Numerical Investigation of Polymerase Chain Reaction Design: Microfluidic Chip Simulation, Liquid Cooling Optimization, and Buoyancy-Driven Amplification
Mohammad Yaghoub Abdollahzadeh Jamalabadi*
Department of Microbiology. Biological Faculty, Moscow State University them MV Lomonosov. 119234. Lenin’s Hills, 1, 12. Moscow, Russia
Submission:February 10, 2026; Published: February 19, 2026
*Corresponding author:Mohammad Yaghoub Abdollahzadeh Jamalabadi, Department of Marine Engineering, Chabahar Maritime University, Chabahar, Iran
How to cite this article:Mohammad Yaghoub Abdollahzadeh Jamalabadi. Numerical Investigation of Polymerase Chain Reaction Design: Microfluidic Chip Simulation, Liquid Cooling Optimization, and Buoyancy-Driven Amplification. Adv Biotech & Micro. 2026; 19(2): 556010.DOI:10.19080/AIBM.2026.19.556010
Abstract
Humans have successfully landed on both Moon as well as Mars under very extreme environmental situations, not favorable for pedogenic or photopedogenic processes for soil development as normally possible on Earth. However, almost like Earth, basalt and other similar rocks and their small pieces are abundantly available on Moon and Mars surfaces. Such broken dust materials both from Moon and Mars were brought to Earth by giving a common name “regolith” (<2mm size, mm) for simulation studies on the Earth. More emphasis was given to basalt, since basalt is more generative through its mode of weathering under extreme environment existing on Moon and Mars surfaces besides possible scope with microbial communities including lichens. It is not a surprise for a soil to be developed when it’s parent materials, associated climate, sunlight, surrounding atmosphere, roughness of surface (topography) including vegetative cover (organic matter) and biodiversity are present. On moon and mars, the rocks dominated by basalt are present on both. However, the physical as well as environmental conditions are very distinct and extreme for pedogenic weathering. The existence of microorganism especially lichens may tolerate harsh conditions on Mars. Lichens being phototropic require sunlight for photosynthesis and the weak sunlight on Moon and Mars. Lichens are symbiotic organisms composed of fungi and algae or cyanobacteria. The existence of lichens on basalt or other mineral surface lead to promote organic matter and carbon and enhance possibilities of weathering of rocks and minerals. These challenges lie on survival and weathering of basalt surface. Such weathering following oxidation of basaltic minerals changes basalt from dark grey or black to yellowish or reddish or brownish. The result of weathering of basalt on colour may be yellowish-brown or red identical to the common colour of honey. The so developed soils on Mars or even on lunar surface to look like the colour of honey as such the actual soil especially on Mars in days to come would referably resemble the colour of honey. As such the proposed soil would be designated as Hanisols under WRB, although such tentative nomenclature would be verified in days to come when such Hanisols would be studied on Mars and Mon exhaustively. Artificial Intelligence (AI) unlocks the mystery of possible soil genesis on Moon and Mars.
Keywords: Lunar Soils; Martian Soils; Photopedogenesis; Harsh Environment; Hanisols; Basalt; Lichens; Artificial Intelligence
Abbreviations: IARI: Indian Agricultural Research Institute; BAU: Bihar Agricultural University; NRL: Nuclear Research Laboratory; NASA: National Aeronautics and Space Administration; UNO: United Nations Organisation; WRB: World Reference Base for Soil Resources
Introduction
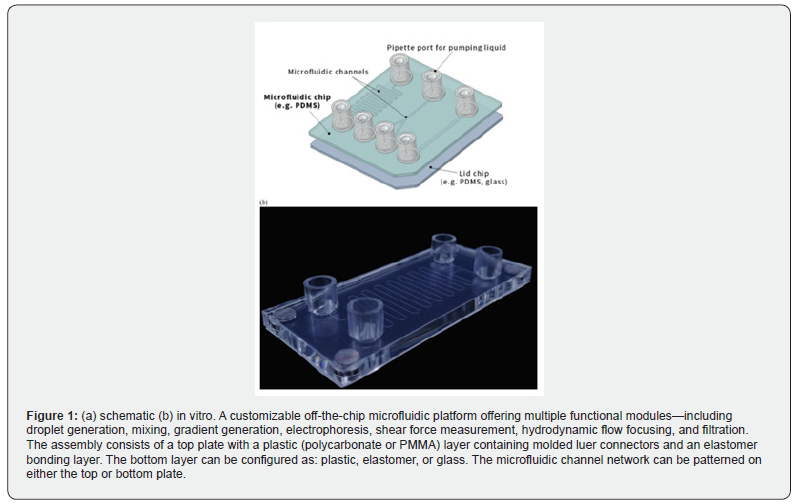
Molecular diagnostic methods have long served as foundational tools in clinical and industrial analysis. Among these, the polymerase chain reaction (PCR) stands out as a particularly sensitive and specific technique capable of amplifying minute quantities of target DNA from complex biological matrices. First conceptualized by Kary Mullis in 1983, PCR operates by cycling through three thermal phases-denaturation, annealing, and extension-that collectively produce exponential copies of a defined nucleic acid sequence [1,2]. Its ability to detect extremely low template concentrations confers a sensitivity that substantially exceeds conventional immunoassay approaches such as ELISA [3,4]. The molecular basis of PCR lies in the selective enzymatic amplification of specific nucleic acid sequences. The process exploits the thermal stability of Taq DNA polymerase to replicate a defined region of the target genome across repeated denaturation, annealing, and extension cycles. PCR is designed to amplify pathogen-specific DNA segments that are conserved across target organisms, allowing for both detection and species differentiation through carefully engineered oligonucleotide primers. The amplified products can subsequently be analyzed via gel electrophoresis or through real-time fluorescence-based quantification, enabling sensitive and specific identification of nucleic acids even at very low concentrations [5,6] (Figure 1).
Despite its analytical power, conventional PCR instruments rely on air-based thermal management systems-typically small fans and heat sinks-to regulate sample temperatures between cycle steps. While functional, these systems impose practical limitations: the comparatively low thermal conductivity of air (approximately 0.0242 W/m·K) restricts the maximum heat removal rate, resulting in cooling steps that can require 20 seconds or more per cycle [7,8]. Over a standard 35-cycle amplification run, this inefficiency accumulates into several minutes of excess processing time. Moreover, the adoption of PCR in field settings remains constrained by its dependence on sophisticated benchtop instrumentation, trained personnel, reliable cold chains for reagent storage, and uninterrupted power supply [9,10]. Microfluidic technology offers a compelling avenue to address these limitations by miniaturizing and integrating multiple analytical functions-including sample preparation, nucleic acid amplification, and detection-onto a compact chip platform [11,12]. Such devices are capable of reducing reagent consumption, shortening processing times, and enabling operation by non-specialist users in decentralized settings. The primary objective of microfluidic PCR chip design is to emulate the performance of conventional PCR assays while significantly reducing device footprint and operational complexity [13,14].
Liquid cooling offers a complementary enhancement. Water and other coolants conduct heat far more effectively than air, and when channeled through microfabricated passages-socalled microchannel heat sinks-they can exploit exceptionally high surface-area-to-volume ratios to extract heat rapidly from compact substrates [15,16]. Previous investigations have explored numerous parameters that govern microchannel thermal performance, including channel patterning, inlet/outlet manifold design, cross-sectional geometry, and the use of enhanced working fluids such as nanofluids or phase-change materials [17,18]. A third strategy for PCR miniaturization is natural convectionbased amplification, in which temperature-induced density differences drive fluid circulation without requiring an external pump. In this approach, a PCR mixture cycles between hot and cold regions within a closed microreactor via buoyancy-driven flow, eliminating the need for active flow control systems [19,20].
The present work integrates three complementary computational investigations: (1) the design and simulation of a miniaturized microfluidic PCR chip using ANSYS CFD, with systematic evaluation of channel geometry and material selection; (2) a numerical comparison of fifteen microchannel cross-sectional geometries for PCR liquid cooling using a performance evaluation criteria (PEC) index that balances heat transfer enhancement against pressure-drop penalties; and (3) a buoyancy-driven natural convection PCR model using COMSOL Multiphysics that tracks spatio-temporal variation of DNA component concentrations. Together, these studies provide a unified computational framework for PCR platform design and thermal management. This work builds upon an expanding body of literature on microfluidic diagnostic platforms. Prior studies have explored microfluidics-based detection of foodborne pathogens [21], molecular diagnostics for infectious diseases [22], ferro-microfluidic cell separation systems [23,24], and organ-onchip modeling [25]. The present work extends this foundation by addressing chip geometry, material selection, cooling optimization, and reaction kinetics within a single integrated framework.
A review of the recent studies reveals distinct yet interconnected research themes, primarily focusing on microfluidic device design [26-35], fabrication [36-40], and application (particularly for PCR and biosensing) [41-50], heat transfer optimization in micro-scale systems [51-55], and molecular detection techniques [56-60]. In the domain of micro-scale heat transfer and fluid dynamics, computational and experimental studies extensively investigate the optimization of microchannel and micro pin fin geometries for enhanced thermal performance. Research consistently shows that geometrical parameters-such as cross-sectional shape, aspect ratio, and configuration-profoundly influence thermohydraulic characteristics [28, 29, 30, 31, 41]. Optimization frameworks, including constructal theory, are applied to determine optimal geometries for microchannel arrays [42]. Advanced designs, such as microchannels with oblique grooved fins, are proposed to further improve performance by inducing secondary flows [43]. The internal physical properties, like viscosity, also affect molecular dynamics in these flows, as studied in the context of DNA [27]. Numerical simulations using tools like ANSYS are fundamental to this field, applied not only to heat exchangers [34] but also to structural analyses like gear optimization [33].
