Genetic Diversity Based on Multivariate analysis m for Yield and it's Contributing Characters in Bread Wheat (Triticum aestivum L.) Genotypes
Birhanu Mecha1, Sentayehu Alamerew2, Alemayehu Assefa3, Ermias Assefa4* and Dargicho Dutamo5
1Wachemo University, Ethiopia
2Jimma University, Ethiopia
3Ethiopian Institute of Agricultural Research (EIAR), Ethiopia
4Ethiopian Biotechnology Research Institute, Ethiopia
5Mizan-Tepi University, Ethiopia
Submission: July 13, 2016; Published: July 28, 2017
*Corresponding author: Ermias Assefa, Ethiopian biotechnology institute, Ethiopia, Email: ethioerm99@gmail.com
How to cite this article: Birhanu M, Sentayehu A, Alemayehu A, Ermias A, Dargicho D. Genetic Diversity Based on Multivariate analyses for Yield and it's Contributing Characters in Bread Wheat (Triticum aestivum L.) Genotypes. Agri Res & Tech: Open Access J. 2017; 8(5): 555748. DOI: 10.19080/ARTOAJ.2017.08.555748
Abstract
Breed wheat is one of the most important export and strategic cereal crop in the world and in Ethiopia in terms of production and utilization. Information about genetic diversity is important for selection of promising parents in hybrid variety and inbred line development, in assigning inbred lines to heterotic groups, variety registration and protection and to estimate the potential of genetic gain in a breeding program. The overall objective was to assess the genetic diversity of wheat genotypes by multivariate analyses (Cluster and Principal component). The experimental material consisted of sixty four breed wheat genotypes tested in 8x8 Simple Lattice Design at Ginchi. Analysis of variance revealed highly significant differences among accessions for all traits. Cluster analysis revealed that the 61 genotypes and 3 checks were grouped into eight clusters. The crosses between genotypes selected from cluster-V with cluster-VIII and cluster V with cluster VII are expected to produce better genetic recombination and segregation in their progenies. Five principal components (PC1, PC2, PC3, PC4 and PC5) exhibited eigen value greater than one and accounted 72.78% of the total variation.
Keywords: Inbread line; Cluster; Principal component; Heterotic group
Introduction
Bread wheat (Triticum aestivum L. em Thell. 2n=6x=42), a self-pollinating annual plant in the true grass family Gramineae (Poaceae), is the largest cereal crop extensively grown as staple food source in the world [1].
Cluster analysis is a multivariate method, which aims to classify a sample of subjects based on a set of measured variables into a number of different groups such that similar subjects are placed in the same group. It sorts genotypes into groups, or clusters, so the degree of association will be strong between members of the same cluster and weak between members of different clusters. The cluster analysis was performed using a measure of similarity levels and Euclidean distance [2,3].
Different researcher grouped bread wheat genotypes using cluster analysis. Hailegiorgis et al. [4] reported that the cluster analysis grouped the 49 bread wheat genotypes into 22 different clusters. This indicates the presence of wide diversity among the tested genotypes. Khodadadi et al. [5] determined the genetic diversity of 36 winter wheat cultivars from Iran and by using cluster analysis, seven clusters were determined.
Principal component analysis makes it possible to transform a given set of characteristics (variables), which are mutually correlated, into a new system of characteristics, known as principal components, which are not correlated. The obtained variables may also be used for further analysis, where the assumption of no co-linearity is required. Moreover, the analysis is characterized by the fact that it includes the total variance of variables, explains maximum of variance within a data set, and is a function of primary variables.
Precise knowledge about germplasm diversity and genetic relationship among breeding materials is a pre-requisite for crop improvement programs as it helps in the development of superior recombinants [6]. Genetic divergence analysis estimates the extent of diversity existed among selected genotypes [7]. Precise information on the nature and degree of genetic diversity helps the plant breeder in choosing the diverse parents for purposeful hybridization [4]. Generating fertile genetic diversity information among wheat genotypes is very important tool because the information will help the wheat breeders to bread for many characters (earliness, yield increase, drought tolerance, etc). Within the investigated material and this new material will serve as a new stock for the improvement of wheat breeding program for traits of interest. Therefore, the objective of this study was to assess the genetic diversity of durum wheat accessions by multivariate analyses.
Materials and Methods
Description of the study area
The experiment was conducted at Ginchi, West Shewa in 2012/13 cropping season. Ginchi Agricultural Research Sub Center is located at an altitude of 2240 meters above sea level, 84 kilometers (kms) to the West of Addis Ababa, and at a Latitude and Longitude of 09°03'N and 38°15'E, respectively. It is the center where the cereal crops like Teff, barley and wheat are grown. The maximum and minimum temperatures of the area are 24.72 °C and 8.76 °C, respectively, whereas the mean annual rainfall is 1080.4mm. The major soil types are black (Vertisol) and clay loam with pH of 6.4, which is heavy clay with 0.911.32% organic matter (HARC, Soil Analysis and Plant Physiology Team, 2012).
