Assessing Fungal and Bacterial Microbiome Diversity in The Black Soldier Fly (Hermetia illucens L.) Gut and Its Different Feeding Media
Noha Shokry1, Moustafa M Eldakak2, Esmat Hegazi1*, Abdel Aziz M Nour3 and Mustafa Alseqely4
1Department of Entomology, Alexandria University, Faculty of Agriculture, Egypt
2Department of Genetics, Alexandria University, Faculty of Agriculture, Egypt
3Department of Animal and Fish Production, Alexandria University, Faculty of Agriculture, Egypt
4Department of Microbiology and Immunology, Alexandria University, Faculty of Pharmacy, Egypt
Submission:March 06, 2023; Published: March 21, 2023
*Corresponding author: Esmat Hegazi, Department of Entomology, Alexandria University, Faculty of Agriculture, Egypt
How to cite this article: Noha Shokry, Mostafa M El Daqaq, Essmat Hegazy, Abd El Azize M Nour and Mustafa Alseqely. Assessing Fungal and Bacterial Microbiome Diversity in The Black Soldier Fly (Hermetia illucens L.) Gut and Its Different Feeding Media. Adv Biotech & Micro. 2023; 17(2): 555958 DOI:10.19080/AIBM.2023.17.555958
Abstract
Now is the time to turn waste from a source of environmental pollution into a treasure of abundance throughout the world, these concerns are even more important when alternative feed ingredients, new product developments and innovative feeding trends, like insect-meals, are considered. The black soldier fly is the key to our food secure future, mainly as an alternative protein source. The research on transfer of contaminants from different substrates to the insects, as well as the possible occurrence of toxin-producing fungi and contaminated bacteria in the gut of non-processed insects are very limited. Accordingly, we investigated the impact of the substrate/diet on the intestinal microbiota of H. illucens larvae. The results were Impressive to make us sure that this incredible insect lead us for the way of our food secure. We had found many fungai which can decay harmful bacteria and toxin-producing fungi as Ascomycete fungi and Pichia fungi. Moreover, many species bacteria, and fungi for the first time they were isolated from the larvae of the black soldier fly for example, the bacterium Ignatzschineria larvae, species of fungi Pichia sporocuriosa, Pichia cecembensis, and Candida thaimueongensis have not been isolated from the intestines of the larvae before. So, in this paper you will find the answers of the most two important questions: first, Do the feeding food that the larvae fed on affect the micro biotic community within the larval intestines?
Second, how diverse are these microbes inside the intestines, whether fungi or bacteria? I wish an enjoyable and interesting reading of the rest of the research paper that opens the horizon for you to continue after us the road to the future of food security for the whole world.
Keywords: Black soldier Fly; Larval gut; Biowastes; Fungi; Bacteria; First isolation; Metabarcoding
Introduction
The past decade has seen the rapid expansion of the industrial insect rearing sector in response to the global increasing demand in high-quality protein for human consumption [1]. One of the economically most important and most promising farmed insect species is the black soldier fly (Hermetia illucens; Diptera: Stratiomyidae). The black soldier fly (BSF), Hermetia illucens (L.), is a promising insect for organic waste management. BSF larvae (BSFL) can be reared on a large scale and feed on various organic wastes, such as animal manure, plant materials, and food waste [2]. Recently, they have been used globally for recycling organic waste into insect biomass [3]. The BSFL has a life cycle which comprises five stages. These are namely, egg, larvae, pre-pupae, pupae, and adult. In fact, the entire life cycle of the BSFL is principally contributed by both stages of the larval and pupal. On the other hand, according to [4], the adult and egg hatching stages of the BSFL life cycle are characterized by being comparatively short. The feeding activity of BSFL remarkably reduces huge volumes of organic waste and environmental pollutants [5,6]. Treating livestock manure with BSFL significantly reduces the numbers of pathogenic bacteria, such as Salmonella [7] and Escherichia coli [8] in feces.
As a saprotrophic insect, the BSF lives in environments that, in general, have high microbial loads. How the BSF defends itself against pathogenic microorganisms and interacts with intestinal microbes remains largely unknown. Jeon et al. first reported the intestinal bacterial community of BSF fed with food waste using pyrosequencing [9]. Zheng et al. [10] analyzed the dynamic changes of bacterial diversity in the gut of BSF for the first time and showed that the most dominant bacterial phyla associated with the BSF are Bacteroidetes and Proteobacteria. Bacteria in the gen-era Bacteroides, Dysgonomonas, Providencia, Sanguibacter, and Sphingobacterium were found in larvae, prepupae, and pupae. Providencia spp. were probably transmitted vertically as they were present in all life stages of the BSF. As reported in different studies, the bacterial compositions in the so-called core microbiome of BSF were different. Nevertheless, how do we define the core microbiome in the guts of BSFL, and do these microbes play important roles in BSF growth and development? Using bacteria isolated from different insects, Zheng et al. showed that bacteria might influence the attraction and oviposition of the adult BSF [11].
