Study of Genetic Polymorphism of Cd18 Gene in Tunisian Holstein Cows
Jemmali B1*, Baccouche R2, Bouraoui R1, Kamon M1, Ben Gara1 and Rekik B1
1Development of Animal Productivity and Food Resources, University of Carthage, Tunisia
2Institut National Agronomique 43, Tunisie
Submission: January 28, 2017; Published: February 09, 2017
*Corresponding author: Jemmali B, 1Development of Animal Productivity and Food Resources, Development of Animal Productivity and Food Resources, Higher School of Agriculture of Mateur, University of Carthage, Tunisia, Tel: 0021672486074; Fax: 0021672485088; Email: bornijemali@yhoo.fr
How to cite this article: Jemmali B, Baccouche R, Bouraoui R, Kamon M, Ben G, Rekik B. Study of Genetic Polymorphism of Cd18 Gene in Tunisian Holstein Cows. Agri Res & Tech: Open Access J. 2017; 3(5): 555623. DOI:10.19080/ARTOAJ.2017.03.555623
Abstract
The Leukocyte cell-adhesion molecule common beta-chain (CD18) was considered as one of quantitative trait loci (QTL) related to mastitis resistance. Some punctual mutations were used for cow^s selection to improve their genetic potential for mastitis resistance. This study aimed to identify the presence-absence of point mutation at position 383pb. A total of 160 blood samples were collected from dairy breed Holstein localized in four governorates of Tunisia: Bizerte, Nabeul, Kairouan and Kasserine. The objective of this work is to study the polymorphism of the gene encoding CD18 by PCR-RFLP technique. CD18 is a glycoprotein that has an important role in the defenses of the mammary gland. PCR-RFLP revealed the presence of the restriction site for the enzyme TaqI encoding gene CD18. It has two different allelic variants A and B and three genotypes AA, AB, BB with a dominance of the allele A. The variability in the restriction profile confirms the existence of a point mutation at position 383pb. To address the problem of mastitis in dairy cattle herd, it is recommended to select normal homozygous animals carrying the mutated allele B.
Keywords: Genetic polymorphism; CD18; Dairy cows; Mastitis; Office of state lands (OTD)
Introduction
Mastitis is a disease that is the cause of many economic losses of dairy cattle in the world. In Tunisia the problem has increased following the introduction of dairy cattle aboveground. CD18 is a glycoprotein which is involved in the defense mechanisms of the mammary gland; this protein is regulated by a gene located on chromosome 1. The polymorphism of this gene is due to a point mutation (G:A) at 128pb position Gerardi, et al. [1]. As for Patel, et al. [2], it shows the existence of a silent point mutation (C: T) at 775pb position.
This study aims to analyze the genetic polymorphism of the CD18 in 160 Tunisia Holstein dairy cows by PCR-RFLP technique.
Animal material
Materials And Methods
The movements performed during this work hit four farms belonging to the Office of State Lands (OTD) and are located in four governorates namely; Nabeul, Kairouan, Bizerte, Kasserine (Table 1).
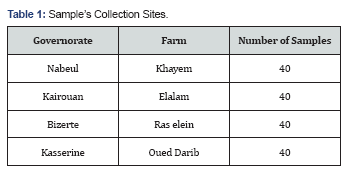
The total number of samples collected is 160 blood samples were taken from the jugular vein of Holstein dairy cows. Subjects were taken randomly. The amount of blood sample is 5 ml; the blood was placed in evacuated tubes containing K3EDTA and stored at -20 ° C for the genomic DNA extraction.
Extraction of genomic DNA
The method for the extraction of genomic DNA is that developed by Sambrook, et al. [3] based on the saline extraction.?
Estimation of the quantity and quality of the DNA
The verification of the quality of the genomic DNA was made by electrophoresis on agarose gel [0.8% (W / V)]. As for the estimation of the amount of DNA was made using the measurement of the optical density at a wavelength of 260 nm using a spectrophotometer or comparing to ladder quantity Jemmali, et al. [4].
Dilution of the DNA
This step allows for equal concentrations of DNA. The total DNA was diluted and the final chosen concentration in this work is 50 ng/^.