For molecular detection, particularly PCR, the literature covers both foundational techniques and advanced microfluidic integration. Early work established sensitive pathogen DNA detection via real-time quantitative PCR [26]. The performance of conventional PCR can be influenced by factors such as amplicon size [59] and thermal cycler efficiency, which requires monitoring [60]. Recent advancements focus on integrating PCR into microfluidic platforms to create compact, efficient diagnostic systems. This includes convection–reaction-diffusion models for PCR in natural convection systems [32], electromagneticallydriven platforms for virus detection (e.g., SARS-CoV-2 via RTLAMP) [44], and highly efficient photonic PCR using plasmonic heating from nanomaterials [48,49]. Digital and droplet-based quantification methods offer high precision, as seen in droplet digital PCR chips with passive bubble removal [35] and pumpfree duplex droplet digital LAMP chips [50]. Microfluidic chip fabrication and material science are critical for enabling these applications. Common substrate materials include polymers like polypropylene [36] and polycarbonate (PC). PC’s susceptibility to hydrolysis is a known consideration [37], but it can be modified for chip bonding and hydrophobicity [38].
Polydimethylsiloxane (PDMS) is another ubiquitous material, often bonded via oxygen plasma treatment, with studies investigating its wettability and bonding mechanism [40] or its machining with CO2 lasers [39]. Comparative assessments evaluate materials like silicon, glass, FR4, PDMS, and PMMA for specific functions such as acoustic manipulation [57]. Fabrication strategies aim for low-cost MEMS technologies [56], and chips are functionalized for specific uses, such as creating hydrophobic surfaces for liquid phase microextraction [51] or fabricating composite chips (e.g., hydroxyapatite-PDMS) for cell culture and drug screening [58]. Finally, integrated microfluidic biosensors represent a major application frontier. These devices often combine sample preparation (e.g., filtration and immunomagnetic separation [45]) with detection for foodborne pathogens [45,52]. Detection leverages various strategies, including virulence genebased PCR for E. coli [46], CRISPR/Cas12a-assisted PCR for Vibrio parahaemolyticus [47], and dual-aptamer assays for Bacillus cereus [52]. For clinical diagnostics, sophisticated chips enable pump-free SERS amplification for lung cancer ctDNA detection [53], virus isolation via cell screening [54], and integrated analysis of exosomal RNA for lung cancer diagnosis [55]. These references collectively illustrate the evolution from fundamental studies to highly integrated, application-specific microfluidic systems.
The novelty of this work lies in its integrated, multi-faceted
computational approach that simultaneously addresses three
critical and often separately studied aspects of microfluidic PCR
design within a single, cohesive framework. While previous
studies have typically focused on isolated components-such as
chip geometry, cooling optimization, or reaction modeling-this
research uniquely combines:
a) Chip-Level Design with Material Screening: A
systematic simulation-based comparison of looped vs. straight
channel geometries, paired with an evaluation of polymeric
materials (PP, PC, PDMS) for thermal performance, leading to the
specific recommendation of polypropylene for optimal thermal
fidelity.
b) Microchannel Cooling Optimization Across Multiple
Geometries: A comprehensive numerical comparison of fifteen
distinct microchannel cross-sections under consistent operating
conditions, identifying the circular geometry as optimal through
a performance evaluation criteria (PEC) index that balances heat
transfer enhancement against pressure-drop penalties-a direct
and practical design guideline not commonly consolidated in PCRfocused
studies.
c) Buoyancy-Driven Reaction Modeling Integrated
with Design: The development and validation of a convectionreaction-
diffusion model for natural convection PCR, explicitly
linking temperature-induced flow to DNA amplification kinetics
and quantifying a doubling time (~60 seconds) consistent with
experimental benchmarks-thereby bridging fluid dynamics with
biochemical process validation.
By unifying chip design, active cooling optimization, and passive convection-driven amplification into one study, this work provides a holistic design toolkit that enables performance trade-off analysis and co-optimization-a step forward toward the development of compact, efficient, and field-deployable PCR systems.
The overarching goal of this work is to establish a cohesive computational framework for the rational design and optimization of microfluidic PCR platforms. To achieve this, the study is structured into three interconnected parts. First, the design and thermal-fluidic performance of a miniaturized PCR chip are analyzed, with a focus on channel geometry and material selection. Second, a systematic optimization of microchannel cross-sectional shapes is conducted to enhance liquid cooling efficiency. Third, a buoyancy-driven natural convection model is developed to validate PCR amplification kinetics without external pumping. Together, these sections provide a comprehensive approach-from chip architecture and thermal management to reaction modeling-culminating in a set of design guidelines and performance benchmarks for next-generation microfluidic PCR systems.
Microfluidic PCR Chip Design and Simulation
This section presents a framework for the preliminary design and physical understanding of the proposed microfluidic PCR device. The analytical model serves three main purposes: (i) providing closed-form relations for key design parameters, (ii) guiding the selection of geometric and operational ranges prior to numerical simulations, and (iii) validating the CFD results presented in subsequent sections. The flow within the microfluidic channel is modeled under the following assumptions: steady, incompressible flow; Newtonian fluid behavior; laminar regime with low Reynolds number; no-slip boundary condition at all solid walls; and rectangular microchannel geometry. The governing momentum equation under these conditions reduces to the Stokes equation: ∇p = μ ∇²u. For a rectangular microchannel satisfying h ≪ w (shallow channel approximation), the volumetric flow rate is given by the Hele-Shaw relation:
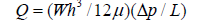
where w is the channel width, h is the channel height, μ is the dynamic viscosity of the PCR mixture, Δp is the applied pressure drop, and L is the channel length. The cross-sectional average velocity follows directly: ū = Q / (w h). Temperature distribution within the chip is governed by the energy equation, which in its general unsteady form reads:
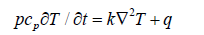
where ρ is the fluid density, cₚ is the specific heat capacity, k is the thermal conductivity of the chip material, and q̇ is the volumetric heat generation rate from the heater plates. Under steady-state conditions and for one-dimensional heat conduction along the channel length, the energy equation simplifies to: k d²T/ dx² + q̇ = 0. The general solution yields the parabolic temperature profile:
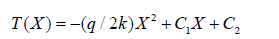
where C₁ and C₂ are integration constants determined by the imposed wall temperatures. This solution directly underpins the relative deviation analysis: a material with higher thermal conductivity k produces a flatter T(x) profile, reducing temperature non-uniformity and improving PCR fidelityconsistent with the superior performance of polypropylene (k = 0.80 W/m·K) over polycarbonate (k = 0.24 W/m·K) and PDMS (k = 0.15 W/m·K).
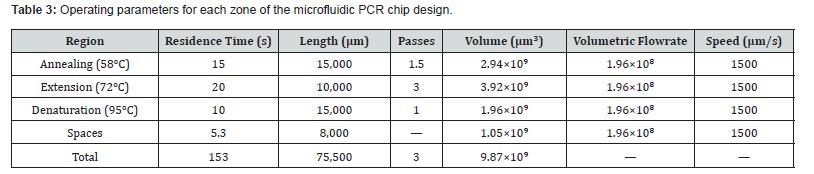
A critical design constraint is that the fluid must remain within each thermal zone long enough for the corresponding PCR reaction step to complete. The residence time in each zone is: tᵣ = L / ū. The design condition tᵣ ≥ tₚᶜᵣ must be satisfied for each zone, where tₚᶜᵣ is the minimum required reaction time for denaturation (10 s), annealing (15 s), and extension (20 s) as specified in Table. Using the design velocity of ū = 1500 μm/s and the zone lengths defined herein, the computed residence times confirm that the channel dimensions and flow velocity satisfy the PCR timing requirements for all three stages. Two dimensionless groups govern the physics of the device. The Reynolds number characterizes the ratio of inertial to viscous forces: Re = ρ ū Dₕ / μ, where Dₕ = 2wh/(w+h) is the hydraulic diameter. For the design parameters employed, re ≈ 4.5 × 10⁻´, confirming fully laminar, viscosity-dominated flow throughout the chip. The Péclet number characterizes the ratio of convective to diffusive thermal transport: Pe = ρ cₚ ū L / k. For the polypropylene chip with L = 15,000 μm, Pe ≈ 50.6, indicating that streamwise heat advection substantially exceeds axial conduction. This confirms that interzone thermal cross-contamination is negligible under normal operating conditions.
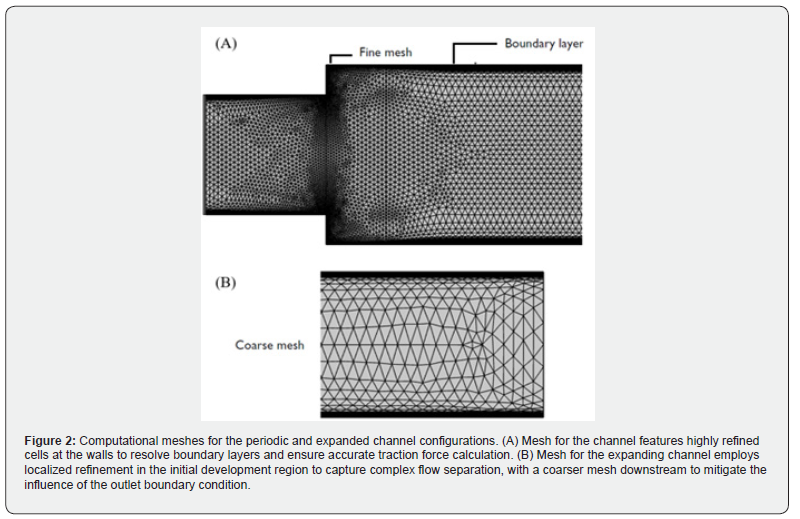
The computational meshes employed for the two validation geometries are illustrated in (Figure 2). For the periodic channel configuration (Figure 2A), near-wall mesh refinement is implemented to accurately resolve the boundary layer and capture the steep velocity and stress gradients that develop adjacent to the channel walls. This refinement is essential for the precise calculation of wall traction forces. For the expanding channel configuration (Figure 2B), the mesh is selectively refined in the initial development region immediately downstream of the geometric expansion, where flow separation, recirculation zones, and elevated shear layers demand high spatial resolution. A progressively coarser mesh is employed toward the channel outlet, which serves as a buffer region to mitigate the influence of the imposed outlet boundary condition on the upstream flow field of interest. In both cases, the mesh topology reflects a physicsdriven design strategy: resolution is concentrated where flow gradients are steepest, ensuring computational accuracy while minimizing unnecessary overhead in low-gradient regions.