There are currently only a few studies describing the effect of BSFL on the fungal component of substrates. Thus, the mycobiome of chicken manure fermented by BSFL has been investigated by [12] the rearing of BSFL results in a sharp decrease in the abundance of phytopathogenic and endophytic fungi. We assumed that such changes would be even more pronounced in the substrates with a predominance of plant components. Such substrates are commercially attractive for the industrial rearing of BSFL. However, the abundance of adverse endophytic fungi of genera such as Alternaria, Pyrenophora, Ustilago, and Nigrospora can significantly complicate the practical use of BSFL biomass and the processed substrates. The thorough sterilization of substrates represents a major and difficult task, which is very expensive and time consuming. A lot of microorganisms (including yeasts) are also beneficial as probiotics for the subsequent use of composts. Therefore, it seems extremely important to receive comprehensive data on the fungal community, not only regarding the larval biomass but also representatives of the fungal community should be attributed to the species level (usually obtained in metabarcoding) which can be achieved by either shotgun sequencing or a combination of metabarcoding and culture based techniques [13].
With all these benefits and environmentally friendly features, the BSF is the most exploited species worldwide in the growing insect farming industry [14].
When the idea of this research paper was manifested, the first aim was recognizing the variation between the microbial load in the larval feeding media and in the larval gut. Second to ensure that these larvae are clean and that they are a safe source of feed and food, whether for poultry, animals or even humans, and that they help effectively build a new thought and a new food culture for all countries around the world, which will change people’s view of new food sources, even if they are from an unfamiliar insect source, especially in the Middle Eastern countries, including Egypt. The impact of this study is starting actual projects based mainly on the production of black soldier fly larvae to spread a new food culture that God has blessed us with from a simple source but infinitely rich in many outputs that will also be based on many vital and important industries for our people and the whole world.
Materials and Methods
Insect rearing
The Black Soldier Fly has been bought from another researcher at faculty of agriculture from Alexandria university and reared at department of applied entomology and zoology in September 2020. The larvae reared on plastic plates at 27-30°C and 60-65% humidity. The prepupal stage featured by auto harvest to anther dry plates to prepare itself for the pupal stage that has been transferred to 40*40cm cage. When the adults get emerged the cage get supported by cotton socked on water, attractive media for females, wooden sticks with interspace 1-1.5cm as an egger. The egg was collected every day and transfer the clutches to wet brane 1-1.5 brane to water in a plastic plate.
The culture media

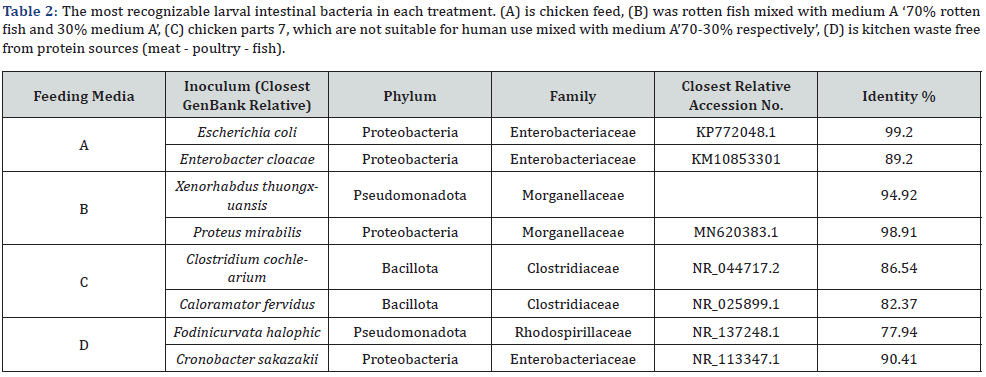
Four different media were used during these experiments. The first medium (A) is chicken feed from Al-Iman Feed Company, with 21% protein content. The second medium (B) was rotten small tilabia fish from the waste of fish farms, mixed with medium A 70% rotten fish and 30% medium A. The third medium (C) is the chicken parts which are not suitable for human use e.g., nonedible meat parts and guts mixed with medium A 70-30% respectively, and I bought them from poultry sellers in the residential area where the researcher lives. The last medium (D) is kitchen waste free from protein sources (meat - poultry - fish) which the researcher used to save from the home kitchen waste. The following figure shows then main components of each type of waste. (Figure 1).
Biological experiment
The feeding of larvae was initially on different food environments starting from the second larval stage, as it is the first larval stage that can be discerned with the naked eye. Five replicates were made for each treatment. Each replicate contains 20 larvae placed in small transparent plastic dishes, 6 cm in diameter and 3 cm in height and they are covered with black cloth. The food presented with a weight of 7 g/replicate in each treatment starting from the second larval age and is increased until the sixth larval age to reach 14 g food. The symbol (A`) was given to the larvae that fed on the first medium, which is chicken feed, while the symbol (B`) was given to the larvae that fed on the second medium, which is a mixture of rotten fish and poultry feed, while the symbol (C`) was given to the larvae that fed on the third medium, which is the mixture between rotten chicken meat and poultry feed, and finally the symbol (D`) was given to the larvae that fed on the kitchen waste medium devoid of any source of protein.
Microbial community isolation from the larval gut and different feeding media
BSFL sterilization: Twelve larvae were collected from each food environment at six instars. They were washed from the outside with sterile distilled water and then with 70% alcohol to ensure that there is no contamination of the intestine caused by any external bacteria. Then, they were dissected by binocular dissecting microscope at 4x magnification power, where the body wall of the larva was cut from both sides. After that the larva was fixed from the front and end of the body with a dissection pin, and the upper layer of the body wall was removed, then the digestive tract was removed from its two ends (the first at the esophagus after the oropharyngeal cavity and the second from the end of the rectum at the colon). Finally, 12 digestive canals are isolated, crushed, and placed in the LB medium.