Experimental material
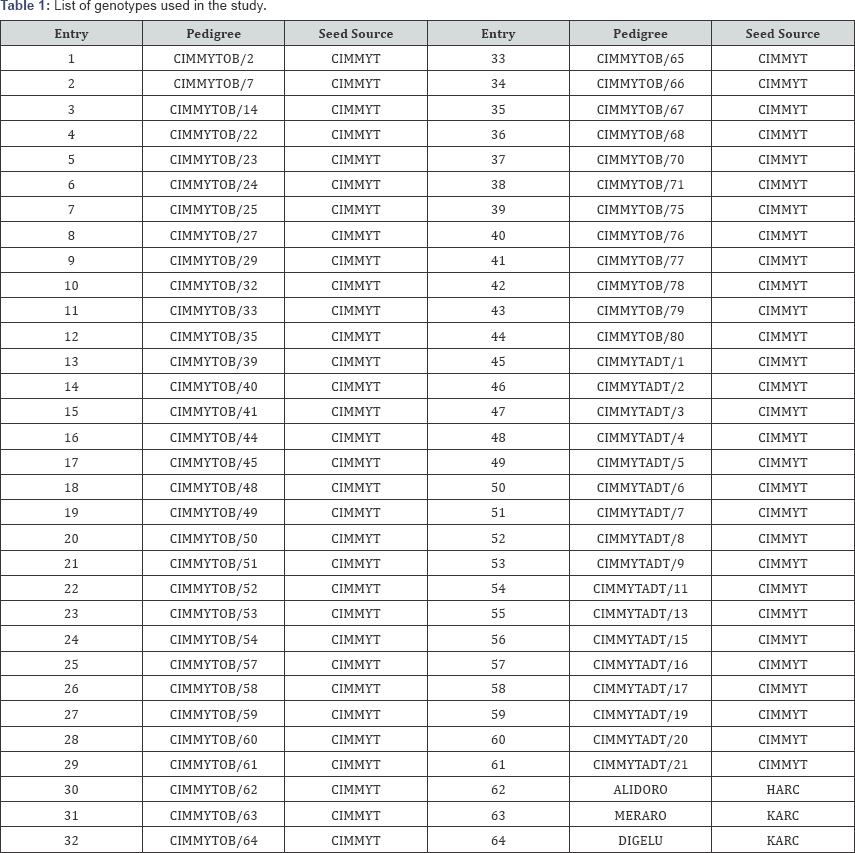
A total of sixty four bread wheat (Triticum aestivum L.) genotypes that include three standard checks and sixty one exotic bread wheat accessions introduced from CIMMYT were included in this study. The accessions were obtained kindly from HARC. The three released cultivars Digelu, Alidoro and Meraro were used as a standard checks. They were selected based on their agronomic performances and suitability to the growing conditions (Table 1). Cipal component, genetic recombination, segregation.
Experimental design and trial management
The experiment was carried out in 8x8 Simple Lattice Design at random. The genotypes were grown under uniform rain fed conditions. The plot size was six rows of 2.5m length with 0.2m row spacing i.e. 1.2mx2.5m=3m2 (standard plot size for variety trial). Planting was done by hand drilling on July 06, 2012. Seed rate was 150kg/ha (45 g/plot). Recommended fertilizer rate of 100/100kg/ha N/P2O5 in the forms of Urea and DAP was applied to each plot in the shallow furrow depths and mixed with soil at the same time during sowing. For data collection, the middle four rows were used (2 m2 area). The central four rows were harvested for grain yield and biomass yield from each plot leaving boarder rows to avoid boarder effects. All other agronomic practices were undertaken uniformly to the entire plot as recommended for wheat production in the area during the growing season.
Description of data collected
The data on the following attributes was collected on the basis of the central four rows in each plot per replication.
Days to 50% heading (DH): The numbers of days from sowing to 50% of plants have started heading.
Days to 75% maturity (DM): The numbers of days from date of sowing to a stage at which 75% of the plants have reached physiological maturity or 75% of the spikes on the plots turned golden yellow color.
Grain filling period: The grain filling period in days was computed by subtracting the number of days to heading from the number of days to maturity.
Thousand kernels weight (TKW) (g): The weight of 1000 kernels from randomly sampled seeds per plot measured with sensitive balance.