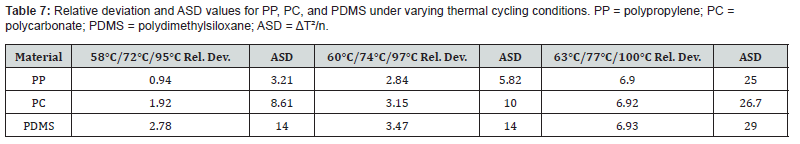
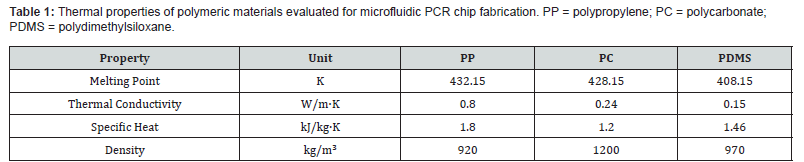
The chip architecture was derived from the amplification protocol of a validated genus-specific real-time PCR method targeting the 18S rRNA gene. The procedure requires a total reaction volume of 30 μL, comprising 25 μL of PCR master mix and 5 μL of extracted DNA template. The fluid subsequently traverses thermally defined zones corresponding to the three PCR stages: annealing, extension, and denaturation. Relevant supplementary data on the PCR protocol and viscosity estimation are available in the associated supplementary files [26]. Three candidate polymers were screened for suitability as chip fabrication materials: polypropylene (PP), polycarbonate (PC), and polydimethylsiloxane (PDMS). The thermal characteristics of each material are summarized in Table 1.
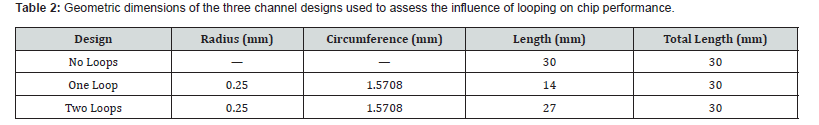
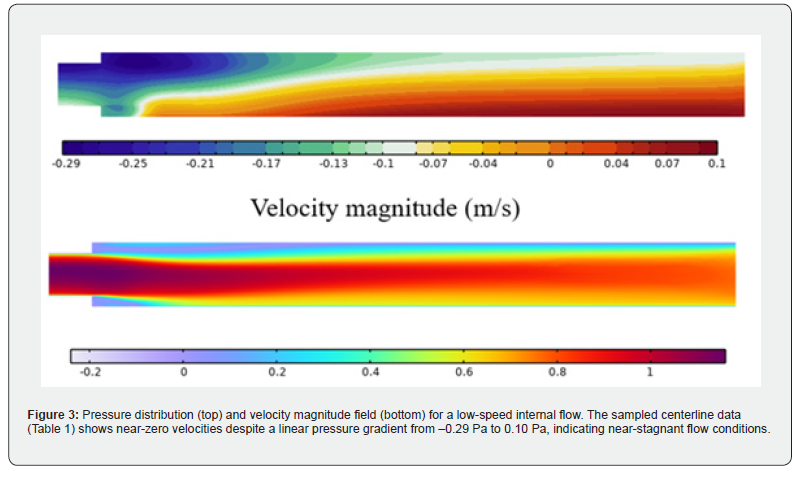
To determine the impact of channel looping on fluid temperature uniformity and device efficiency, three geometric variants were modeled and simulated: a straight-channel design (no loops), a single-loop configuration, and a dual-loop configuration. All designs were constrained to equivalent total path lengths to isolate the effect of looping geometry. All three configurations were assigned a copper plate temperature of 58°C during simulation. The geometric specifications are listed in Table 2, and the resulting flow field is illustrated in Figure 3. The pressure field varies linearly from approximately −0.29 Pa to 0.10 Pa across the channel domain, consistent with fully developed viscous flow under a weak imposed pressure gradient. The centerline velocity magnitude remains near zero, which is attributable to the extremely low Reynolds number (Re ≈ 4.5 × 10⁻´) characterizing flow in this microscale geometry. This nearstagnant condition reflects the dominance of viscous forces and confirms that the flow is entirely in the Stokes regime, consistent with the analytical framework presented in Section 2.
The overall microfluidic chip was modeled with dimensions of 73 mm × 45.5 mm. The denaturation and annealing zones were each assigned a length of 15 mm, while the extension zone was set to 10 mm, with 2 mm spacing between zones. Due to software licensing constraints on the academic version of ANSYS, the total mesh element count was limited to 512,000 elements. Mesh quality was maintained at a minimum threshold of 0.10 to ensure simulation reliability. The viscosity of the PCR reaction mixture was estimated using the Falkenhagen equation, following the approach of Yang et al. [27]. Additional operating parameters for each thermal zone are presented in Table 3.
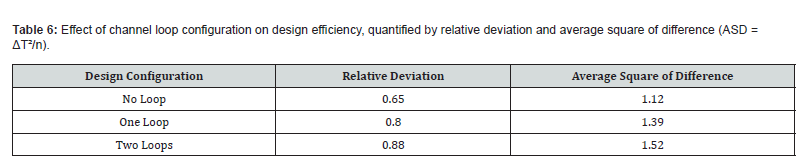
Following the loop geometry assessment, each of the three polymeric materials was incorporated into the 2-cycle chip simulation and evaluated across three sets of PCR temperatures: 58/72/95°C, 60/74/97°C, and 63/77/100°C (corresponding to annealing, extension, and denaturation temperatures, respectively). Performance was quantified using relative deviation and the average square of difference (ASD) relative to target temperatures. Once the most efficient material was identified, a secondary comparison was performed to evaluate the effect of varying channel cross-sectional area. Two designs with differing diameters and channel lengths were simulated at identical inlet velocities. Design 1 had a diameter of 300 μm, a channel length of 200 μm, and a cross-sectional area of 130,686 μm². Design 2 used a diameter of 500 μm, a length of 300 μm, and a cross-sectional area of 346,350 μm².
Liquid Cooling Microchannel Optimization
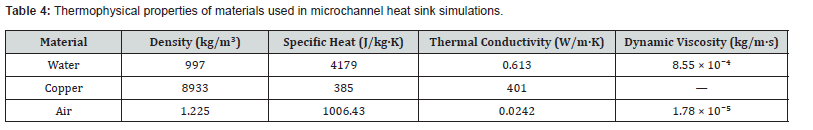
The heat sink substrate measured 8 mm (width) × 8.26 mm (length) × 1 mm (height) and was modeled as copper, while water served as the working fluid. Thermophysical properties used in all simulations are summarized in Table 4. Fifteen cross-sectional geometries were evaluated, encompassing the most commonly encountered forms in microchannel literature [28,29]. For all shapes, the hydraulic diameter was held constant at 533 μm and the wall width at 800 μm, following the normalization approach of Deng et al. [30]. A uniform heat flux of 103,057 W/m² was imposed on the top substrate surface; the bottom surface and sidewalls were thermally insulated.
Fluid flow and heat transfer were modeled under the following simplifying assumptions: steady-state, single-phase, laminar, incompressible flow; negligible viscous dissipation; negligible gravitational effects; and no natural convection or radiation. Under these conditions, the governing conservation equations for mass, momentum, and energy-separately for the fluid and solid domains-were solved in their standard incompressible forms. For the buoyancy-driven convection PCR model (Section 3c), temperature-dependent fluid properties for water are employed throughout. The Nonisothermal Flow interface in COMSOL Multiphysics models the fully coupled heat transfer and fluid flow. The incompressible Navier-Stokes equation with a buoyancy source term governs the momentum balance and accounts for the density-driven lifting force responsible for natural convection:
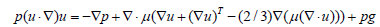
In Eq. 4, the dependent variables for flow are the fluid velocity vector u and the pressure p. The constant g denotes gravitational acceleration, ρ gives the temperature-dependent density, and μ is the temperature-dependent dynamic viscosity. The local bulk fluid temperature was determined by a cross-sectional massflux- weighted integral, and the average wall temperature was taken as the spanwise mean across the channel width. The local heat transfer coefficient at each axial position was calculated from the imposed heat flux divided by the product of the local temperature difference (wall minus bulk fluid) and the ratio of base area to wetted area. Local and average Nusselt numbers were then derived using the hydraulic diameter and fluid thermal conductivity as characteristic scales. Pressure drop was evaluated as the difference between inlet and outlet static pressures. No-slip boundary conditions apply on all boundaries. At steady state, the heat transfer within the fluid is given by:
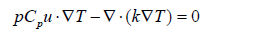
where T is the temperature, k denotes the thermal conductivity, and Cₚ is the specific heat capacity of the fluid. The mass transfer in the channel is governed by convection and diffusion using the Transport of Dilute Species interface:
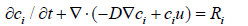
where Dᵢ denotes the diffusion coefficient (10⁻⁷ cm²/s) and cᵢ is the species concentration (mol/m³). The term Rᵢ (mol/(m³·s)) corresponds to the species› net reaction rate.
A uniform inlet velocity and temperature were prescribed at the channel entrance, while an atmospheric pressure condition was applied at the outlet. No-slip and temperature-continuity conditions were enforced at all solid-fluid interfaces, and adiabatic conditions were applied to the bottom surface and lateral walls.