Bacterial community isolation: A LB medium (Luria Broth) was prepared by dissolved 4 grams of LB: 200 ml of distilled water. They were distributed equally into eight glass flasks of 120ml volume, so that each flask contains a 25ml LB medium. Then this medium was sterilized in an autoclave for 20minutes at 121°C. Next, the media was left in incubation for 72 hours until it is ensured that we follow good sterilization procedures. Then, the larval guts (A`, B`, C`, and D`) and a sample of the feeding media (A, B, C, and D) of sterilized of BSFL was transferred to the flasks that contain the LB media under sterilized conditions in a laminar air follow caped. Then, the eight groups were inculpated at 28°C for 48 hours as to allow the growth of bacterial communities as to be ready to DNA extraction.
Fungal community isolation: A Potato Dextrose Agar (PPA) environment was prepared by a 20 grams of potato Dextrose agar was mixed with 75ml of sterile distilled water, and they were sterilized. Then we made 250ml environment rent they were poured into eight Petry dishes and left in the incubator for 72 hours to ensure that no contamination would occur and the safety of the environment before transplanting. Then, the larval guts (A`, B`, C`, and D`) and a sample of the feeding media (A, B, C, and D) of sterilized of BSFL was transferred to PPA media and they have been incubated for one week at 28°C. After this period, the fungi growing on the media then, were isolated in 1.5ml Eppendorf tubes and the DNA of the fungi growing in the liquid environment was isolated by following protocol B from the kit’s instruction book.
DNA metabarcoding of the microbial communities in the larval gut and the feeding media
Genomic DNA Extraction was made using G-spin™ Total DNA Extraction Kit according to manufactory instructions of protocol B for the bacterial community and fungal community grown on LB media and PPA respectively for ether the larval gut or the feeding media. The purity and concentration of the resultant DNA was determined using a Nanodrop 2,000/2,000c spectrophotometer (Thermo Fischer Scientific, Wilmington, United States). PCR, and Sequencing of the media Before being rinsed with distilled water, each individual insect larva was surface sterilized with 70% alcohol. The Isolate II Genomic DNA Kit (Bioline, London, and the United Kingdom) was used to extract genomic DNA in accordance with the instructions provided by the manufacturer.
PCR was performed to amplify the 16s rRNA region for bacterial DNA and ITS barcode region for fungal DNA in a total reaction volume of 20μL containing 2x topsimple dymixntaq (enzynomics), 10pmol/μl of primers16s515FGTGCCAGCMGCCGCGG, 16s907R CCGTCAATTCMTTTRAGTTT, ITSF CTTGGTCATTTAGAGGAAGTAA and ITS R TCCTCCGCTTATTGATATGC as shown at table 1 and 50ng/μl of DNA template in a peqlab primus 25 advanced thermal circular Machin (Germany). The following cycling conditions were used: initial denaturation for 5 min at 95°C, followed by 40 cycles of 1 s at 95°C, 1 s annealing at 52°C, extension for 1 min at 72°C, and a final elongation step of 10 min at 72°C. Purified PCR products were shipped to Macrogen Europe BV (Meibergdreef, Amsterdam, Netherlands) for bi-directional sequencing. Each PCR product for each sample was sequenced by two different sequencing reactions to identify the two most prominent microbes in each sample. Retrieved sequences were aligned to NCBI database. (https://www.ncbi.nlm.nih.gov) using a nucleotides blast suite (blast in) from blast tools (Basic Local Alignment search tool) for identifying the most prominent species to microbial community either bacteria or fungi on larval gut and feeding media.
Phylogenetics analyses
Four Phylogenetics trees were constructed.
a) The most prominent bacterial species of the bacterial community on the feeding media.
b) The most prominent bacterial species of the bacterial community on the larval gut.
c) The most prominent fungal species of the fungal community on the feeding media.
d) The most prominent fungal species of the fungal community on the larval gut.
e) Combined the most prominent fungi and bacteria had been found in this study.
Using the MEGA11 software (molecular evolutionary genetics analysis) as to analyze the Phylogenetic relationship between the microorganisms in each group. The evolutionary history was inferred using the UPGMA method [15]. The evolutionary distances were computed using the Maximum Composite Likelihood method [16] and are in the units of the number of base substitutions per site. This analysis involved 8 nucleotide sequences. All ambiguous positions were removed for each sequence pair (pairwise deletion option). There was a total of 440 positions in the final dataset. Evolutionary analyses were conducted in MEGA11 [17].

