Problems in Recovering Total RNA from the Soil
Abbas A*
Department of Plant Pathology, The University of Agriculture Peshawar, Pakistan
Submission: May 17, 2017; Published: June 05, 2017
*Corresponding author: Abbas A, Department of Plant Pathology, The University of Agriculture Peshawar, Pakistan, Email: aqlpath@gmail.com
How to cite this article: Abbas A. Problems in Recovering Total RNA from the Soil. Agri Res & Tech: Open Access J. 2017; 7(4): 555717.DOI: 10.19080/ARTOAJ.2017.07.555717.
Abstract
Microbial gene expression provides information about their activities in the soil. The extraction of total RNA from the soil is a key step in the detection of those gene expressions. Currently several methods of total RNA extraction have been reported. However to recover total RNA from certain soils is still a challenge. In this mini-review we highlight those challenges.
Keywords: Soil; RNA extraction; Soil impurities; rRNA
Introduction
The isolation and identification of microorganism from the soil is a never ending task. Most of the microorganism cannot be isolated by culture methods. Culture based methods are no doubt have limitations for the survey of microbial populations in soil. Researchers have tried new culture independent procedures [1].Polymerase chain reaction (PCR) was invented in the mid- 1980 to detect genes of microorganism of soil. With the passage of time small subunit and large subunit ribosomal RNA genes have been detected by PCR. With the development of molecular technique such as various typesof PCRs it was found that the microbial world is genetically and functionally more complex [2].Moreover Hugh diversity is found than on the basis of culture based techniques. Moreover using molecular techniques large amount of information regarding microorganism was revealed and this information was utilized for identifying newly microbial species [3]. There are several reports of RNA extraction from soil by using kits (Table 1) however until now there is no any method which can be used to extract RNA from all types of soil. The lack of universal RNA extraction methods hinder the study of all microbial expression in the soil [4]. The following are few difficulties in recovering RNA from the soil.
Soil impurities
- Humic acids
- Fulvic acids
- Humin
Most of humic acids are soluble under alkaline conditions whereas fulvic acids are soluble under all pH conditions. However humin is insoluble fraction and also it cannot be extraction along with RNA. So the predominant compounds which are coextracted with RNA are humic and fulvic acids. Fulvic acids at higher concentration usually inhibit PCR amplification. Humic acids possess more difficulties in biological experiments as well as it interferes with the enzymatic reactions as well as various PCR techniques [6]. Therefore the removal of humic substances from the soil is very important, however the complete removal of humic substances is rather difficult.
Soil types
There may be several successful cases of the RNA extraction from the Diverse soil however to extract RNA from Andosols seem to be more difficult. Andosols is actually volcanic ash soils and found in throughout the world. Various kits has been proven to extract RNA from anodsols still RNA extraction failed in some of andosols. These soils may have severae RNase activities however currently there are reports that RNA can survive in the presence of RNase in the soil [7]. Even the intact bacterial rRNA can be recovered from andosols by using various extraction buffers. The failure of RNA extraction from andosols is possibly because of RNA adsorption by soil but not because of degradation of RNA by RNases. The problem of RNA is that it adsorbs to soil rapidly About half of the adsorbed RNA molecules were absorbed by clay within one hour and 85% of the maximum adsorption occurred on the major components in andosols called allophance within half an hour. Mononucleotides, nucleosides, bases, phosphate and ribose sugars are parts of RNA though having different no of phosphate groups still adsorbed by the soil allophone [8]. The DNA could also be absorbed by soil however that RNA is more difficult to recover from soil. This may be because of ribose of RNA having on more OH group than the deoxyribose sugar of DNA. The increase in hydroxyl group may cause the stronger adsorption of RNA on soil than the DNA [9]. Secondly there are some free extra cyclic functional groups are present in the single strand structure of RNA and because of these functional group base pairing sometime occurs in some regions of RNA molecules.Some of these functional groups form hydrogen bonds with the soil surface. There is an extraction buffer called skim milk amended extraction buffer which can be used to successfully recover the DNA from the soil but failed to recover RNA [10]. Recently it was found that clay contents found in particular soil type significantly affects the RNA yield as well as PCR analysis. Clay content also negatively affect all RNA isolation methods. In case of andosols still it is unclear whether the clay content are playing major role in adsorption of RNA on soil [11]. Therefore more research is needed to explore the RNA adsorption by andosols.