Development of PCR-RFLP technology
Development of the PCR reaction: Seyfert, et al. [5]
Preparation of primers: The lyophilized primers (Invitrogen) were dissolved in dilution buffer to give a final concentration of 100 pmol/μl.
Preparation of the reaction mixture: The reaction mixture consists of two primers, dNTPs, MgCl2, H2O, Taq polymerase and buffer PCR is performed with a final volume of 25μl. Reagents of the amplification solution are presented in (Table 2).
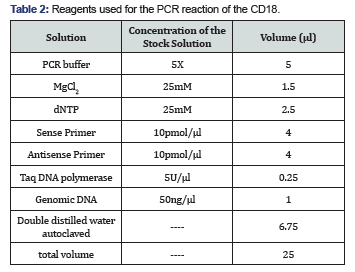
Implementation of a cyclic amplification: After some tests a suitable hybridization temperature was found. The initial denaturation was performed at 95°C for 5 min. 39 amplification cycles each cycle consisting of denaturation at 95°C for 30s, followed by primer annealing at 61 ° C for 30 s, elongation at 72 ° C for 45s, and then a final extension at 72°C for 10min.
Control amplification: To determine the efficiency and specificity of the amplification reaction, an electrophoresis on 2% agarose gel was performed.
Application of RFLP
Development of RFLP: After amplification, 8^l of the PCR product were digested by the enzyme in a reaction volume of 25mul. The reagents of the enzyme solution and their concentrations are described in (Table 3). Incubation lasts 24 hours at 65°C (in a water bath), restriction enzymes hydrolyze the PCR products.

Enzymatic digestion of control: This step is done by migration on a 2% agarose gel. 12μl of the digest were placed in each well. Each series comprises an undigested PCR product made in the first well. The last well contains the size marker in order to know the size of the digested products.
Allele Frequencies: F (A1) = (2n1 + n2)/2N = p
F (A2) = (2n 3 + n2)/2N = q
Genotype frequencies:
A1A1 = p2x N A1A2 = 2 pq xN A2A2= q2xN
The frequency of the A1 allele p is the proportion of the A allele among all genes of the population that may occupy the locus (A1, A2). Similarly, q is the proportion of alleles A2, since the locus (A1, A2) is necessarily occupied by either A1 or A2.
Results and Discussion
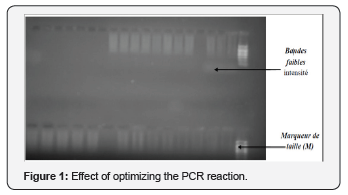
DNA fragments containing the CD18 gene were amplified via the PCR technique. The optimization of the PCR reaction affected several parameters including those related to the amount of DNA. Indeed, it happens that 1μ of DNA is sufficient to have better PCR- RFLP profiles, beyond that amount that is to say the number of bands 2μl decrease and primer annealing is inhibited therefore the choice of 1μ of DNA was made. Furthermore, the amount of Taq polymerase used is 1U/mu.l which prove to be an adequate amount. Following several trials, the amount of MgCl2 is selected from 1.5μl, which is the appropriate amount for the optimization. In addition, a change of the thermal program was created in order to improve the efficiency of amplification. 61°C is the temperature at which the amplification profiles are clearer It follows, therefore, this program has been considered for the PCR reaction (Figure 1).
The amplification results of the genomic DNA are positive for the majority of individuals tested, confirming the existence of the coding sequence for the CD18 gene (Figure 2). Indeed, electrophretic analysis of amplification products resulted in the stringent conditions, to a band of 367 bp in the study subjects which are consistent with results found by Patel, et al. [2], and band obtained are of high intensity, which allows the use of this protocol as a standard.

PCR-RFLP revealed the presence of the restriction site for the enzyme Taq I, of the CD18 gene. The locus of the gene encoding CD18 has two different allelic variants A and B and three genotypes AA, AB and BB. These results are similar to those found by Balcan, et al. [6], Patel, et al. [2] and Oner, et al. [7].
By comparing the allele and genotype frequencies found in literature, it appears that the normal homozygous animals were observed on the restriction profile by two bands (313 and 54 bp) and the BB genotype, and heterozygotes show three bands of different sizes (376, 313 and 54 bp).
