Analysis of Genetic Variability and Correlation in Fennel (Foeniculum Vulgare Mill.) Germplasm
Rohit Kumar, RS Meena, AK Verma, Hemant Ameta and Alka Panwar*
ICAR-National Research Centre on Seed Spices, India
Submission: October 29, 2016; Published: January 23, 2017
*Corresponding author: Alka Panwar, ICAR-National Research Centre on Seed Spices, Beawar Road, Ajmer, Rajasthan 305006, India, Email: simiran.saini@rediffmail.com
How to cite this article: Rohit K, R Meena, A Verma, Hemant A, Alka P. Analysis of Genetic Variability and Correlation in Fennel (Foeniculum Vulgare Mill.) Germplasm. Agri Res & Tech: Open Access J. 2017; 3(4): 555616.DOI: 10.19080/ARTOAJ.2017.03.555616.
Abstract
Fifty germplasm were used to study the genetic variability, heritability, genetic advance and correlation for growth and yield contributing characters in fennel. Experiment laid out in an Augmented Block Design at National Research Centre on Seed Spices, Ajmer for yield and its yield attributing characters in 2014-15. The analysis of variance revealed significant differences among the germplasms for number of primary branches, number of umbels per plant, number of umbellate per umbel, number of seed per umbellate, test weight (g) and seed yield (5 plant g). The phenotypic coefficient of variance (PCV) was higher than genotypic coefficient of variance (GCV) for most of the characters. Nnumber of umbels per plant, number of umbels per umbellate per umbel, number of seed per umbellate, test weight, seed yield and number of secondary branches exhibited high genetic advance as percentage of mean along with high heritability. Nnumber of primary branches (0.75***), number of secondary branches (0.63***), umbel per plant (0.87***), umbellate per umbel (0.63***), seeds per umbellate (0.70***) and test weight (0.52***) exhibited positive and significant correlated with the seed yield. Therefore, greater emphasis should be given on these characters for genetic improvement of fennel.
Keywords: Variability; Heritability; Genetic advance; Correlation; Fennel
Introduction
Fennel (Foeniculum vulgare Mill.), belongs to family Apiaceae. It is a diploid species with chromosome number, 2n=22 and native of Europe and Mediterranean region Agarwal et al. [1]. It's mainly cultivated for its seeds in the state ofRajasthan and Gujarat. The leaves and seeds of fennel are used in many culinary traditions Ehsanipour et al. [2]. Mature fennel fruits and essential oil are used as flavouring agents in food products such a liqueurs, bread, pickles, pastries, and cheese Zoubiri et al. [3]. The root is regarded as a purgative. Fennel fruits are used in diseases like cholera, bile disturbances, nervous disorders, constipation, dysentery and diarrhea and also used for control of diseases attacking chest, lungs, spleen, and kidney and in colic pain. Since most of the yield attributing characters are quantitatively inherited and highly affected by environment it is difficult to judge whether the observed variability is heritable or not. The starting point of any systemic breeding programs is the collection of a large germplasm. Furthermore, information on association among different morphological characters and with seed yield is necessary for formation of suitable selection criteria for producing high yielding varieties.The volatile oil is used in the manufacture of cordials and enters into the composition of fennel water, which in commonly given to infants as medicine.
Keeping this in view, the present investigation was made to explore the genetic variability, by determining the magnitude of genetic coefficient of variation, heritability estimates and expected genetic advance of different biometric traits, their correlation and effects in 54 fennel genotypes including checks.
Materials and Methods
Fifty four germplasm of fennel include checks were evaluated in Augmented Block Design at National Research Centre on Seed Spices, Tabiji, Ajmer (Rajasthan). The centre lies on 74° 35' 39” E to 74° 36' 01” longitude and 26° 22' 12” to 26° 22' 31" N latitude at an altitude of 460.17 m above mean sea level. In each single line genotype were sown in a plot of 3m x 2m size accommodating 6 row of 2 m length spaced 50 cm maintained. All the recommended package of practices was followed to raise a good crop. Five competitive plants were marked in each single line and observations were recorded on these plants for days to germination, days to 50% flowering, king umbel anthesis, plant height (cm), number of primary branches, number of secondary branches, number of seed per umbellate, number of umbel per plant, number of umbellate per umbel, king umbel diameter, test weight (g) and seed yield g (5 plants). Analysis of variance [4]. Genotypic coefficients of variation (GCV) were computed was done by the method suggested by Panse & Sukhatme according to Burton [5].