Results
The feeding media bacterial core
The most prominent bacterial species of the bacterial communities in the four different feeding media were identified by using 16S rRNA sequence alignment shown in table 1 as follows Starting with the first medium in which the larvae were fed, the most defined bacterium is Escherichia coli with sequence identity of 99.20%. The second kind of bacteria at the same medium A is Enterobacter cloacae that identified with 89.20%. In the feeding medium B (which was a rotten fish unfit for human use mixed with chicken feed). The most defined bacterium was Xenorhabdus thuongxuansis that is a genus of gram-negative, motile bacteria that belong to the Morganellaceae family. When moving to the third feeding medium on which the larvae feeded, we find the two types Calostridium cochlearium and Caloramator fervidus They are the most defined in the feeding medium with rates of 86.54% and 82.37% respectively, which was a mixture of rotten chicken unfit for human use and chicken feed. the last medium is the kitchen food waste without any protein resources and the most identification bacteria were Cronobacter sakazakii and Fodinicurvata halophic with 90.41% and 77.94% respectively according to (Table 2 & Figure 2)

The intestinal larvae core bacteria
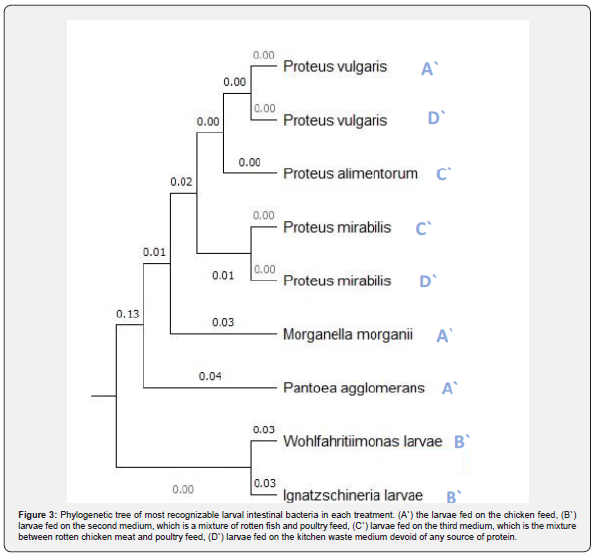
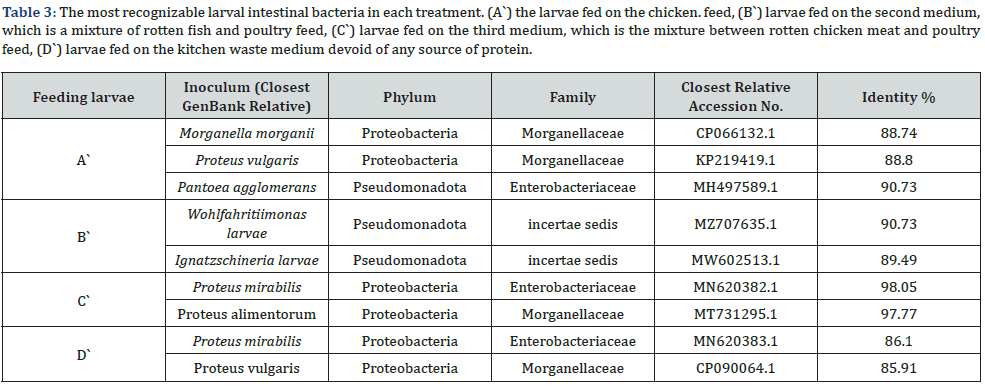
We have found some verities between the different treatments in the gut core bacteria which almost belong to Pseudomonadota and Proteobacteria phyla as showed at Table 3. Four groups of samples were identified by using 16S rRNA sequence alignment as follows: the group of treated larvae A` was identified as Patoea agglomerans with 88.80% identity, Morganella morganii with 88.74% identity, and Proteus vulgaris with 88.61% identity. While the treated larvae’s B` samples showed 90.73% identity for Wohl-fahritiimonas larvae and 89.49% for Ignatzschineria larvae and it is the first time isolate this bacterium from the BSFL gut. In the group of treated larvae’s C` sample, the bacteria Proteus mirabilis was identified with 98.05% and 97.77% identity for Proteus alimentorum. The last sample D` was also identified with Proteus mirabilis as in larvae treated with treatment C` but with 86.10% identity and Proteus vulgaris as in larvae treated with treatment A` but with 85.91%. (Table 3 & Figure 3).

The feeding media fungal core
Four groups of samples were identified by using ITS rRNA sequence alignment shown at (Table 1) as follows: As shown in table 4 the most identification at feeding medium A fungi were Cyphellophora phylostchydis and Pichia sporocuriosa with 100% identity ratio and 95.43% for Pichia sporocuriosa. Then, Aspergillus fugnigatus in treatment B and C with identity ratio 99.83% and 98.48% respectively. Then at treatment B we had found Penicillium ornatum that belong to Family Trichocomaceae with identity ratio 90.23% and 88.60% respectively and Hamigera brevicompacta in feeding media C 88.60% identity ratio. Finally, the Pichia paraexigua we had found it at medium D with 97.87% and 96.72% identification ratio respectively (Table 4 & Figure 4).