A simplified first-order reaction mechanism is used to
represent the PCR process:
dsDNA → 2 ssDNA (Denaturation, 95°C)
ssDNA → aDNA (Annealing, 55°C)
aDNA → dsDNA (Extension, 72°C)
The rates of change in concentration of DNA components are determined from stoichiometric balances. The denaturation, annealing, and extension reaction rates are given by kd, ka, and ke, respectively. These reaction rate constants are spatially modulated using a Gaussian mapping function to localize each reaction in its respective temperature zone [32]. The characteristic reaction rate k* (= kᵢ/κᵢ) was taken to be 5.564 s⁻¹. The dimensionless reaction rate constants κᵢ were taken to be 1, 1, and 0.5 for denaturation, extension, and annealing respectively. The mapping function is defined as:
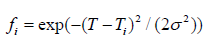
where σ (= 1°C) is the standard deviation of the reaction rate with respect to temperature and Tᵢ is the ideal temperature for each reaction. The initial value of dsDNA template is set to 1 mol/ m³ with no-flux boundary conditions.
Mesh independence was established by testing four tetrahedral mesh densities (206,000; 485,000; 663,000; and 800,000 elements) at Re = 500. The local Nusselt number profile converged with increasing element count, and the deviation between the 663,000- and 800,000-element meshes was below 0.63%. The 663,000-element mesh was therefore adopted for all subsequent simulations, balancing accuracy and computational cost. The numerical model was validated against published data from Alfaryjat et al. [31], who experimentally and numerically characterized circular, hexagonal, and rhombus cross-sections. Agreement was satisfactory across the Reynolds number range, with maximum deviations of approximately 4% in the average heat transfer coefficient and 10% in the pressure drop at Re = 1000-both within the tolerance reported in comparable studies. To compare cross-sections on equal footing, all heat transfer and pressure-drop results were normalized relative to the horizontal rectangular cross-section, which represents the standard baseline geometry in commercial microchannel heat sinks.
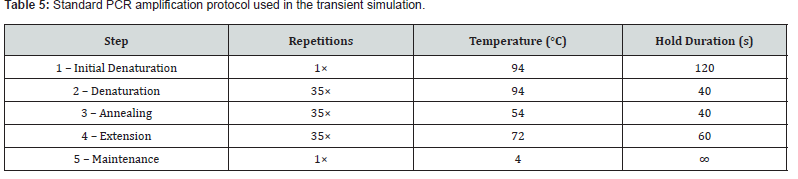
The heat transfer enhancement factor (E𝒩𝙊) and the pressure-drop penalty factor (E𝒳) were defined as the ratios of each geometry’s Nusselt number and pressure drop, respectively, to those of the reference rectangle. The overall performance evaluation criteria (PEC) index η was computed as E𝒩𝙊 / E𝒳¹⁄³. A value of η > 1 indicates that the net thermohydraulic benefit of a given cross-section exceeds that of the rectangular reference, accounting for both heat transfer gain and pressure-drop penalty. Having identified the optimal cross-sectional geometry, a transient simulation was conducted on a representative single-well PCR assembly. The assembly dimensions were 32.25 mm × 8 mm × 8.26 mm (height, width, length), with symmetry boundary conditions applied to all four lateral faces. A time-step independence study using four step sizes confirmed that Δt = 0.1 s produced converged temperature histories. The standard PCR thermocycle protocol (Table 5) was prescribed as a time-varying heat flux on the heater surface.
Results and Discussion

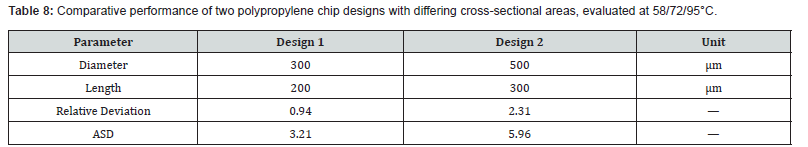
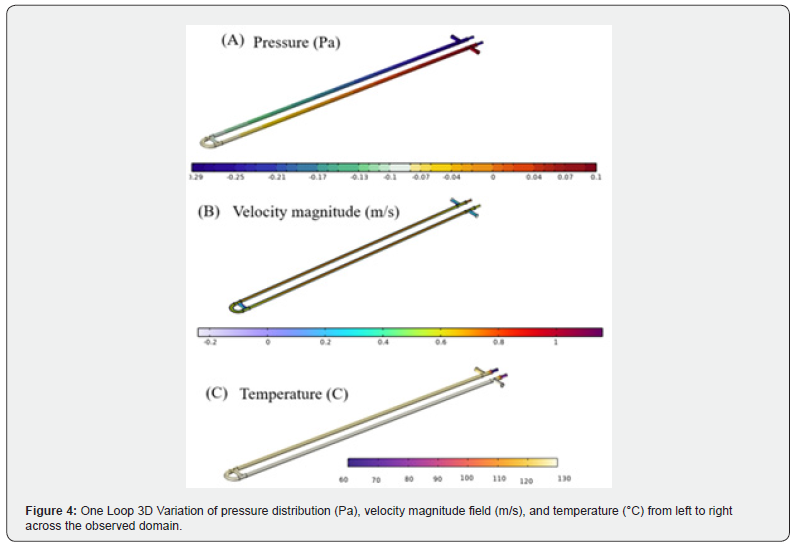
The results of the three computational investigations are presented in sequence, corresponding to the chip design, cooling optimization, and buoyancy-driven amplification studies described in Section 2 and Section 3. ANSYS CFD was employed to analyze fluid behavior, heat transfer, and reactive transport within the microscale environments characteristic of lab-on-chip devices, leveraging its capabilities for multiphase flow modeling, species transport, and surface-reaction kinetics [33,34]. A key advantage of the simulation-first approach is the early identification of performance limitations-including bubble entrapment, flow nonuniformity, localized thermal gradients, and dead-zone formationprior to physical fabrication, thereby reducing development costs and experimental iteration cycles. Three designs-one with no loops, one with a single loop, and one with two loopswere simulated at a copper plate temperature of 58°C. The resulting relative deviations and average squares of difference are presented in Table 6. (Figure 4) presents the three-dimensional spatial distributions of pressure (Pa), velocity magnitude (m/s), and temperature (°C) for the single-loop configuration, displayed side by side for direct comparison. The coupled nature of the flow and thermal fields is evident: elevated fluid velocities in the loop interior correspond to localized pressure gradients, while thermal contours confirm that the fluid approaches the prescribed copper plate temperature within each zone. Together, these field distributions validate the numerical fidelity of the CFD model and illustrate the interplay between hydrodynamics and heat transfer in the looped channel geometry.
The data indicate a mild but consistent increase in relative deviation as loop count increases-from 0.65 with no loops to 0.88 with two loops. This trend reflects a minor reduction in temperature uniformity, attributable to the altered flow dynamics and additional contact area introduced by the loop geometry. However, the magnitude of this degradation is minimal, and the looped configurations remain thermally stable relative to the target temperature. From a practical standpoint, incorporating loops into the chip design offers meaningful advantages. Looping reduces the total linear length of the chip while preserving the required total fluid path length, thereby enabling a more compact chip footprint. Additionally, channel loops promote vortex formation and secondary flow patterns that enhance reagent mixing, potentially improving reaction homogeneity. The introduction of redundancy through multiple loop circuits may further contribute to reaction stability by enabling the system to compensate for localized flow perturbations.
The proposed microfluidic PCR chip was modeled in ANSYS as a 44-cycle configuration encompassing full thermal cycling across all three PCR stages. The design incorporates five thermally active copper plates maintained at specific temperatures corresponding to the annealing, extension, and denaturation conditions required for DNA amplification. Temperature conditions for the three simulation runs were set as follows: 58/72/95°C (first simulation), 60/74/97°C (second), and 63/77/100°C (third), applied in sequence across the five plate positions. In microfluidic PCR applications, the number of amplification cycles directly governs the sensitivity and detection limits of the assay. An elevated cycle count supports exponential accumulation of target sequences, which is particularly advantageous when starting from low-copy-number samples. The design specification of 44 cycles exceeding the 36-cycle threshold established for conventional PCR at this sensitivity level-was adopted to account for the reduced sample volume processed in the microfluidic format [35]. Due to software licensing restrictions on the academic version of ANSYS, the full 44-cycle chip could not be meshed within the available computational budget. Accordingly, a 2-cycle chip was used as a simulation surrogate, with the results taken as representative of the full-scale design behavior. The performance of each polymeric material under three sets of temperature conditions is summarized in Table 7.
Among the three materials tested, polypropylene at the 58/72/95°C temperature setting produced the lowest relative deviation (0.94) and ASD (3.21), indicating the best combination of precision and accuracy for temperature delivery within the chip. These findings align with the known physicochemical properties of the materials. Polypropylene’s relatively high thermal conductivity (0.80 W/m·K) compared to PC (0.24 W/m·K) and PDMS (0.15 W/m·K) enables more responsive heat transfer between the copper plates and the fluid channel [36,37]. Its chemical inertness, biocompatibility, and favorable manufacturability further support its suitability for PCR chip fabrication. Polycarbonate offers advantages in terms of optical transparency and chemical resistance, which could be valuable for real-time fluorescence-based detection [38]. PDMS, while highly flexible and amenable to soft lithographic fabrication of intricate channel geometries, exhibited the greatest thermal deviation in this study, likely due to its very low thermal conductivity limiting effective heat transfer across the channel wall [39,40]. Following identification of polypropylene as the optimal material, the influence of channel geometry was examined by comparing two design variants differing in channel diameter and length while maintaining the same inlet flow velocity. Results are shown in Table 8.