Results and Discussion
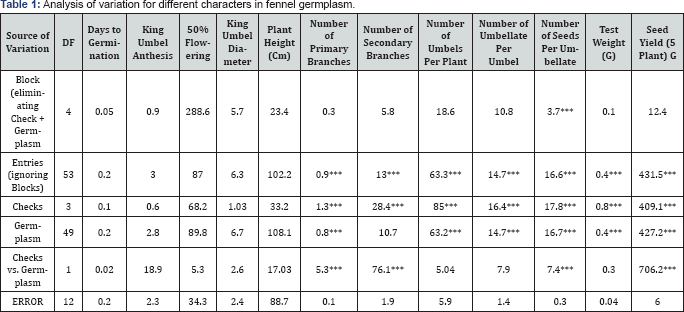
*** Significant at p =0.001
Analysis of variance indicated significant for these characters are given viz., number of primary branches, number of umbels per plant, number of umbellate per umbel, number of seed per umbellate, test weight (g) and seed yield (5 plant g) was found significant (Table 1). The observed general mean for plant height was (180.7) and ranged from 160 to 201.8, seed yield (5 plant g) (146) and ranged from 112.5 to 178.1 while 50% flowering (114.4) and ranged from 102-132 (Table 2).
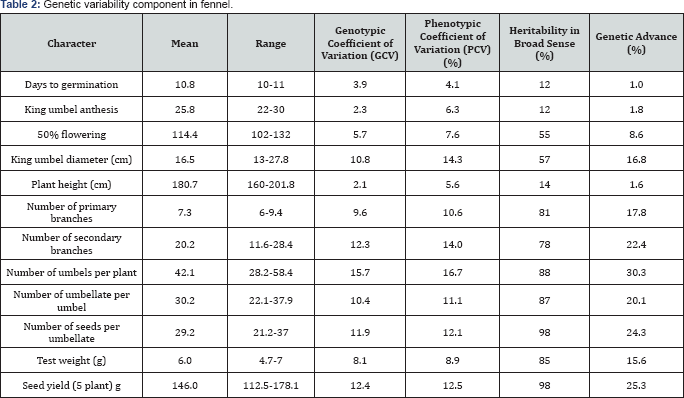
The extent of variability present in 50 germplasm of fennel was measured in terms of range, mean, PCV, GCV, heritability in broad sense and genetic advance. All the germplasm differed significantly with respect to different characters studied. Wide range of variation was observed in all the characters. Munshi & Behra [6], Warshamana et al. [7] and Gupta et al. [8] also reported wide range of variation for most of the characters.
The GCV and PCV were high for number of umbels per plant (15.7 & 16.7) and number of secondary branches (12.3 & 14), number of umbellate per umbel (10.4 &11.1) and number of primary branches (9.6 & 10.6) indicating greater diversity for these traits. GCV in general, were lower than the PCV which indicated close association between phenotype and genotype. These results are in agreement with those reported by Munshi & Behra [6], Singh et al. [9] and Gupta et al. [8]. Low GCV and PCV were recorded for test weight (8.1 & 8.9) and Kumar et al. [10], 50 % flowering (5.7 & 7.6).These results are in conformity with the findings of Singh et al. [9] and Samadia [11].
Heritability is parameter of tremendous significance to the breeders as its magnitude indicates the reliability with which a genotype can be recognized through its phenotypic expression. Johnson et al. [12] stresses that for estimating the real effect of selection, heritability estimates along with genetic advance are more meaningful. Heritability in broad sense was observed to be high for all the traits studied. High heritability estimates were also reported earlier by Verma et al. [13] and Samadia [11]. Heritability estimates alone are not an ideal parameter for predicting the effect of selecting the desired individual. Heritability estimates alone with genetic advance are more useful than heritability values alone in predicting the selection of best individuals. In the present investigations number of umbels per plant, number of umbels per umbellate per umbel, number of seed per umbellate, test weight, seed yield and number of secondary branches exhibited high genetic advance as percentage of mean along with high heritability. These results indicated the influence of additive gene action. Similar results were also recorded earlier by Dashora et al. [14], Jain et al. [15], Shukla et al. [16], Lal [17], Dashora & Sastry [18], Abou [19].