The intestinal larvae core fungi
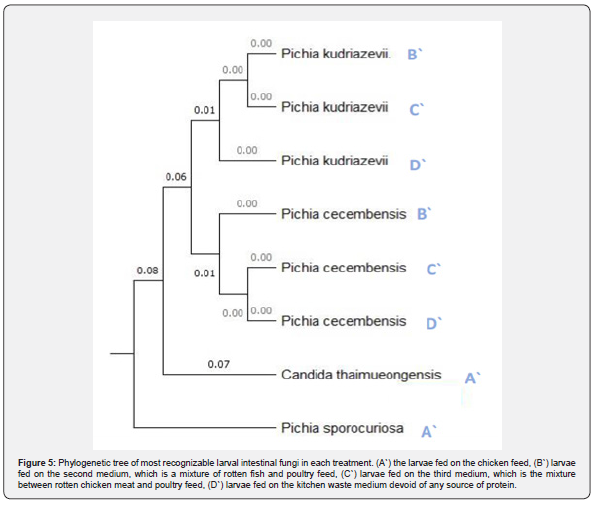
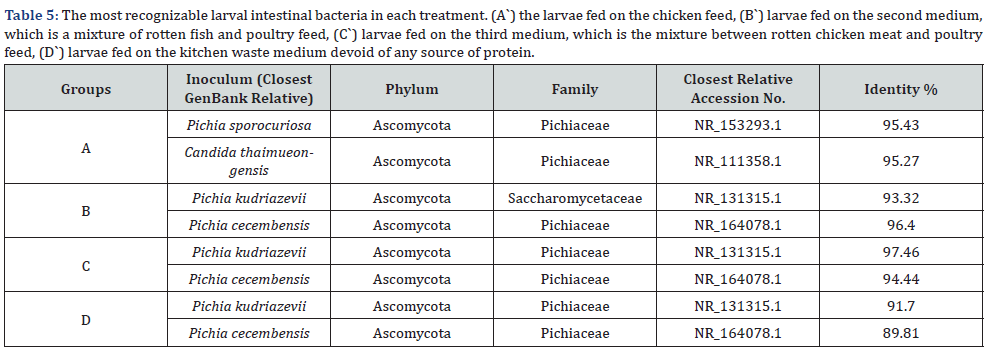
According to table 5 the larvae that reared on treatment A` had two most identified fungi Pichia sporocuriosa and Candida thaimueongensis with 95.43% and 95.27% identification ratio, both belong to Ascomycota phylum and pichiaceae and Saccharomycetaceae family respectively and they are first time isolated from the Black Soldier Fly Larval gut. But if we talk about the genus of pitchia which has been repeated by different species in all treatments it is a genus of yeasts in the family Pichiaceae. First the Pichia kudriazevii that that had been found at each group of treated larvae B`, C`, and D` with identity ratio 93.32%, 97.46% and 91.70% respectively. Second the Pichia cecembensis not only repeated at each group of treated larvae B`, C`, and D` with high identity ratio 96.40%, 94.44% and 89.81% respectively but although first time isolated from the Black Soldier Fly Larval gut (Table 5 & Figure 5).
Discussion
The feeding media bacterial core
In the feeding medium B which was a Rotten fish unfit for human use mixed with poultry feed the most defined bacterium was Xenorhabdus thuongxuansis that is a genus of motile, gram-negative bacteria from the family of the Morganellaceae. The flowing studies explained how the feeding media containing many entomological pathogenic bacteria, The biological control agents used against Aedes mosquitoes include Bacillus thuringiensis israelensis or B. sphericus and their toxins and Xenorhabdus/Photorhabdus [18,19] Xenorhabdus and Photorhabdus have potential for the biological control of Aedes mosquitoes. These entomopathogenic bacteria are used in the control of mosquito larvae [20,21]. Xenorhabdus and Photorhabdus have been successfully used to reduce the development of several insect pests in laboratory conditions [22]. The Calostridium species are obligate or facultative anaerobic bacteria, producing endospores that are highly resistant to heat and other environmental factors [23,24]. Some Clostridium species are well-known pathogens [25]. The Cronobacter sakazakii which before 2007 was named Enterobacter sakazakii [26,27] is an opportunistic Gram-negative, rod-shaped, pathogenic bacterium that can live in very dry places, otherwise known as xerotolerance. Fodinicurvata halophic a Gram-stain-negative, rodshaped, facultatively anaerobic, moderately halophilic bacterium the genus Fodinicurvata belongs to the family Rhodospirillaceae, within the order Rhodospirillales [28] of the class Alphaproteobacteria.
The intestinal larvae core bacteria
The microbes in the fly larvae gut have multiple functions that are important to larval development [6] The functions of gut microbiota impact the development, pathogen resistance, nutrition, and physiology of the host. Insects’ unique intestinal biotransformation system is poorly understood, particularly in terms of how the various symbiotic microorganisms in the intestine function. The potential science and application values, as well as the relationships between insects and symbiotic microorganisms, have been rationalized; However, with regard to the study of mammal’s gut bacteria [29] there remains much room for development concerning the compounds from insect gut microbes.
We now move on to the treated larvae groups which had been fed on the feeding media we talked about earlier. The Patoea agglomerans that a gamma proteobacterium of plant origin, possesses many beneficial traits that could be used for the prevention and/or treatment of human and animal diseases, combating plant pathogens, promotion of plant growth and bioremediation of the environment. It produces a number of antibiotics (herbicolin, pantocins, microcin, agglomerins, andrimid, phenazine, among others) which could be used for combating plant, animal and human pathogens or for food preservation [30]. In accordance with our study, the scientist Zhu and others confirmed the significance of larval gut microorganisms in the degradation of biowaste, the strains of P. agglomerans could also be useful in the acquisition of energy from various alternative sources, such as waste recycling. An example is the salt-tolerant P. agglomerans BH-18 strain isolated by Zhu et al. [31] from mangrove sludge, which possesses the ability to produce hydrogen.