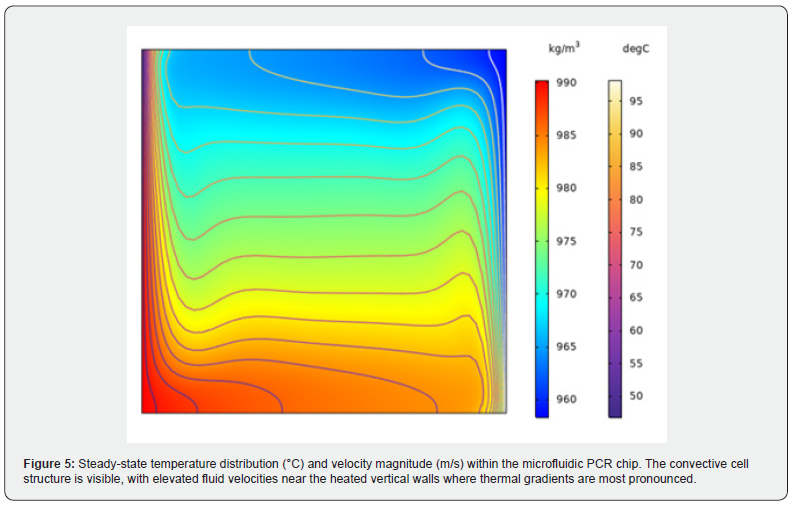
Increasing the channel diameter from 300 μm to 500 μm and the length from 200 μm to 300 μm resulted in a marked increase in relative deviation (0.94 to 2.31) and ASD (3.21 to 5.96). This deterioration in thermal performance is attributable to the increased volume of fluid that must be heated or cooled within each zone, which reduces the efficiency of temperature equilibration when the volumetric flow rate is held constant. The smaller cross-sectional design therefore provides more favorable heat-transfer kinetics and is recommended for use in the physical fabrication of the chip. (Figure 5) illustrates the steady-state velocity profile alongside temperature contours within the microchannel. A prominent convective cell dominates the flow, with fluid moving more rapidly along the vertical walls where the temperature gradient is most pronounced.
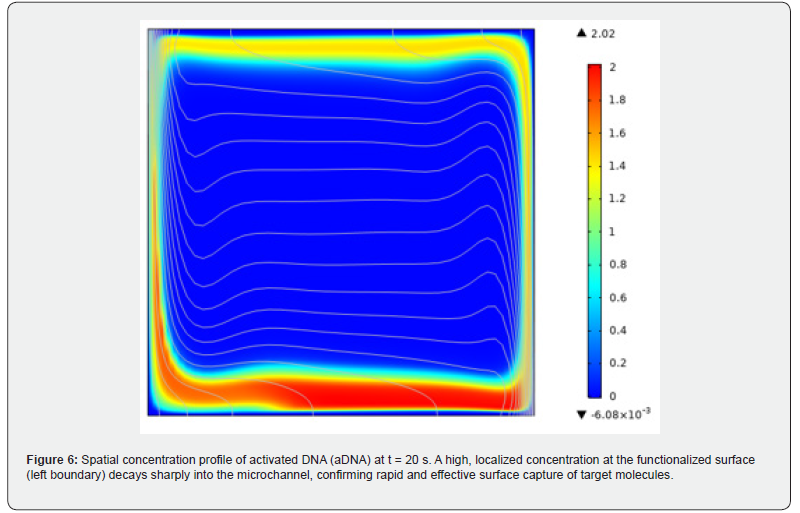
Two factors primarily govern heat transfer magnitude in microchannels: the ratio of substrate base area to total wetted surface area (Aˇ/NAâ), and the uniformity of temperature distribution within the channel. The circular and trapezoidal cross-sections exhibit the highest Aˇ/NAâ values among all shapes tested, which promotes convective heat transfer. However, the trapezoidal geometry produces pronounced thermal hotspots at its acute corners, leading to uneven temperature distributions that partially offset the benefit of its large, wetted area. The circular cross-section, by contrast, offers both a favorable wetted area ratio and the smallest fluid-to-wall temperature differential, yielding the highest Nusselt numbers across the entire Reynolds number range examined (350–1150). At Re = 1150, the circular crosssection surpasses the square by approximately 25% in average Nusselt number [41]. (Figure 6) presents the spatial concentration profile of activated DNA (aDNA) 20 seconds into the simulated assay. The aDNA, representing the target molecule bound to the surface activator, shows a high, localized concentration at the functionalized surface (left boundary), forming a steep gradient that decays sharply into the microchannel. This profile confirms the successful and rapid capture of target molecules at the sensor interface.
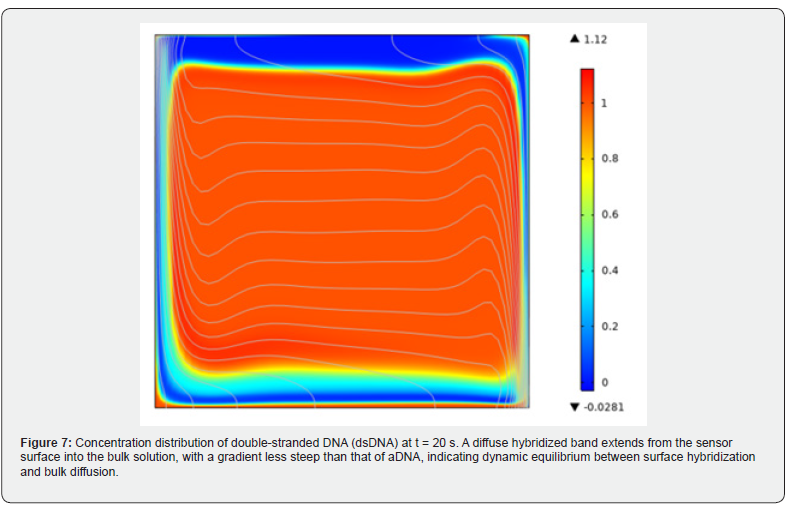
Cross-sections with sharp interior angles-including delta, gradient, and various trapezoidal configurations-consistently exhibited lower heat transfer performance, attributable to recirculation zones and stagnant dead zones forming at the corners. These regions impede convective transport and promote local thermal energy accumulation, reducing overall thermal performance. (Figure 7) presents the concentration distribution of double-stranded DNA (dsDNA) at the 20-second mark. The profile reveals a diffuse band extending from the sensor surface into the bulk solution, indicating the presence of hybridized surface complexes. The gradient is less steep than that of aDNA, consistent with a dynamic equilibrium between hybridization at the surface and diffusion or dissociation into the bulk channel volume. Pressure drop across all cross-sections increases monotonically with Reynolds number, as expected for laminar flow. Among all geometries, the trapezoidal cross-sections generate the greatest pressure drop, while the circular cross-section produces the lowest values at every Reynolds number tested. The PEC index η normalizes results to the rectangular cross-section, and the circular geometry consistently achieves the highest η values across the entire Reynolds number range-confirming that its heat transfer gains more than compensate for any pressure-drop increase. Conversely, the trapezoidal cross-sections yield the lowest η, reflecting the disproportionate pressure-drop penalty associated with their corner geometry [42].


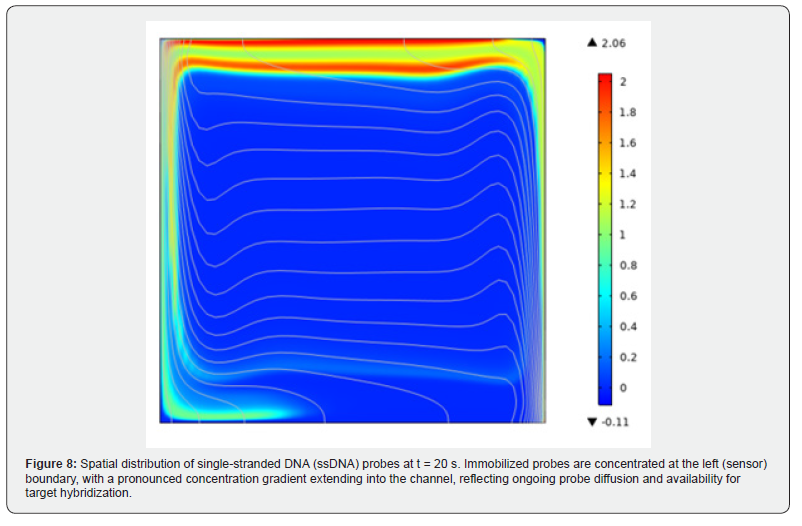
When the circular microchannel heat sink was positioned beneath a single-well PCR device and the standard thermocycle protocol was applied, transient simulation results indicated that the sample temperature reaches the target annealing setpoint in 16.5 seconds per cooling step. By comparison, a benchmark airfan cooling system requires approximately 20.97 seconds per equivalent step-a difference of approximately 4.5 seconds per cycle. Multiplied over 35 amplification cycles, this differential accumulates to a total time saving of approximately 157.5 seconds (2.6 minutes) per amplification run [43]. (Figure 8) presents the spatial distribution of single-stranded DNA (ssDNA) probes after 20 seconds. A high concentration is maintained at the left boundary (sensor surface), representing the immobilized probe layer. A pronounced concentration gradient extending into the channel reflects the ongoing diffusion of unbound ssDNA and confirms probe availability for continued target hybridization.