Grain yield per plot (GYP) (g): The grain yield per plot was measured using sensitive balance after moisture of the seed is adjusted to 12.5%. Total dry weight of grains harvested from the middle four rows was taken as grain yield t.
Biomass yield per plot (BMYP): It was recorded by weighing the total above ground yield harvested from the four central rows of each experimental plot at the time of harvest.
Harvest index (%): It was estimated by dividing grain yield per plot to biological yield per plot.
Hectoliter weight (HLW) (kg/ha): It is grain weight of one hectoliter volume random sample of wheat grain for each experimental plot.
Ten plants were randomly selected from the four central plots for recording the following observations:
Plant height (cm): The average height of ten randomly taken plants at the maturity time from the middle four rows of each plot of the replication was measured from the ground level to the top of the spike excluding the awn.
Number of productive tillers per plant: The numbers of tillers per plant bearing productive heads were counted at the time of harvest and average was recorded for the ten randomly taken plants from the middle four rows.
Spike length (cm): The average spike length of ten randomly taken plants from the base of the main spike to the top of the last spikelet excluding awns was recorded from four central rows of each plot.
Number of spikelets per spike: Total number of spikelets on main spike of all ten plants from four central rows was counted at the time of maturity and average was recorded.
Number of kernels per spike (NKPS): Total number of grains in the main spike were counted at the time of harvest from ten randomly taken plants and expressed as average and recorded from four central rows of each plot.
Statistical analysis
Analysis of variance (ANOVA): The data collected for each quantitative trait were subjected to analysis of variance (ANOVA) for simple lattice design. Analysis of variance was done using Proc lattice and Proc GLM procedures of SAS version 9.2 (SAS Institute, 2008) after testing the ANOVA assumptions.
The mathematical model for Simple Lattice Design is:
Yijr = μ + Ar + Gij +Bir +Bjr + eijr,
Where Yijr = the value observed for the plot in the rth replication containing the genotype Gij, μ= grand mean, Gij = genotype effect in the ith row and jth column, Ar= replication effect, Bir=ith block effect, Bjr =jth block effect, eijr, = the plot residual effect.
Cluster analysis
Clustering the genotypes into different groups was carried out by average linkage method. The appropriate number of clusters was determined from the values of Pseudo F and Pseudo T2 statistics using the procedures of SAS computer software version 9.2 to group sets of genotypes into homogeneous clusters [8].
Genetic divergence analysis
Genetic divergence between clusters was determined using the generalized Mahalanobis D2 statistics [1]. The D2 analysis performed based on the mean values of all traits by using SAS software program. In matrix notation, the distance between any two groups was estimated from the following relationship.
D2ij = (Xi-xj)S-1(Xi-Xj)
Where, D2ij = the square distance between any two accessions i and j; Xi and Xj = the vectors for the values for accession ith and jth genotypes; and S-1= the inverse of pooled variance covariance matrix within groups.
Testing the significance of the squared distance values obtained for a pair of clusters was taken as the calculated value of X2 (chi-square) and tested against the tabulated X2 values at n-2 degree of freedom at 1% and 5% probability level, where n= number of characters used for clustering genotypes.
Principal component analysis (PCA)
The principal component analysis was performed using the proc princomp procedure of SAS version 9.2 software [8]. Statistical inference was computed by taking into account all the factors at a time. In this study, investigation of suitable multivariate technique for analyzing data for all the characters is proposed. The general formula to compute scores on the first component extracted in a principal component analysis:
PC1=b11(X1]+b12+---b1p=(Xp]
Where, PC1 = the subject's score on principal component 1 (the first component extracted], blp = the regression coefficient (or weight) for observed variable p, as used in creating principal component 1 and Xp = the subject's score on observed variable p.