The authors proposed using this strain for the biological treatment of marine aquaculture wastewater and marine organic waste, associated with the production of biohydrogen that represents a promising alternative source of energy due to its reproducibility, non-polluting nature, and high energy yield. In a subsequent study, the authors demonstrated that the yield of hydrogen could be significantly greater (by 36.94%) if the P. agglomerans BH-18 strain is used in a mixed culture with a Candida tropicalis BH-6 salt-tolerant strain, which had been isolated from the same mangrove ecosystem[32] be noted that we have found the genes candida within the fungal load rang in A larval intestinal tested by its primer Candida thaimueongensis which explain the interaction between the gut microbes. the acidic antibiotics called agglomerins which are moderately active against a wide variety of anaerobic bacteria (including Clostridium difficile, C. perfringens, Propionibacterium acnes) and weakly active against aerobic Gram-positive bacteria (including Streptococcus pyogenes, S. pneumoniaz [33]; the pseudopeptide antibiotic andrimid which is active against both Gram-negative and Gram-positive bacteria, including methicyllin-resistant Staphylococcus aureus (MRSA), vancomycin-resistant Enterococcus (VRE) and Klebsiella pneumoniae, and displays also limited antiproliferative activity against human tumor cell lines [34].
D-alanylgriseoluteic acid (AGA), a potent phenazine antibiotic produced by P. agglomerans strain Eh1087 which reveals a broad spectrum of antimicrobial activity and is particularly active against Gram-positive pathogens, such as Streptococcus pneumoniae [35,36]. the recently isolated in Korea phenazine antibiotic from P. agglomerans apple strain R190, active against various spoilage bacteria, including Pectobacterium carotovorum subsp. carotovorum, Clavibacter michiganensis, and Burkholderia andropogonis, as well as against foodborne pathogens such as Escherichia coli O157:H7 and Salmonella enterica, and other human pathogens such as Klebsiella pneumoniae and Yersinia enterocolitica [37]. To review the rest of the results of the bacteria, these bacteria can be considered to explain to us how the different types of bacteria found in organic waste are digested and it is also a confirmation that the larvae of the black soldier fly are amazing in treating organic waste.
Now we will discuss the species of the genus Proteus, which were common in the intestines of larvae that fed on the four environments of organic waste and the extent of their importance for the insect to digest the pathogens in these wastes, which are a source of environmental pollution and thus negatively affect human and animals’ health. Microorganisms belonging to the genus Proteus were first described in 1885 by a German microbiologist Gustav Hauser, who had revealed their ability to swarm on solid surfaces. The name Proteus came from Homer’s BOdyssey^ and its character Proteus, who could change his shape and had an ability of endless transformation. Hauser described two species of the genus: Proteus vulgaris and Proteus mirabilis [38]. It is postulated that human intestines are a reservoir of Proteus bacteria, especially those belonging to prevailing P. mirabilis species, and they are members of natural fecal microflora of several percent of human population so [39] reported the presence of Proteus spp. bacteria (one P. mirabilis and one P. vulgaris strain) in fecal samples from 4 % of healthy Spanish volunteers. [40] explained that not only rats but also many wild and domestic animals (mammals, birds, reptiles, amphibians, insects, and Seafood) are the hosts of Proteus spp. bacteria. At the same time, P. mirabilis protects the larvae from harmful microorganisms because it is antagonistic to some bacteria that maggots remove from wounds [41].
One of the most interesting kind of bacteria was Ignatzschineria larvae that first time isolated from BSFL and not discussed its importance to BSFL well yet [42]. Describe the Ignatzschineria spp as a genus of aerobic, gram-negative, non-sporeforming, non-hemolytic rod-shaped bacteria that belongs to the class Gammaproteobacteria. Three recognized species: I. indica, I. larvae, and I. ureiclastica which are commonly recovered from the larvae of parasitic spotted flesh fly Wohlfahrtia magnifica found in Europe, Asia, and North Africa [43]. The Ignatzschineria spp. thrive in the digestive tract of the larvae with Providencia [43]. Ignatzschineria is a recently identified genus of bacteria that has been isolated from the digestive tract of multiple flies associated with decomposing tissue. Species within this genus are rarely implicated in human disease, and less than 10 cases worldwide have been documented in the literature. So, we can say that this bacterium is benefit to both larvae and humane health and recovery.
The Wohlfahrtiimonas larvae has been isolated to the first time in 2014 by the scientist lee [44] and he said that A novel, Gram-negative, facultative anaerobic, motile and short rod-shaped bacterium, strain KBL006T was isolated from the larval gut of Hermetia illucens, Black soldier fly The description of genus Wohlfahrtiimonasis followed: Gram-negative, straight, short-rod shape. Catalase and oxidase reactions are positive. Main fatty acids are C18:1andC14:0. Main polar lipids are phosphatidylglycerol, phosphatidylethanolamine, and phosphatidylserine [44]. demonstrating the significance of the bacteria in the larval gut for human health as well as for larvae, the research Maggot Wound Therapy Associated with Wohlfahrtiimonas chitiniclastica. Blood Infection by [45] who tested that the Maggots secrete defensins which are proteins like those produced by circulating human white blood cells that may result in the potential antimicrobial properties of maggot therapy.