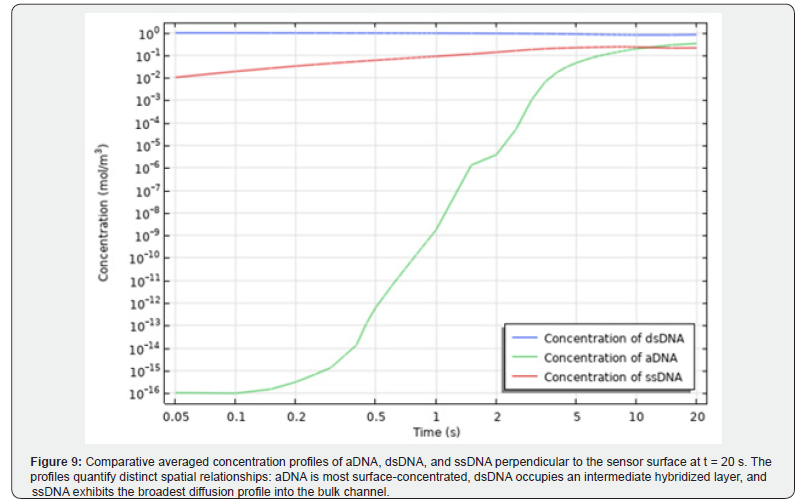
This time reduction carries direct practical implications: in high-throughput diagnostic laboratories processing large numbers of samples, shortening each amplification run by several minutes translates into substantially increased daily throughput. Furthermore, the microchannel heat sink integrates compactly with the standard 96-well PCR plate format, eliminating the bulky external enclosures and fans associated with conventional aircooled systems. (Figure 9) presents a comparative summary of the averaged concentration gradients of aDNA, dsDNA, and ssDNA measured perpendicular to the sensor surface at 20 seconds. The composite profile highlights distinct spatial relationships: aDNA is highly surface-concentrated, dsDNA forms an intermediate hybridized layer, and ssDNA exhibits the broadest diffusion profile extending furthest into the bulk channel. Together, these profiles provide quantitative confirmation of the binding and hybridization kinetics established during the early stage of the assay.
The steady-state velocity profile analysis confirms that a large convective cell occupies the channel. The fluid flows along the boundaries and is faster at the vertical walls where the temperature variations are highest, consistent with Rayleigh- Bénard type convection in the vertical geometry [32]. Reaction rate analysis after 120 seconds demonstrates that denaturation, annealing, and extension reactions are localized around 95°C, 55°C, and 72°C respectively, validating the Gaussian mapping function approach for temperature-dependent reaction localization. Initially, only denaturation is taking place in the channel, but after a transition period, annealing and extension effects become visible as the convective circulation establishes steady flow patterns. Concentration profile analysis of DNA components over a 120-second simulation confirms the characteristic PCR kinetics: the concentration of double-stranded DNA template (dsDNA) initially decreases due to denaturation but later exponentially increases due to amplification in the extension region. The doubling time of approximately 60 seconds is of the same order of magnitude as reported in the literature [32] and compares favorably with the doubling time of a conventional laboratoryscale PCR system of approximately 2 minutes.
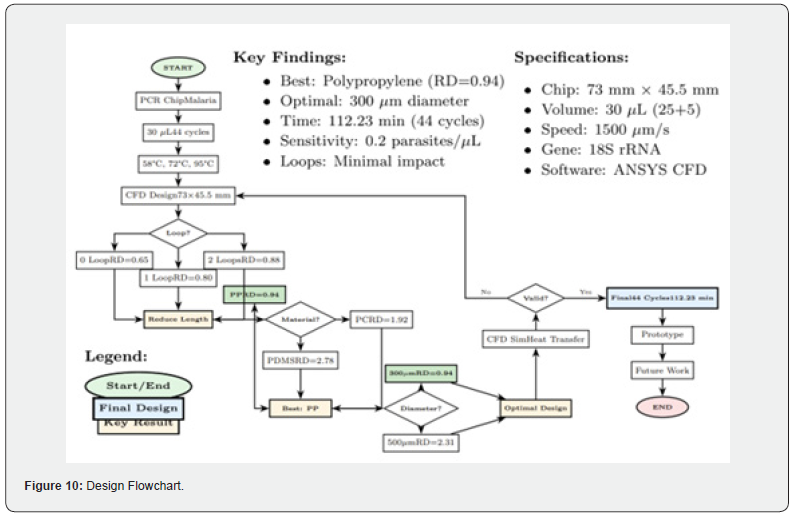
Based on the zone-specific residence times defined in Table, the proposed 44-cycle microfluidic PCR chip requires approximately 112.23 minutes to complete a full amplification run. While this is longer than some rapid molecular diagnostic platforms, it remains within an acceptable range for point-of-care settings where the alternative involves sending samples to a centralized laboratory. Further design optimization-particularly in reducing transition times between thermal zones, combined with the liquid cooling enhancements identified in Part II-may enable significant reductions in total cycle time in future iterations. (Figure 10) presents a design flowchart summarizing the systematic optimization process for the microfluidic PCR chip. Beginning from defined target specifications (30 μL reaction volume, 44 amplification cycles, 1500 μm/s flow velocity), the framework proceeds through three sequential optimization stages: (i) loop configuration assessment, comparing zero-, one-, and two-loop geometries; (ii) material selection, evaluating PP, PC, and PDMS on the basis of thermal performance metrics; and (iii) cross-sectional geometry optimization, selecting a 300 μm diameter over a 500 μm alternative. The validated design parameters are subsequently carried forward into CFD heat transfer simulation and provide a basis for experimental prototype fabrication, culminating in a chip capable of completing a full 44-cycle amplification run in 112.23 minutes.
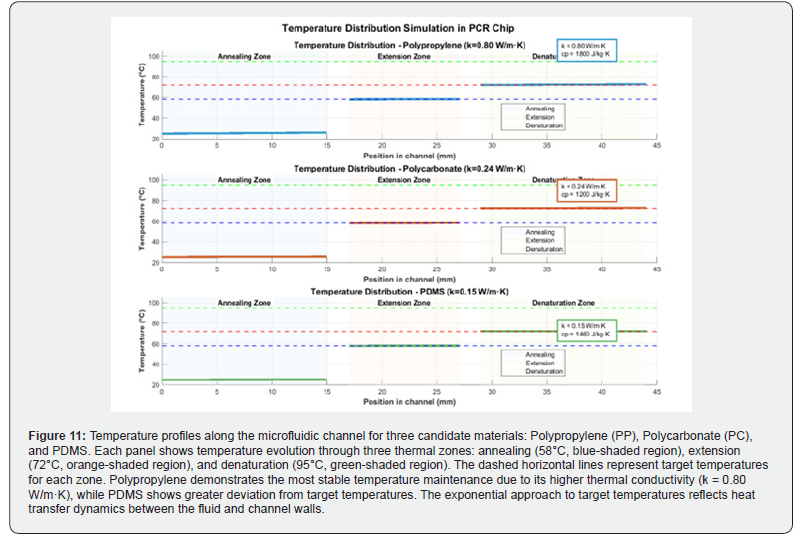
Figure 11 presents the simulated axial temperature profiles along the microfluidic PCR channel for three candidate polymeric materials: polypropylene (PP), polycarbonate (PC), and PDMS. Each panel corresponds to one material and displays the thermal evolution of the fluid as it traverses the three sequential heating zones: annealing (58°C), extension (72°C), and denaturation (95°C), indicated by blue, orange, and green shaded regions, respectively, with dashed horizontal lines denoting target temperatures. The fluid temperature follows an exponential approach toward each setpoint, governed by the convective heat transfer between the fluid and the heated channel walls. Polypropylene (top panel) achieves the closest agreement with target temperatures across all three zones, attributable to its superior thermal conductivity (k = 0.80 W/m·K), which facilitates efficient heat exchange through the channel wall. Polycarbonate (middle panel) shows intermediate performance, while PDMS (bottom panel) exhibits the greatest lag and deviation from target temperatures, particularly in the annealing and extension zones, consistent with its low thermal conductivity (k = 0.15 W/m·K). These profiles directly support the material selection decision and underscore the critical role of substrate thermal conductivity in achieving the precise temperature control required for reliable PCR amplification.
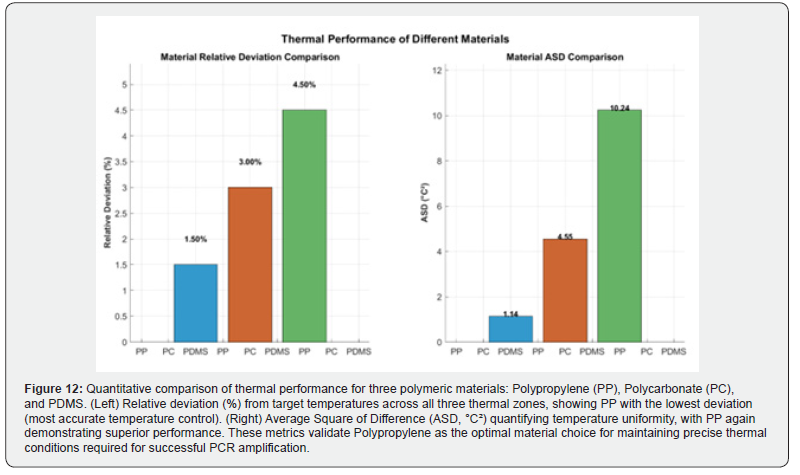
Figure 12 presents a quantitative comparison of thermal performance for the three candidate materials using two complementary metrics: relative deviation (%) and the Average Square of Difference (ASD, °C²). The left panel displays bar charts of the relative deviation from target temperatures across all three thermal zones, with polypropylene (PP) yielding the lowest value (approximately 3.33%), followed by polycarbonate (PC) and PDMS. The right panel presents the corresponding ASD values, confirming the same ranking. These results collectively establish that PP provides the most accurate and thermally uniform performance among the materials evaluated, directly validating its selection as the preferred fabrication material for the microfluidic PCR chip.