Results and Discussion
Analysis of variance (ANOVA)
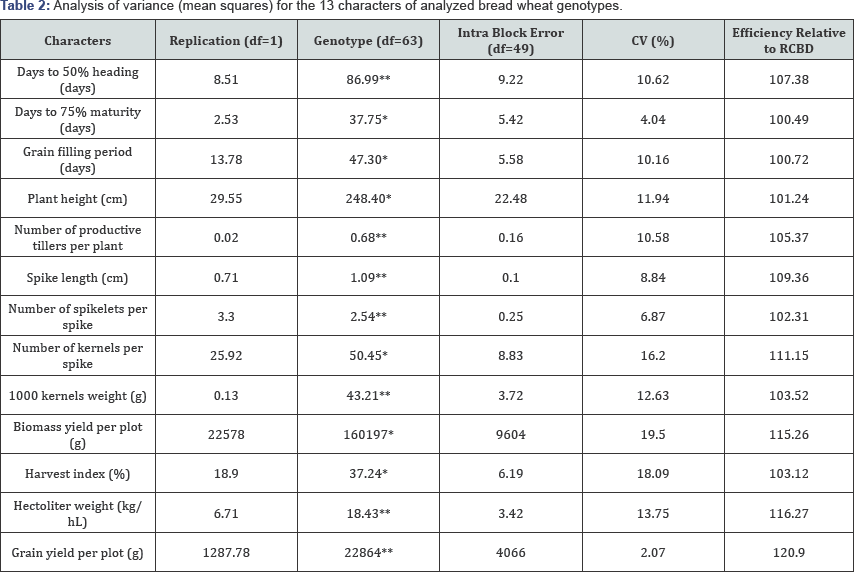
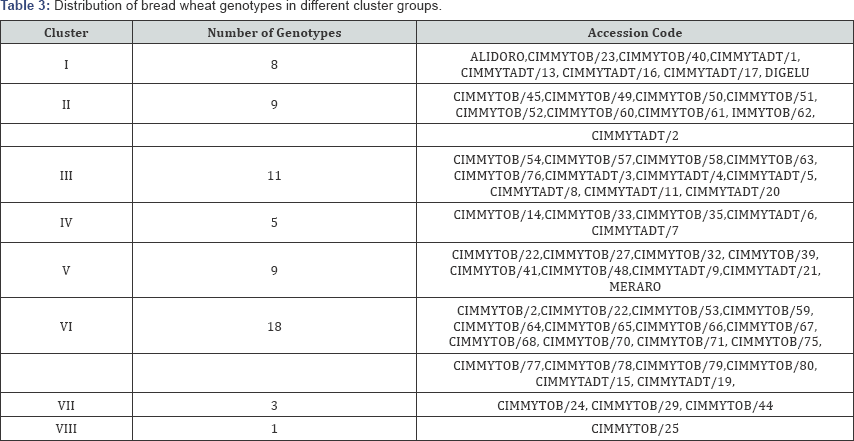
Mean squares of the 13 traits from analysis of variance (ANOVA] are presented in Table 2. Highly significant differences among genotypes (P<0.01) were observed for seven characters (days to heading, number of productive tillers per plant, spike length, number of spikelets per spike, 1000 kernel weight, grain yield plot-1 and hectoliter weight or test weight], significant at (P<0.05) for the rest six characters; namely, days to 75% maturity, grain filling period, plant height, number of kernels spike-1, biomass yield and harvest index. This result indicates that there is variability among the studied genotypes (Table 3).
Genetic divergence
Genetic divergence analysis quantifies the genetic distance among the selected genotypes and reflects the relative contribution of specific traits towards the total divergence. The genetic improvement through hybridization and selection depends upon the extent of genetic diversity between parents [9].
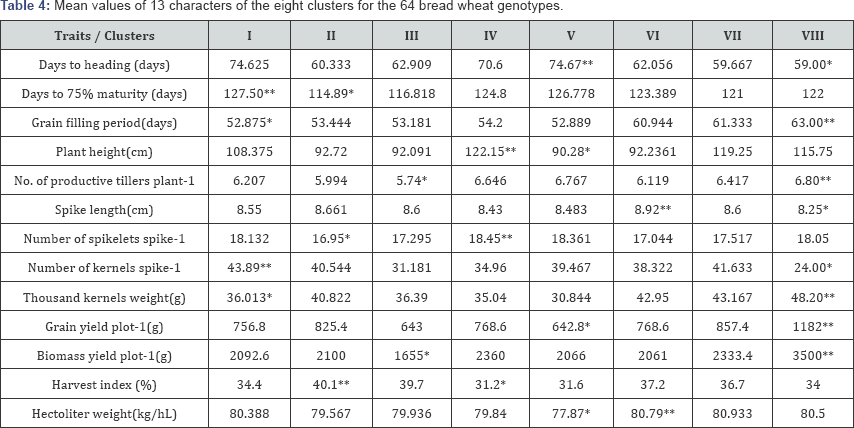
* =Lowest value, **= Highest value.
Cluster mean analysis: The mean values of the thirteen traits in each cluster are presented in Table 4. Cluster I exhibited the highest days to maturity and the delayed maturity time (127.5 days) with largest number of kernels per spike (43.89); thus containing late maturing genotypes. Cluster II could be characterized by the earliest maturity time (114.890 days) and low number of spikelets per spike (16.95) but with the highest harvest index (40.1%). Cluster III had the lowest number of productive tillers per plant (5.736 with the lowest biomass yield per plot (1654.6g).