The relationship between the bacterial load on feeding media and larval gut
The relatively abundant bacteria and the dynamics of bacterial compositions during the development of BSFL fed with a food waste diet and an oil waste diet were different compared with BSFL fed with a chicken feed diet. Imputation of metabolic pathways indicated that metabolic capability might be the key factor for the changes of intestinal bacteria [46]. Unlike the other studies, no significant differences among the gut microbiome of BSFL fed with 3 different diets (chicken feed, freshly cut grass, and fruit/vegetables) were observed, indicating that a core microbiome (Actinomyces, Dysgonomonas, Enterococcus, and another unclassified Actinomycetales) exists in the gut of BSFL Klammsteiner et al. [47]. A survey of the dynamic changes of bacteria in the intestinal tract of H. illucens showed that the gut microbiome was relatively stable over the course of larval development, whereas the microbiome in the feed residue changed significantly. The core microbiome includes Citrobacter, Enterobacter, Klebsiella, Morganella, Proteus, and Providencia. Dysgonomonas was also dominant, except in the early larval stage.
Among the 6 bacteria studied, Proteus (BSF4) was the exception. The larval and prepupal weights of BSF inoculated with this bacterium were even less than those of the germ-free group. The prepupae and eclosion rates were also very low. The durations of the larval and pupal stages were even longer than those of the germ-free BSF, indicating that this bacterium did not promote BSF growth and development. This result conflicts with a study on the management of chicken manure with the combination of BSF and Proteus, in which Proteus promoted BSFL weight gain Mazza et al. [48]. The differences may be linked to the distinct diets or substrates used in the 2 different studies. The interactions between Proteus and the complicated microbial community in chicken manure may be beneficial for BSF growth and the biotransformation process. Previous studies have shown that the microbes in the guts of BSFL interact with the microbial community in the substrate after feeding to form a new intestinal microbial community [49]. This new community was beneficial to BSFL growth and development and the biotransformation of the substrate [50, 51]. Our study support strongly This phenomenon provides further evidence that the interactions between BSFL and their feeding environment led to the formation of a specific microbial community structure, which might be beneficial for BSF growth and development [46] (Table 6).
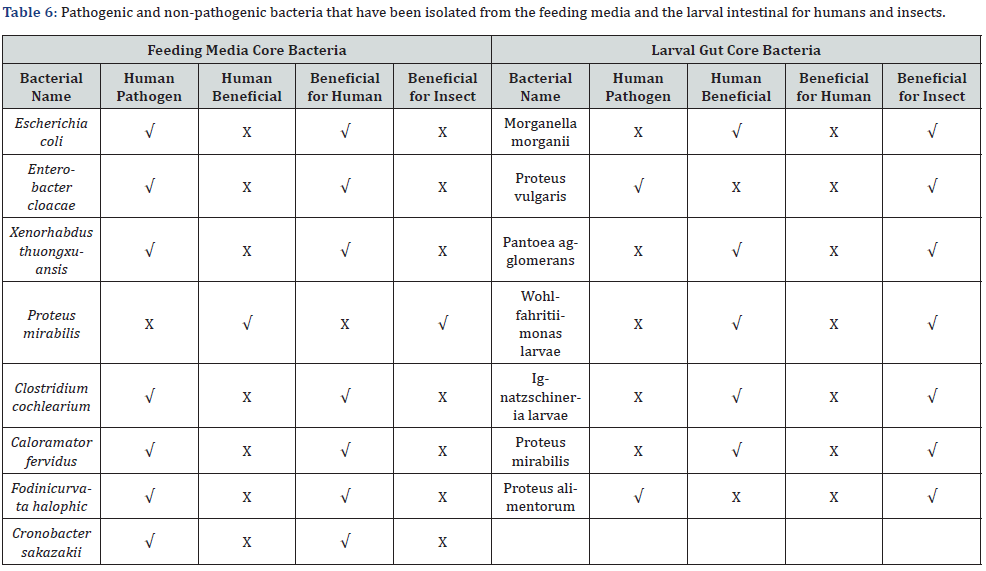
The intestinal larvae core fungi
Then we noted the existence abundance and remarkably the Ascomycete fungi that attack bacteria and produce these fungi antibiotic penicillin and is used during organ transplantation to prevent cellular rejection and the manufacture of cheese Roquefort produces important enzymes in the dairy industry and food preservation. Yeasts of the genus Candida, one of the largest genera in terms of numbers of species, are widely distributed in nature. Species of this genus have been isolated from various sources in terrestrial and aquatic habitats [52-54]. Reported that the name Candida thaimueangensis sp. nov is proposed to accommodate these new strains. But if we talk about the genus of pitchia which has been repeated by different species in all treatments it is a genus of yeasts in the family Pichiaceae with spherical, elliptical, or oblong acuminate cells. Pichia is a teleomorph, and forms hat-shaped, hemispherical, or round ascospores during sexual reproduction.