The data on correlation showed that most of the correlation coefficients at genotypic level were greater than the corresponding phenotypic correlation coefficients. This suggested the predominate ofgenotypic correlation of nine yield and yield attributing characters presented (Table 3) indicated thatseed yield was positively correlated with number of primary branches (0.75***), number of secondary branches (0.63***), umbel per plant (0.87***), umbellate per umbel (0.63***), seeds per umbellate (0.70***) and test weight (0.52***).
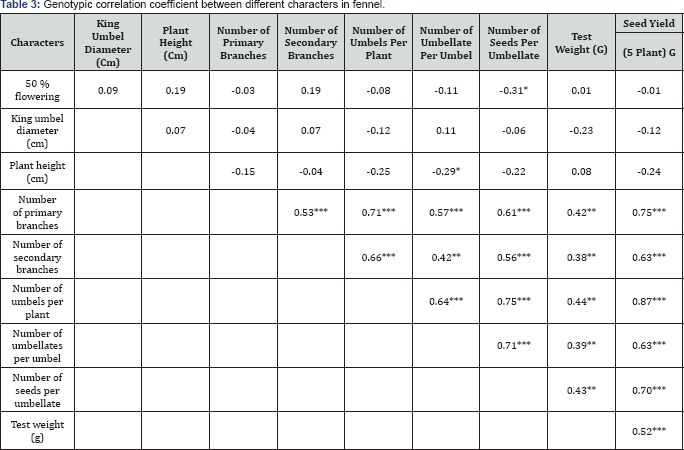
*** Significant at p =0.001 ** Significant at p =0.01 * Significant at p =0.05
50 % flowering was found significant and negative correlated with number of seeds per umbellates (-0.31), while it was also found non-significant but positively correlated with king umbel diameter (0.09), plant height (0.19), no of secondary branches (0.19) and test weight (0.01). Diameter of king umbel was found non-significant positive correlated with plant height (0.07), number of secondary branches (0.02) and no. of umbellate per umbel. Plant height (cm) had found significant and negative correlation number of umbellates per umbel (0.29), while it was also found non-significant but positively correlated with test weight (0.08). Primary branches had found significant and positive correlation with number of secondary branches per plant (0.53), number of umbels per plant (0.71), number of umbellate per umbel (0.57), number of seeds per umbellates (0.61), test weight (0.42) and seed yield (gm) (0.75). Number of secondary branches per plant had found significant and positive correlation with number of umbels per plant (0.66), number of umbellate per umbel (0.42), number of seeds per umbellates (0.56), test weight (0.38) and seed yield (gm) (0.63). Number of umbels per plant had also recorded significant and positive correlated with number of umbellate per umbel (0.64), number of seeds per umbellates (0.75), test weight (0.44) and seed yield (gm) (0.87). Number of umbellates per umbel was found significant and positively correlated with number of seeds per umbellates (0.71), test weight (0.39) and seed yield (gm) (0.63). Number of seeds per umbellates had also recorded significant positive correlated with test weight (0.43) and seed yield (gm) (0.70).
The association analysis at both genotypic and phenotypic level revealed that the seed yield was significantly and positively correlated with umbel per plant, number of primary, number of secondary branches, number of umbellates per umbel and number of seeds per umbellates. Similar results were found in the findings of Meena et al. [20], Dashora & Sastry [18], Meena et al. [21]. While the correlation of test weight showed positive but non-significant correlation with seed yield. Similar result was found in the findings of Abou [19], Yogi [22-26].
References
- Agarwal AK, Hatfield J, Kemery E, Valenti S, Esnault L (2001) Web- based education: Changing the equilibrium in managing information technology in a global economy. Proceedings of the IRMA International Conference, Idea Group Publishing: Hershey London Melbourne Singapore, Toranto, Canada, pp. 1197.
- Ehsanipour A, Razmjoo J, Zeinali H (2012) Effect of Nitrogen Rates on Yield and Quality of Fennel (Foeniculumvulgare Mill.) Accessions. In-dustrial Crops Products 35(1): 121-125.
- Zoubiri S, Baaliouamer A, Seba N, Chamouni N (2014) Chemical Composition and Larvicidal Activity of Algerian Foeniculum vulgare Seed Essential Oil. Arab J Chem 7(4): 480-485.
- Panse VG, Sukhatme PV (1985) Staistical Methods, for Agricultural Workers (2nd edn), ICAR, New Delhi, India, pp. 361.