A relatively larger range between clusters was displayed for plant height. Cluster V had the shortest plant height (90.28cm) and cluster IV revealed the highest (122.15cm) with the highest number of spikelets per spike (18.45), but it showed the lowest harvest index (31.2%). The highest number of days to fill grain was observed in cluster VIII (63.00 days) and the lowest goes for cluster I (52.875 days). The ranges of number of spikelets spike-1, thousand kernels weight (g), harvest index (%), spike length (cm) and hectoliter weight (kg/hL) were low.
Cluster V showed the shortest plant height (90.28cm), the lowest yield per plot (642.8g) and the lowest hectoliter weight (77.87kg/hL) but revealed the latest days to heading (74.667 days) and maturity (126.778 days). Cluster VI showed the longest spike length (8.92cm) and the highest hectoliter weight (80.79kg/hL). Cluster VII revealed intermediate characteristics for all of the traits. Cluster VIII revealed the highest grain yield (1182g), the heaviest thousand kernels weight (48.20g) and the highest biological yield per plot (3500g) but the lowest number of kernels per spike (24.00).
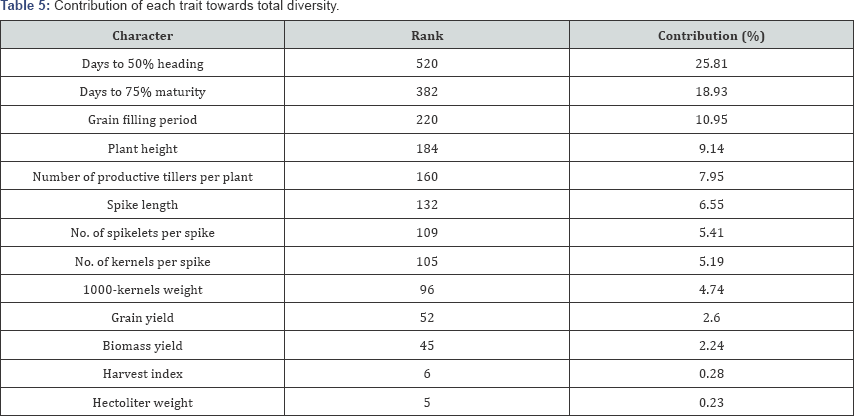
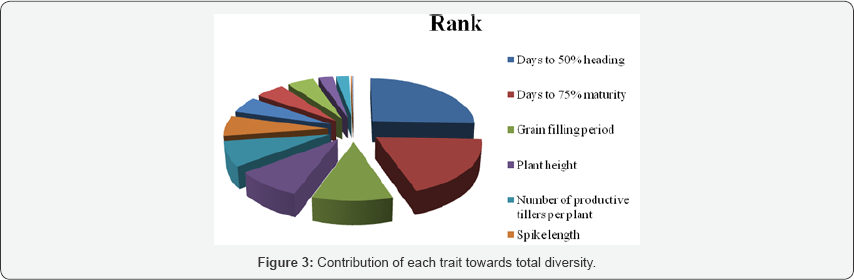
Relative contribution of each character towards divergence: The analysis of the contribution of each character towards the expression of genetic divergence (Table 5) indicated that days to 50% heading contributed maximum followed by days to 75% maturity to the total genetic divergence in the genotypes studied. These two traits followed by grain filling period, plant height, number of productive tillers per plant, days to 75% maturity, spike length, number of spikelets per spike, number of kernels per spike and thousand kernels weight totally accounted for 94.65% of total genetic divergence in the materials studied. Grain yield, biomass yield, harvest index and hectoliter weight were the least contributor to the divergence.
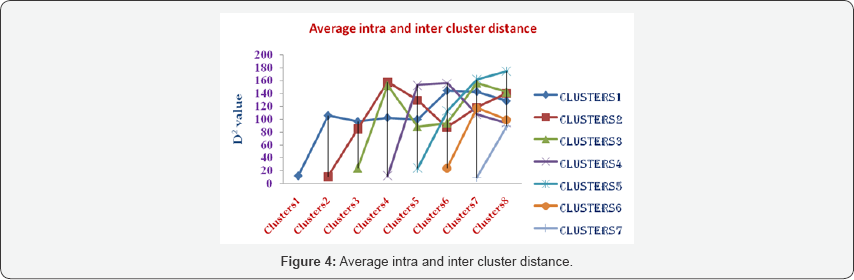
Average intra and inter cluster distance (D2): The average intra and inter cluster distance D2 values are presented in Table 6. Maximum average intra cluster D2 was shown by cluster III (24.35) followed by cluster VI (23.99).