Presence of rRNA
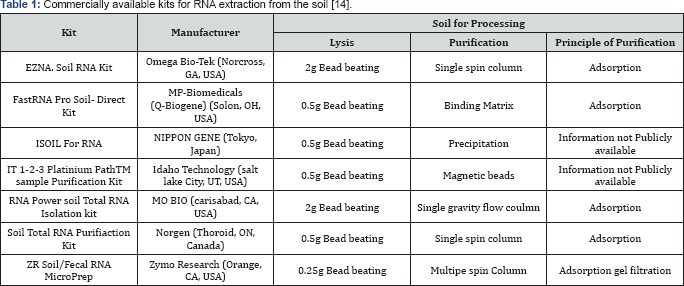
In case of prokaryotes the mRNA does not normally have a poly (A) tail which sometime makes the purification of bacterial mRNA difficult. Recent research revealed that there are some bacteria which mRNA molecules contain a poly (A) tail. Unluckily these mRNA is part of whole transcriptome and therefore prone to rapid degradation [12]. Therefore to obtain bacterial mRNA without the contamination of rRNA is one of the major challenge. For PCR analysis (RT and Qrt) the presence of thus, obtaining bacterial mRNA without the contamination of rRNA is a challenge. For RT-PCR and qRT-PCR, the existence of rRNA in an RNA sample is not a problem as specific primers are used. It is rather difficult to recover total RNA from the soil and the yield of RNA extracted from one gram of soil ranges from nanogram to several micrograms. Low yield of RNA may be because of several reasons the amount of microorganism, inhibition by humic substances or degradation of RNA during purification. For RT-PCR and Qrt-PCR analysis, the quantity of RNA in nanogram could be sufficient however microarray and high-throughput put sequencing analysis needs microgram level of RNA especially for the detection of rare sequences. A large amount of RNA as well as more purification procedures is essential for the microarray or high throughout sequencing analysis [13]. If the soil amount is increased for RNA extraction the amount of humic acids increases in RNA samples. As a result more purification procedures are required (Table 1)[14].
Conclusion
Removal of humic substances should be possible without loss of total RNA. Secondly the collection of RNA from all the soil types is required. It is also essential to simplify the extraction protocols as well as to develop new universal extraction protocol for all types of soil.
Acknowledgment
The first author is very thankful to Prof. Dr Asad Ali Department of Plant Pathology for his valuable suggestions.
References
- Vartoukian SR, Palmer RM, Wade WG (2010) Strategies for culture of 'unculturable' bacteria. FEMS Microbiol Lett 309(1): 1-7.
- Selenska S, Klingmuller W (1991) DNA recovery and direct detection of Tn5 sequences from soil. Lett Appl Microbiol 13(1): 21-24.
- Taniguchi T, Kataoka R, Tamai S, Yamanaka N, Futai K (2009) Distribution of bacterial species in soil with a vegetational change from Japanese black pine (Pinus thunbergii) to black locust (Robinia pseudoacacia). Microbes Environ 24(3): 246-252.?
- Wang Y, Morimoto S, Ogawa N, Fujii T (2011) A survey of the cellular responses in Pseudomonas putida KT2440 growing in sterilized soil by microarray analysis. FEMS Microbiol Ecol 78(2): 220-232.
- Wang Y, Morimoto S, Ogawa N, Oomori T, Fujii T (2009) An improved method to extract RNA from soil with efficient removal of humic acids. J Appl Microbiol 107(4): 1168-1177.
- Kreader CA (1996) Relief of amplification inhibition in PCR with bovine serum albumin or T4 gene 32 protein. Appl Environ Microbiol 62(3): 1102-1106.
- Goyal S, Sakamoto K, Inubushi K (2000) Microbial biomass and activities along an andosol profile in relation to soil organic matter level. Microbes Environ 15(3): 143-150.
- DeCoste NJ, Gadkar VJ, Filion M (2011) Relative and absolute quantitative real-time PCR-based quantifications of hcnC and phlD gene transcripts in natural soil spiked with Pseudomonas sp. strain LBUM300. Appl Environ Microbiol 77(1): 41-47.
- Cleaves HJ, Jonsson CM, Jonsson CL, Sverjensky DA, Hazen RM (2010) Adsorption of nucleic acid components on rutile (TiO2) surfaces. Astrobiology 10(3): 311-323.
- Hoshino YT, Matsumoto N (2007) DNA-versus RNA-based denaturing gradient gel electrophoresis profiles of a bacterial community during replenishment after soil fumigation. Soil Biology and Biochemistry 39(2): 434-444.
- Novinscak A, Filion M (2011) Effect of soil clay content on RNA isolation and detection and quantification of bacterial gene transcripts in soil using qRT-PCR. Appl Environ Microbiol 77(17): 6249-6252.
- Dreyfus M, Régnier P (2002) The poly (A) tail of mRNAs: bodyguard in eukaryotes, scavenger in bacteria. Cell 111(5): 611-613.
- Urich T, Lanzen A, Qi J, Huson DH, Schleper C, et al. (2008) Simultaneous assessment of soil microbial community structure and function through analysis of the meta-transcriptome. PloS One 3(6): e2527.
- Wang Y, Hayatsu M, Fujii T (2012) Extraction of Bacterial RNA from Soil: Challenges and Solutions. Microbes and Environments 27(2): 111121.






