- Burton GW (1952) Quantitative inheritance of grasses. Proceeding of 6th International Grassland Congress 1: 227-283.
- Munshi AD, Behera TK (2000) Genetic variability, heritability and genetic advance for some traits in chilli. (Capsicum annuum L.). Veg Sci 27: 39-41.
- Warshamana IK, Vikram A, Kohli UK (2008) Genetic evaluation for physicochemical and yield traits in chilli (Capsicum annuum L.) under hilly conditions of Himachal Pradesh. The Hort J 21: 62-66.
- Gupta AM, Singh D, Kumar A (2009) Genetic variability, genetic advance and correlation studies in chilli (Capsicum annuum). Indian J Agri Sci 79: 221-223.
- Singh MD, Laisharam JM, Bhagirath T (2005) Genetic variability in local chillies (Capsicum annuum L.) of Manipur. Indian Journal of Horticul-ture 62(2): 203-205.
- Kumar K, Baswana KS, Partap PS (1999) Genetic variability and heri- tability studies in chilli (Capsicum annuum L.). Haryana J Hort Sci 28: 125-126.
- Samadia DK (2007) Genetic variability studiesin chilli germplasm under hot arid ecosystem. Indian J Hort 64: 477-479.
- Johnson HW, Robinson HF, Comstock RE (1955) Estimates of genetic and environmental variability in soyabean. Agronomy Journal 47(7): 314-318.
- Verma SK, Singh RK, Anja RR (2004) Genetic variability and correlation studies in chilli. Prog Hort 36: 113-117
- Dashora A, Sharma RK, Sastry EVD, Singh D (2002) Varietal diallel analysis for yield and yield related traits in fennel (Foeniculum vulgare Mill.) Journal of Spices and Aromatic Crops 11(1): 18-21.
- Jain UK, Singh D, Jain SK (2002) Assessment of genetic variability in coriander. Annals of plant and soil Research 4(2): 329-330.
- Shukla S, Singh PK, Garg VK (2003) Study on genetic variability and heritability in fennel grown on sodic soils. Journal of Medicinal and Aromatic Plant Sciences 25(4): 956-958.
- Lal RK (2007) Association among agronomic traits and path analysis in fennel (Foeniculum vulgare Mill.). Journal of Sustainable Agriculture 30(1): 21-29.
- Dashora, Abhay, Sastry EVD (2011) Variability, character association and path coefficient analysis in fennel. Indian Journal of Horticulture 68(3): 351-356.
- Abou El-Nasr THS, Sherin A, Mahfouze Al Kordy MAA (2013) Assessing Phenotypic and Molecular Variability in Fennel (Foeniculum vulgare) Varieties. Australian Journal of Basic and Applied Sciences 7(2): 715722.
- Meena RS, Kakani RK, Anwer MM, Panwar Alka (2010) Variability of some morphological characters in fennel (Foeniculum vulgare). Indian Journal of Agricultural Sciences 80(8): 710-712.
- Yogi R, Meena RS, Kakani RK, Panwar A, Solanki RK (2013) Variability of some morphological characters in fennel (Foeniculum vulgare Mill.). International Journal of Seed Spices 3(1): 41- 43.
- Agnihotri P, Dashora SL, Sharma RK (1997) Variability, corrilation and path coefficientanalysis in fennel (Foeniculumvulgare Mill). Journal of Spices and Aromatic Crops 6(1): 51-54
- Dewey DR, Lu KH (1959) A correlation and path coefficient analysis of components of crested wheat grass seed production. AJ 51(9): 515518.
- Meena SK, Singh Bhuri, Meena AK (2013) Variability in fennel (Foenic- ulum vulgare Mill.) for yield and yield attributes. Indian Research Journal Genetics & Biotechnology 5(2): 117-124.
- Rajput SS, Singhania DL, Singh D, Sharma KC, Rathore VS (2004) Assessment of genetic variability in fennel (Foeniculumvulgare Mill.) germplasm. In: NationalSeminar on New Perspectives in Commercial Cul- tivation, Processing and Marketing Seed Spices and Medicinal Plants, p.25-26
- Yadav PS, Pandey VP, Yadav Y (2013) Variability studies in fennel (Foeniculum vulgare Mill.). Journal of Spices and Aromatic Crops 22(2):203-208.






