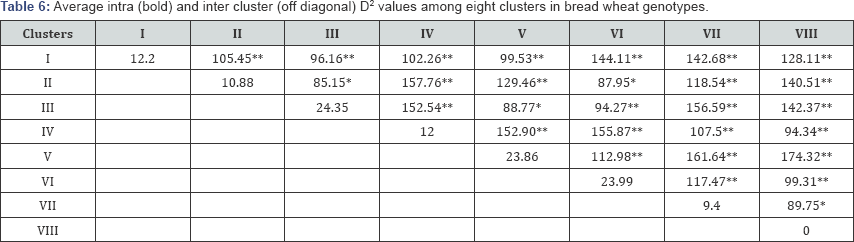
x2=82.529 at 5% probability level and x2=92.010 at 1% probability level, *= Significant at 0.05 probability level ,**= Highly significant at 0.01 probability level.
The lowest intra cluster distance D2 was recorded in cluster VII (9.40), which shows the presence of less genetic variability or diversity within this cluster.
The diversity among clusters or inter cluster distance D2 ranged from 85.15 to 174.32. Cluster V and VIII showed maximum inter cluster distance of 174.32, followed by that between clusters V and VII (161.64). The lowest inter cluster distance was noticed between clusters II and III (85.15), followed by that between clusters II and VI (88.77). Evaluation of genetic diversity can be useful for the selection of the most efficient genotypes. The results of this study showed the presence of a high genetic divergence among wheat genotypes, similar to the findings of Ali et al. [10] who reported that cluster analysis can be useful for finding high yielding wheat genotypes. According to Rahim et al. [11] who showed that the hybrids of genotypes with maximum distance resulted in high yield, the cross between these genotypes can be used in breeding programs to achieve maximum heterosis. Therefore, more emphasis should be given on cluster V and VIII for selecting genotypes as parents for crossing with the genotypes of cluster, which may produce new recombinants with desired traits. The chi-square test for the clusters indicated that there was a statistically significant difference in all characters (Table 7). The x2- test for the eight clusters indicated that there was a statistically significant difference in all characters.
Principal component analysis
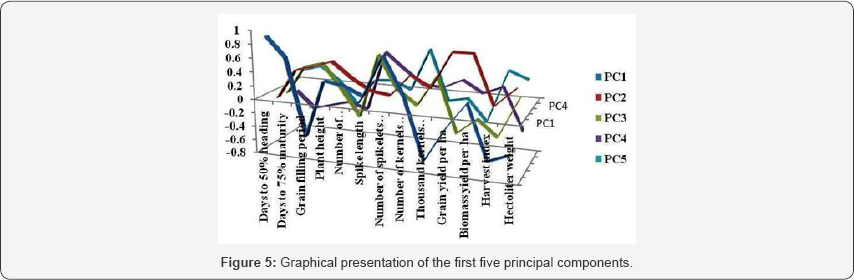
Principal component analysis reflects the importance of the largest contributor to the total variation at each axis for differentiation [12]. The data matrix of 13*64 was prepared for principal component analysis. Out of thirteen principal components (PCs), the first five exhibited eigen value greater than one (significant). The rest eight PCs explained nonsignificant amount of variation. The eigen values are used to determine how many factors to retain. The sum of the eigen values is usually equal to the number of variables. Therefore, in this analysis the first factor retains the information contained in 3.36 of the original variables. The coefficients defining the first five principal components of these data are given in Table 7. The principal component analysis revealed that five principal components (PC1-C5) exhibited eigen value higher than one, with values 3.36, 2.46, 1.43, 1.19 and 1.03, respectively, have accounted for 72.78% of the total variation so these five were given due importance for further explanation.