The results obtained are similar to those found by Patel, et al. [2], and homozygous affected animals are presented with a single intact band of 367 bp and this finding is consistent with that found by Oner, et al. [7]. The variability in the restriction profile highlights the existence of a point mutation at position 383 in the genomic DNA, confirming the hypotheses of Viana, et al. [8]. The electrophoresis profile on agarose gel of PCR products, following digestion with the enzyme Taq I was shown by the (Figure 3).
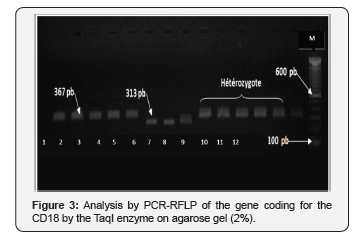
The line M is a molecular weight marker (100 bp), the wells: 1, 2, 3, 4, 5 = the PCR products (367 bp). Wells: 9, 10, 11, 12 = the genotype AB (367et 313 bp and 54 bp). The wells: 6, 7 = BB genotype (313, 54 bp).
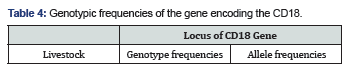

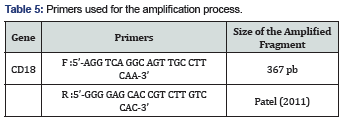
The Table 4 below shows the frequencies of alleles and the sizes of the different populations. At the locus of the gene coding for the CD18 which has two alleles A and B, the dominance of the allele is remarkable and its frequency is 0.92. Our results are similar to Patel, et al. [2], hence the need to select normal animals homozygous carriers of the mutated allele B (Table 5).
Conclusion
The study of polymorphism of the gene encoding CD18 showed that the majority of the test animals are carriers of the gene, after PCR amplification using appropriate primers defining the desired segment to be amplified (376 bp). The TaqI restriction enzyme, the cutting site specific levels and produces a mixture containing two alleles A and B, frequency 96% and 4%. These two alleles control the appearance of the three genotypes AA, AB and BB. Variability encountered levels of digestion of PCR-RFLP profile confirms the existence of a point mutation in the genomic DNA. To address the problem of mastitis in dairy cattle herds it is recommended to select normal animal's homozygous carriers of the mutated allele B.
References
- Gerardi AS (1996) Bovine leukocyte adhesion deficiency: a review of a modern disease and its implications. Res Vet Sci 61(3): 183-186.
- Patel M, Patel RK, Singh KM, Rank DN, Thakur MC, et al. (2011) Detection of genetic polymorphism in CD18 gene in cattle by PCR- RFLP. Wayamba J Anim Sci 3: 110- 111
- Sambrook J, Fritsch EF, Maniatis T (1989) Molecular Cloning. A Laboratory Manual. (2nd edn), Cold Spring Harbor Laboratory Press, USA, pp. 1626.
- Jemmali B, BenGara A, Selmi H, Ammari Z, Bouheni C, et al. (2013) Kappa casein gene polymorphism in Tunisian local goat. Pak J Biol Sci 16(24): 2031-2035.
- Seyfert HM, Luthen F (1998) The structure of the bovine butyrophilin encoding gene differs grossly from mouse concerning promoter localization and exon organization of the 5- untranslated region. Proceedings of the 6th World Congress on Genetics Applied to Livestock Production 25: 51-54.
- Balcan RA, Manea MA, Georgescu SE, Dinischiotu A, Tesio CD, et al. (2007) Evidence of single point mutation inducing BLAD disease in Romanian Holsteinderived cattle breed. Biotechnol Anim Husb 23(5- 6): 375-381.
- Oner Y, Keskin A, Elmacia A (2010) Identification of BLAD, DUMPS, Citrullinemia, Facor XI deficiency in Holstein Cattle in Turkey. Asian Journal of Animal and Veterinary Advance 5(1): 60-65.
- Viana JL, Fernandez A, Iglesias A, Santamarina G (1998) Diagnóstico y control de las principales enfermedades genéticas (citrulinemia, DUMPS y BLAD) descritas en ganado Holstein-Frisón. Med Vet 15(10): 538-544.






























