Electronic Journal of Biology 5(1): 1-4.
- Hassan MG, Badr AT, Mustafa A, El-Galaly MA, Mobarak AA (2003) Genetic diversity among Mentha population in Egypt as reflected by morphological and protein electrophoretic variation. Journal of Agricultural and Food Chemistry 2: 269-286.
- Mustafa A, Badr A, El-Galaly MA, Mobarak AA, Hassan MG (2006) Genetic diversity among Ocimum population in Egypt as reflected by morphological, seed proteins and isozyme polymorphism. International Journal of Botany 2: 261-269.
- Boulter D, Thurman D, Derbyshire E (1967) A disc electrophoretic study of globulin proteins of legume seeds with reference to their systematics. New Phytologist 66(1): 27-36.
- El-Bakatoushi R (2010) Genetic diversity of winter wheat (Triticum aestivum L.) growing near a high voltage transmission line. Romanian Journal of Biology Plant Biology 55(2): 71-87.
- Khurshid H, Rabbani MA (2012) Comparison of electrophoretic protein profiles from seed of different oilseed Brassica cultivars. Practice 4: 6.
- Talei D, Valdiani A, Puad MA (2013) An effective protein extraction method for two-dimensional electrophoresis in the anticancer herb Andrographis paniculata Nees. Biotechnology and applied biochemistry 60(5): 521-526.
- Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227: 680-685.
- Rohlf F (1998) NTSYS-pc version 2.0. Numerical taxonomy and multivariate analysis system. Exeter software, Setauket, New York, USA.
- Perrier X J-CJ (2006) DARwin Software.
- Bechkri S, Medoukali I, Khelifi D (2017) Ecogeographic variability and genetic diversity associated with seed albumins, globulins and prolamins patterns in Vicia taxa from Algeria. Bot Stud 58(1): 27.
- Aharizad S, Rahimi MH, Moghadam M, Mohebalipour N (2012) Study of genetic diversity in lemon balm (Melissa officinalis L.) populations based on morphological traits and essential oils content. Annals Biological Research 3(12): 5748-5753.
- Ghaffariyan S, Mohammadi S, Aharizad S (2011) Patterns of population diversity in Lemon balm (Melissa officinalis L.) as revealed by IRAP markers. Journal of Plant Physiology and Breeding 2: 39-51.
- Bompalli LK, Nallabilli L (2013) Genetic diversity of Ocimum species through biochemical technique and UPGMA cluster analysis. International Journal of pharmaceutical sciences 5(4): 155-159.
- Ghafoor A, Arshad M (2008) Multivariate analyses for quantitative traits to determine genetic diversity of blackgram Vigna mungo L. Hepper germplasm. Pakistan Journal of Botany 40: 2307-2313.
- Vollmann J, Fritz CN, Wagentristl H, Ruckenbauer P (2000) Environmental and genetic variation of soybean seed protein content under Central European growing conditions. Journal of the Science of Food and Agriculture 80: 1300-1306.
- Valdiani A, Kadir MA, Saad MS, Talei D, Tan SG, et al. (2012) Intraspecific hybridization: generator of genetic diversification and heterosis in Andrographis paniculata Nees. A bridge from extinction to survival. Gene 505(1): 23-36.
- Valdiani A, Talei D, Tan SG, Kadir MA, Maziah M, et al. (2014) A classical genetic solution to enhance the biosynthesis of anticancer phytochemicals in Andrographis paniculata Nees. PloS one 9: e87034.






