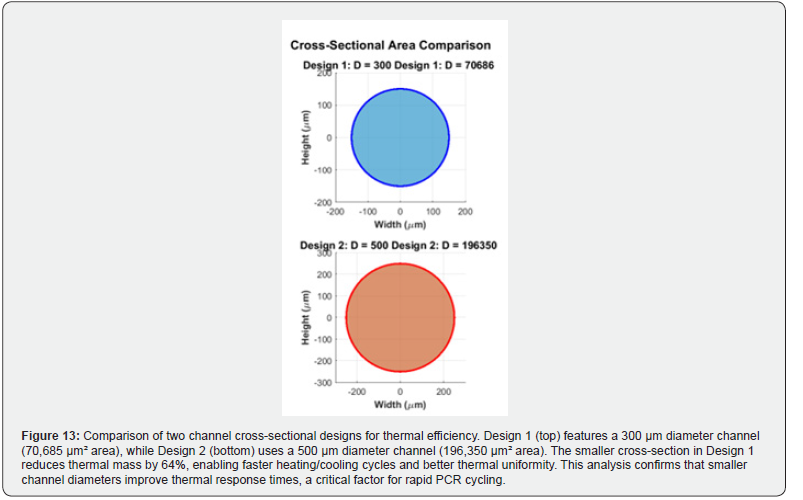
Figure 13 compares two candidate channel cross-sectional designs in terms of their thermal efficiency. Design 1 employs a circular cross-section with a 300 μm diameter (cross-sectional area: 70,685 μm²), while Design 2 uses a 500 μm diameter (crosssectional area: 196,350 μm²). The smaller cross-section of Design 1 reduces the thermal mass by approximately 64% relative to Design 2, enabling substantially faster heating and cooling transients. This improvement in thermal responsiveness is critical for rapid PCR thermocycling, where minimizing the transition time between temperature setpoints directly reduces total assay duration.
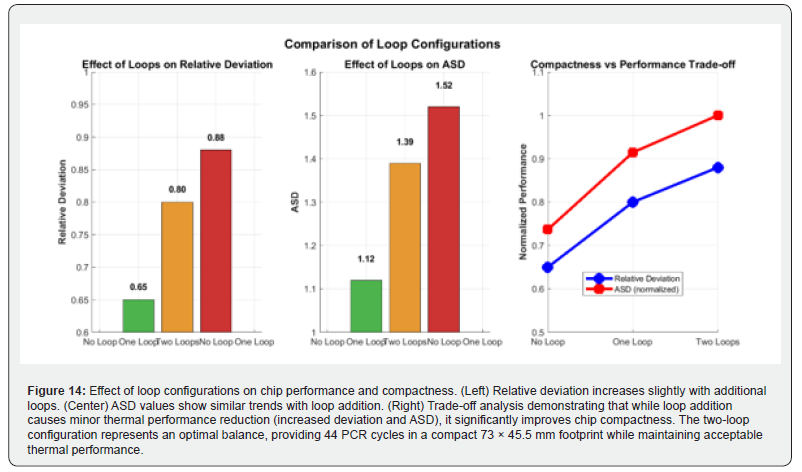
Figure 14 illustrates the effect of loop configuration on chip thermal performance and physical compactness through a three-panel comparative analysis. The left and center panels show the relative deviation and ASD values, respectively, for the no-loop, single-loop, and dual-loop configurations; both metrics increase marginally with loop count, confirming the mild thermal penalty associated with increased geometric complexity. The right panel presents a trade-off analysis in which normalized performance metrics are plotted against compactness: while loop addition causes a modest reduction in thermal uniformity, it yields a substantial gain in device compactness. The two-loop configuration represents the optimal balance, accommodating a full 44-cycle amplification protocol within a 73 mm × 45.5 mm footprint while maintaining thermal performance within acceptable bounds for PCR applications.
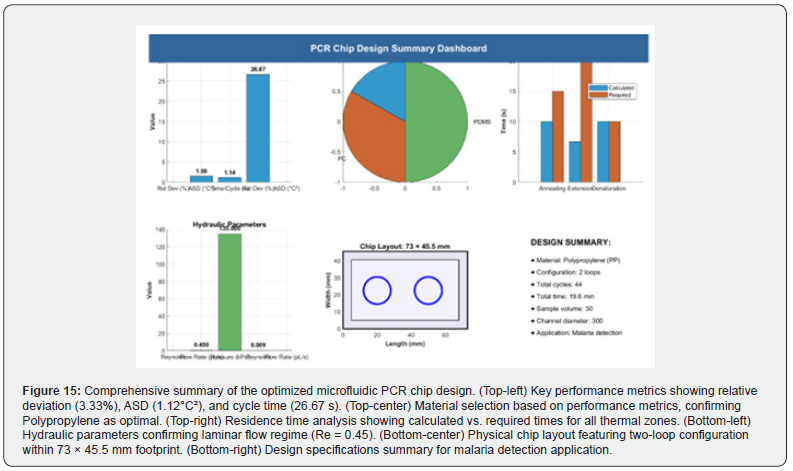
Figure 15 presents a comprehensive summary dashboard that
integrates all key aspects of the final PCR chip design into a single
visualization comprising six panels:
i. Key Performance Metrics: Bar chart showing relative
deviation (3.33%), ASD (1.12 °C²), and time per cycle (26.67 s).
ii. Material Selection: Pie chart confirming Polypropylene
as the chosen material based on its lowest relative deviation.
iii. Residence Time Analysis: Grouped bar chart comparing
calculated versus required residence times for each thermal zone,
all meeting or exceeding requirements.
iv. Hydraulic Parameters: Bar chart displaying Reynolds
number (0.45, confirming laminar flow), volumetric flow rate, and
pressure drop.
v. Chip Layout: Schematic diagram of the physical chip
layout (73 mm × 45.5 mm) with two loops for compactness.
Design Specifications: Text summary listing critical parameters including 44 cycles, total run time, sample volume, channel diameter, and the target application (malaria detection). Together, these panels provide a holistic overview of the optimized microfluidic PCR chip, validating the design decisions and summarizing the operational specifications for practical implementation in diagnosing Plasmodium falciparum at low parasitemia levels.
Conclusion
This integrated computational study establishes a cohesive
framework for the rational design of microfluidic PCR platforms
by simultaneously addressing three critical aspects: chip
architecture, thermal management, and reaction kinetics. The
principal findings and their implications are summarized as
follows:
a) The introduction of channel loops provides a viable
strategy for achieving a compact device footprint, with
only a marginal increase in thermal deviation (from 0.65 to
0.88). Among the candidate polymers, polypropylene (PP)
demonstrated superior performance, yielding the lowest relative
deviation (0.94) and Average Square of Difference (3.21) under
standard PCR conditions (58/72/95°C). This is attributed to its
favorable thermal conductivity (0.80 W/m·K), which ensures
precise temperature control. Furthermore, a smaller channel
cross-section (300 μm diameter) was shown to enhance thermal
responsiveness and fidelity compared to larger designs, providing
a clear guideline for geometric optimization.
b) A systematic comparison of fifteen microchannel crosssections
identified the circular geometry as optimal for PCR
cooling applications. It delivered a 25% higher average Nusselt
number than a square channel at Re = 1150 while simultaneously
generating the lowest pressure drop. The Performance Evaluation
Criteria (PEC) index confirmed its superior thermohydraulic
performance. Transient simulations of a full 35-cycle protocol
revealed that this optimized liquid cooling system reduces the
cooling time per cycle to 16.5 seconds, culminating in a total time
saving of approximately 157.5 seconds (2.6 minutes) per run
compared to conventional air-based cooling-a significant gain for
high-throughput diagnostics.
c) The convection-reaction-diffusion model successfully
validated a pump-free PCR paradigm, where temperatureinduced
density gradients autonomously drive fluid circulation.
The simulation confirmed exponential DNA amplification with
a doubling time of approximately 60 seconds, consistent with
established experimental benchmarks. This demonstrates the
feasibility of simplifying device architecture by eliminating
external pumps, which is advantageous for developing portable,
field-deployable diagnostic systems.
Collectively, these findings are not isolated results but form a
synergistic design toolkit. The recommended polypropylene chip
with a compact looped layout can be effectively paired with the
circular microchannel heat sink for active thermal management,
while the buoyancy-driven model offers a pathway to passive,
pump-free operation. This multi-faceted approach enables
engineers to make informed trade-offs between speed, footprint,
complexity, and performance. To transition from computational
insight to practical implementation, future work should focus on:
i. Full-scale simulation of the 44-cycle chip geometry to
validate extrapolated performance metrics
ii. Experimental prototyping and validation of the
integrated design combining the optimized chip, cooling system,
and buoyancy-driven flow
iii. Advanced material exploration, including biocomposites
and substrates with enhanced thermal properties
iv. Further heat transfer intensification through wavy or
oblique microchannel patterns and the use of nanofluids
v. System-level integration with on-chip sample
preparation and detection modules to create a fully functional
lab-on-a-chip device.
In conclusion, this work provides a comprehensive computational roadmap for developing next-generation microfluidic PCR platforms that are faster, more compact, and better suited for point-of-care and resource-limited diagnostic settings than conventional systems. The integrated framework presented here bridges the gap between isolated component studies and holistic platform design, offering a systematic, simulation-driven basis for advancing miniaturized PCR technology toward practical field deployment.
Author Contribution
Conceptualization, MYAJ; Methodology, MYAJ; Software, MYAJ; Validation, MYAJ; Formal analysis, MYAJ; Investigation, MYAJ; Resources, MYAJ; Data curation, MYAJ; Writing-original draft preparation, MYAJ; Writing-review and editing, MYAJ; Visualization, MYAJ; Supervision, MYAJ; Project administration, MYAJ. All authors have reviewed and approved the final version of the manuscript.
- Rahman MT, Uddin MS, Sultana R (2013) Polymerase chain reaction (PCR): a short review. Anwer Khan Modern Medical College Journal 4: 30-36.
- Mullis KB (1994) The polymerase chain reaction (Nobel Lecture). Angewandte Chemie International Edition 33(12): 1209-1213.