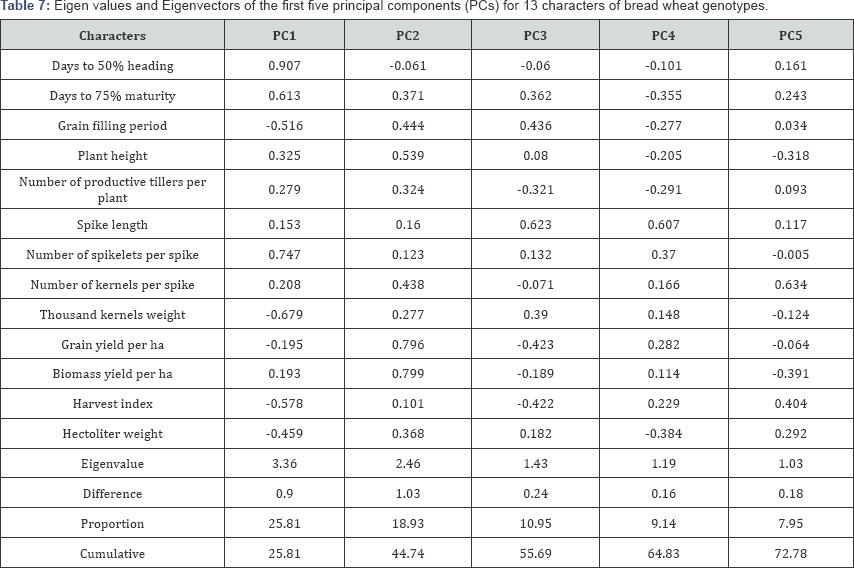
According to Chahal & Gosal [13], characters with largest absolute value closer to zero. Therefore, in the present study, absolute value closer to unity within the first principal differentiation of the genotypes into different clusters was component influence the clustering more than those with lower because of relatively high contribution of few characters rather than small contribution from each character. Accordingly, the first principal component (PC1) which accounted for 2 5.81% of the variability among genotypes were attributed to discriminatory traits such as days to heading, days to maturity, grain filling period, plant height, number of spikelets per spike, thousand kernels weight, harvest index and hectoliter weight. Likewise, 18.93% of the total variability among the tested genotypes accounted for the second principal component (PC2) mainly extracted from variation in days to maturity, grain filling period, plant height, number of productive tillers per plant, number of kernels per spike, grain yield per plot, biomass yield and test weight. Similarly, the major contributing characters for the 10.94% of total variation in the third principal component (PC3) were spike length, grain filling period, grain yield, harvest index, thousand kernels weight, and days to maturity and number of productive tillers per plant [14-18].
Furthermore, the fourth principal component (PC4), which explained 9.14% of total variations was obtained from days to maturity, spike length, number of spikelets per spike and hectoliter weight. Quantitative characters such as plant height, number of kernels per spike, biomass yield per plot and harvest index, explained mainly for the remaining 7.95% of the variations of fifth principal component (PC3) (Figures 1-5).
Summary and Conclusion
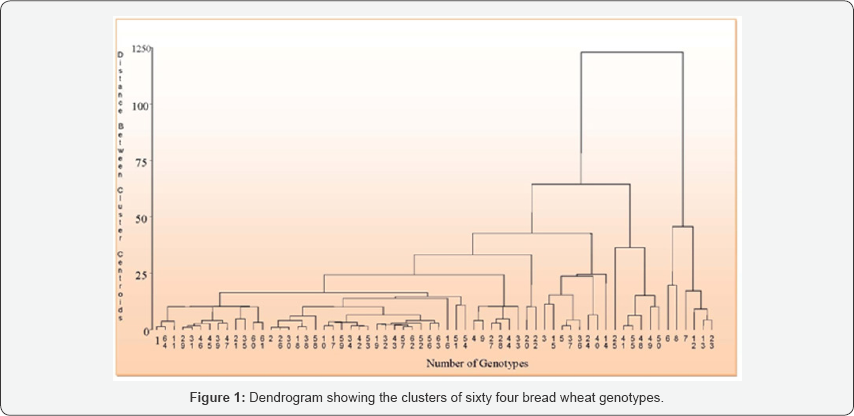
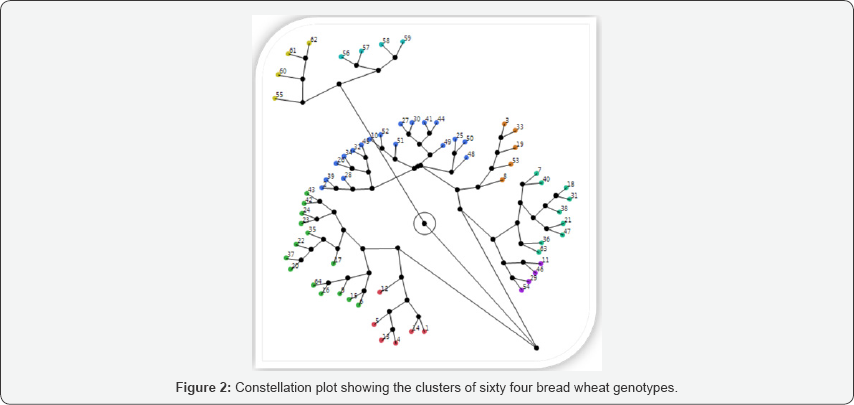
Clustering was made to categorize quantitative traits into components for the sake of understanding the share components contribute to major variation in the study. The cluster analysis based on D2 analysis on pooled mean of genotypes classified the 64 genotypes into eight clusters; which makes them to be moderately divergent. There was statistically approved difference between all the characters. Mean values in each cluster revealed that genotypes in cluster I had relatively moderate in most of characteristics but late heading and maturity days with the shortest time to fill the kernels. Mean values in genotypes grouped in cluster II had low to moderate characteristics with earliest maturity time and high number of kernels per spike with high thousand kernels weight. Mean values of cluster III showed the least number of productive tillers per plant, short time to fill the kernels, the earliest maturing types, low in grain and biomass yields and the lowest number of kernels per spike. The highest inter-cluster distance were exhibited between cluster V and VIII (D2 = 174.32), cluster V and VII (D2 = 161.64) and cluster II and IV (D2 = 157.76), indicates wider genetic divergence among the clusters. Whereas, the shortest squared distance was observed between cluster II and III (D2 = 85.15) at, followed by between cluster II and VI (D2 = 88.77).The crosses between genotypes selected from cluster-V with cluster-VIII, cluster V with cluster VII and cluster II and IV are expected to produce better genetic recombination and segregation in their progenies. Therefore, these bread wheat genotypes need to be crossed and selected to develop high yielding variety. The principal component analysis revealed that principal components PC1, PC2, PC3, PC4 and PC5, accounted for 72.78% of the total variation. This result further confirmed the presence of ample genetic diversity for use in improvement program.