- Mullis KB (1990) The unusual origin of the polymerase chain reaction. Scientific American 262(4): 56-65.
- Cha RS, Thilly WG (1993) Specificity, efficiency, and fidelity of PCR. PCR Methods and Applications 3(3): S18-S29.
- Dong L (2023) A rapid multiplex assay of human pathogenic parasites by digital PCR. Clin Chim Acta 539: 70-78.
- Taghdiri A (2019) Detection of pathogens using nested PCR and investigation of genetic diversity by 18S rRNA gene. Microb Pathog 137: 103782.
- Khater A, Abdelrehim O, Mohammadi M (2021) Thermal droplet microfluidics: from biology to cooling technology. TrAC Trends in Analytical Chemistry 138: 116234.
- Kleiner MB, Kuehn SA, Haberger K (1995) High performance forced air cooling scheme employing microchannel heat exchangers. IEEE Transactions on Components, Packaging, and Manufacturing Technology Part A 18: 795-804.
- Dong X (2021) Rapid PCR powered by microfluidics: a quick review under the COVID-19 pandemic. TrAC Trends Anal Chem 143: 116377.
- Struijk R (2023) Ultrafast RNA extraction-free SARS-CoV-2 detection by direct RT-PCR. Diagn Microbiol Infect Dis 107(1): 115975.
- Zhao X, Li M, Liu Y (2019) Microfluidic-based approaches for foodborne pathogen detection. Microorganisms 7(10): 381.
- Wang X (2022) Microfluidics-based strategies for molecular diagnostics of infectious diseases. Mil Med Res 9(1): 11.
- Bakuova N (2023) Design, simulation, and evaluation of polymer-based microfluidic devices via CFD and cell culture on-chip. Biosensors 13(7): 754.
- Kim E (2023) Rapid and accurate on-site identification of microorganisms using microfluidic chip PCR. LWT 179: 114635.
- Khan WA, Yovanovich MM, Culham JR (2006) Optimization of microchannel heat sinks using entropy generation minimization. Annual IEEE Semiconductor Thermal Measurement and Management Symposium pp: 78-86.
- Dix J, Jokar AA (2008) Microchannel heat exchanger for electronics cooling applications. ASME International Conference on Nanochannels, Microchannels, and Minichannels pp : 1935-1936.
- Zhao N, Guo L, Qi C (2019) Experimental study on thermo-hydraulic performance of nanofluids in CPU heat sink with rectangular grooves. Energy Conversion and Management 181: 235-246.
- Qi C, Tang J, Fan F, Yan Y (2020) Effects of magnetic field on thermo-hydraulic behaviors of magnetic nanofluids in CPU cooling systems. Applied Thermal Engineering 179: 115717.
- Mulberry G, White KA, Vaidya M (2017) 3D printing and milling a real-time PCR device for infectious disease diagnostics. PLOS ONE 12(6): 1-18.
- Alihosseini Y, Bari AR, Mohammadi M (2021) Effective parameters on increasing efficiency of microscale heat sinks and application of liquid cooling in real life. Advances in Microfluidics and Nanofluidics.
- Cao Y (2022) Simultaneous detection of multiple foodborne bacteria by LAMP on a microfluidic chip. Food Control 134: 108694.
- Zhang D (2023) All-in-one microfluidic chip for 30-min quantitative POCT of nucleic acids. Sens Actuators B Chem 390: 133939.
- Hewlin RL, Edwards M, Schultz C (2023) Design of a travelling wave ferro-microfluidic device for magnetophoretic cell separation. Micromachines 14: 889.
- Hewlin R, Edwards M, Smith M (2023) A 2D transient computational model for analyzing magnetic and non-magnetic particle dynamics in a ferro-microfluidic device. J Eng Sci Med Diagn Ther 14: 1-47.
- Bi W (2023) Implementation of blood-brain barrier on microfluidic chip. Ageing Res Rev 87: 101921.
- Hermsen CC (2001) Detection of pathogen DNA in vivo by real-time quantitative PCR. Mol Biochem Parasitol 118: 247-251.
- Yang XD, Melnik RVN (2007) Effect of internal viscosity on Brownian dynamics of DNA molecules in shear flow. Comput Biol Chem 31(2): 110-114.
- Wang H, Chen Z, Gao J (2016) Influence of geometric parameters on flow and heat transfer performance of micro-channel heat sinks. Applied Thermal Engineering 107: 870-879.
- Gunnasegaran P, Mohammed HA, Shuaib NH, Saidur R (2010) The effect of geometrical parameters on heat transfer characteristics of microchannels heat sink with different shapes. International Communications in Heat and Mass Transfer 37(8): 1078-1086.
- Deng D, Pi G, Zhang W (2019) Numerical study of double-layered microchannel heat sinks with different cross-sectional shapes. Entropy 21(1): 1-16.
- Alfaryjat AA, Mohammed HA, Adam NM (2014) Influence of geometrical parameters of hexagonal, circular, and rhombus microchannel heat sinks on thermohydraulic characteristics. International Communications in Heat and Mass Transfer 52: 121-131.
- Yariv E, Ben Dov G, Dorfman KD (2005) Polymerase chain reaction in natural convection systems: a convection-reaction-diffusion model. Europhysics Letters 71(6): 1008-1014.
- Agrawal AP, Ali S, Rathore S (2022) Finite element stress analysis for shape optimization of spur gear using ANSYS. Mater Today Proc 64(3): 1147-1152.
- Malika M, Bhad R, Sonawane SS (2021) ANSYS simulation of a low volume fraction CuO-ZnO/water hybrid nanofluid in a shell and tube heat exchanger. J Indian Chem Soc 98(11): 100200.
- Gao Z (2023) A droplet digital PCR chip with passive bubble removal for absolute nucleic acid quantification. Sens Actuators B Chem 392: 134109.
- Maier C, Calafut T (2008) Polypropylene: The Definitive User's Guide and Databook. Amsterdam: Elsevier Science.
- Bair HE (1981) Hydrolysis of polycarbonate. J Appl Polym Sci 26(6): 1777-1786.
- Su S (2019) One-step bonding and hydrophobic surface modification for polycarbonate-based droplet microfluidic chips. Sens Actuators B Chem 282: 60-68.
- Guler MT (2022) Fabricating plasma bonded microfluidic chips by CO2 laser machining of PDMS. J Manuf Process 73: 260-268.
- Jiang B (2022) Microscale investigation on wettability and bonding mechanism of oxygen plasma-treated PDMS microfluidic chip. Appl Surf Sci 574(1): 151704.
- Tullius JF, Tullius TK, Bayazitoglu Y (2012) Optimization of short micro pin fins in minichannels. International Journal of Heat and Mass Transfer 55: 3921-3932.
- Salimpour MR, Sharifhasan M, Shirani E (2011) Constructal optimization of the geometry of an array of micro-channels. International Communications in Heat and Mass Transfer 38(1): 93-99.
- Alihosseini Y, Zabetian Targhi M, Heyhat MM (2021) Thermo-hydraulic performance of wavy microchannel heat sink with oblique grooved fins. Applied Thermal Engineering 189: 116719.
- Tsai YS (2022) Electromagnetically driven integrated microfluidic platform using RT-LAMP for SARS-CoV-2 detection. Anal Chim Acta 1219: 340036.
- Lee SY, Kim JH, Oh SW (2022) Combination of filtration and immunomagnetic separation based on real-time PCR for foodborne pathogen detection. J Microbiol Methods 201: 106577.
- Brons JK (2020) Fast identification of E. coli in urinary tract infections using a virulence gene-based PCR. J Microbiol Methods 169: 105799.
- Zhang M (2020) Selective endpoint detection of Vibrio parahaemolyticus with CRISPR/Cas12a assisted PCR. Talanta 214: 120818.
- Nabuti J (2023) Highly efficient photonic PCR system based on plasmonic heating of gold nanofilms. Biosens Bioelectron X 14: 100346.
- Higashi T (2015) Nanomaterial-assisted PCR based on thermal generation from magnetic nanoparticles under AC magnetic fields. Chem Phys Lett 635: 234-240.
- Wu C (2022) TriD-LAMP: a pump-free microfluidic chip for duplex droplet digital LAMP analysis. Anal Chim Acta 1233: 340513.
- Parvizi F, Parvareh A, Heydari R (2023) Fabrication of a hydrophobic surface as a new supported liquid membrane for microfluidic liquid phase microextraction. Microchem J 189: 108514.
- Zhou Z (2023) Portable dual-aptamer microfluidic chip biosensor for Bacillus cereus. J Hazard Mater 445: 130545.
- Cao X (2022) A dual-signal amplification strategy based on pump-free SERS microfluidic chip for lung cancer ctDNA detection. Biosens Bioelectron 205: 114110.
- Su W (2022) A microfluidic cell chip for virus isolation via rapid screening for permissive cells. Virol Sin 37: 547-557.
- Lu Y (2022) Multiple exosome RNA analysis methods for lung cancer diagnosis via integrated on-chip microfluidic system. Chin Chem Lett 33(6): 3188-3192.
- Gale BK (2016) Low-cost MEMS technologies. In: Reference Module in Materials Science and Materials Engineering. Amsterdam: Elsevier.
- Açikgöz HN (2023) Assessment of silicon, glass, FR4, PDMS and PMMA for acoustic particle/cell manipulation in microfluidics. Ultrasonics 129: 106911.
- Tang Q (2021) Fabrication of a hydroxyapatite-PDMS microfluidic chip for bone-related cell culture and drug screening. Bioact Mater 6(1): 169-178.
- Huber JA (2009) Effect of PCR amplicon size on assessments of clone library microbial diversity. Environ Microbiol 11(5): 1292.
- Uribe MR (2003) An alternative PCR-based method for monitoring thermal cycler performance. Hum Immunol 64: S177.






