Acknowledgment
The authors would like to thank Jimma University College of Agriculture and Veterinary Medicine (JUCAVM) and Holeta Agricultural Research Center (HARC) for their financial support.
References
- Mahalanobis PC (1936) On the generalized distance in statistics. Proc Nat Inst Sci 2: 49-55.
- Everitt BS (1993) Cluster Analysis. Wiley, New York, USA.
- Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression ilatteriis. Proc Natl Acad Sci U S A 95(25): 14863-14868.
- Hailegiorgis D, Mesfin M, Genet T (2011) Genetic divergence analysis on some bread wheat genotypes grown in ethiopia. 2Bahir Dar University, Amhara, Ethiopia.
- Khodadadi M, Hossein Fotokian M, Miransari M (2011) Genetic diversity of wheat (Triticum aestivum L.) genotypes based on cluster and principal component analyses for breeding strategies. Australian Journal of Crop Sciences 5(1): 17-24.
- Rauf S, Tariq SA, Hassan SW (2012) Estimation of pedigree based diversity in Pakistani wheat (Triticum aestvium L.) germplasm.
- Mondai SK, Khajuria MR (2001) Correlation and path analysis in bread wheat (Triticum aestivum L.) under rainfed condition. Environment and Ecology 18(2): 405-408.
- SAS (2008) Statistical Analysis System. Version 9.2, SAS Institute Inc., Cary, NC, USA.
- Hailu A, Alamerew S, Nigussie M, Assefa E (2016) Study of genetic diversity in different genotypes of barley (Hordeum vulgare l.) based on cluster and principal component analyses. Agricultural Science Research Journal 6(2): 31-42.
- Ali Y, Atta BM, Akhter J, Monneveux P, Lateef Z (2008) Genetic variability, association and diversity studies in wheat (Triticum aestivum L.) germplasm. Pak J Bot 40(5): 2087-2097.
- Rahim MA, Mia AA, Mahmud F, Zeba N, Afrin K (2010) Genetic variability, character association and genetic divergence in murgbean. Plant Omics 3(1): 1-6.
- Sharma PK, Gupta PK, Balayn HS (1998) Genetic diversity in a large collection of wheats (Triticum spp.). Indian Journal of Genetics and Plant Breeding 58(3): 271-278.
- Chahal GS, Gosal SS (2002) Principles and procedures of plant breeding: biotechnology and conventional approaches. Narosa Publishing House, New Delhi, India, p. 604.
- Degewione A, Alamerew S (2013) Genetic diversity in bread wheat (Triticum aestivum L.) genotypes. Pak J Biol Sci 16(21): 1330-1335.
- Rymuza K, Turska E, Wielogórska G, Bombik A (2012) Use of principal component analysis for the assessment of spring wheat characteristics. Acta Sci Pol Agricultural 1(1): 79-90.
- Mollasadeghi V, Shahryari R (2011) Important morphological markers for improvement of yield in bread wheat. Advances Environ Biol 5(3): 538-542.
- Noorka IR, Khaliq I (2007) An efficient technique for screening wheat (Triticum aestivum L.) germplasm for drought tolerance. Pak J Bot 39(5): 1539-1546.
- Shashikala SK (2006) Analysis of genetic diversity in wheat. University of Agricultural Sciences, Dharwad, India.






