Pitchia sporocuriosa is first time isolated from the gut of BSFL. Pichia kudriavzevii is an eurybiont, involved in the fermentation of many natural substrates. It is not a typical pathogen, although it has been reported previously that, in rare cases, this yeast has taken part in candidal vaginitis, mastitis, and candidal arthritis [55]. Finally we would like to refer to the larval hemolymph exerts antimicrobial effects on various pathogens [56]. BSFL extracts, consisting of homogenized larvae with buffer, were shown to inhibit the growth of several Gram-negative bacterial pathogens including Neisseria gonorrhoeae, Klebsiella pneumoniae, and Shigella sonnei [57]. These bacteria were among the bacterial load in the media that the larvae fed on in our study. Both aqueous and methanol- based BSFL extracts exhibited antimicrobial effects against both Gram-negative and Gram-positive human pathogens [58].
The relationship between the fungal load on feeding media and larval gut.
The fungi composition may change in this type of environment through time due to the composting process [59]. they found constancy in the dominance of Candida in most of the environments and larval guts. These results harmonized with previous analyses of the gut fungal community composition of BSF larvae that were fed on agricultural waste [60,61] and of the community composition of the compost environment that was treated with BSF [13]. In these cases, the most abundant fungal generium in the BSF was Pichia which were abundant in our study. This study suggests that the fungal community structure is affected from a ‘core’ fungal community as found in related studies of the bacterial community composition of the BSF [62, 63]. We found that only Pichia presented a high relative abundance and prevailed across all the substrates; the rest of the identified fungal communities were highly substrate specific. The high prevalence and dominance of Pichia in this study as well as in that of [60]. points toward a stable association with BSF larval gut. P. kudriavzevii, the most prevalent species found in our samples, has been reported to encode the antibacterial toxin RY55 that is active against several human pathogens such as E. coli, Enterococcus faecalis, Klebsiella sp., Staphylococcus aureus, Pseudomonas aeruginosa and Pseudomonas alcaligenes [60] (Table 7).
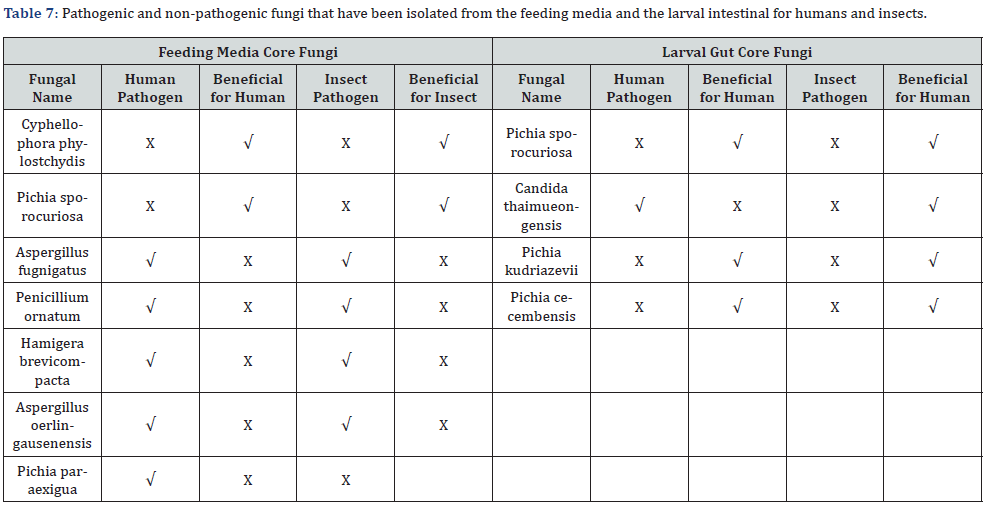
When verifying the results in the rest of the transactions according to (Figure 3 & 5) we find the presence of the same genera and even the same types of fungi inside the stomach of the larvae, which confirms the stability of the fungus community inside the intestines, unlike the bacterial environment, in which the genera and species varied, and this makes us seek to link these results to each other to understand what is the importance of this variation for the larvae? Does it affect the composition of fats and proteins of larvae? Is it this diversity of bacteria and the persistence of fungi that magnifies the value of these larvae and the richness of their precious outputs, which humans have not yet exploited for the richness of the world?
Our interpretation as researchers for this research paper is that yes, all of the above studied on larvae and bloom for us these differentiated results of the bacterial community and similar to the fungus community is what gives these larvae their value and distinguish their outputs and their ability to convert surplus organic waste from human or animal use and polluting the environment into wealth from an insect source and to realize that we as humans have within our intestines bacterial and fungal communities that serve our biological development and help us enjoy good health and without them or even the occurrence of any defect In its rates inside our intestines we become sick we suffer from dysfunction of our bodies as well as the insect (Figure 6).

Conclusions
We have found out how miraculous it was to introduce organic rich waste in bacterial and fungi pathogens, whether for the insect itself or for humans and animals, or even as source of polluting emissions for the environment and digested into rich source of protein and oils that has potential in food and pharmaceutical industry. The results indicated that intestinal bacteria influenced BSF growth and development significantly and provided insights into using these beneficial microbes to promote BSF applications in the waste management industry. We hope in the coming period to expand worldwide in the form of productive farms of these amazing larvae full of Treasures that have not yet been appreciated by humans, which will make a big progress in the field of natural industry once we pay attention to them and use them in our lives and recognize their value. This research and previous studies of this insect are an encouraging a powerful start with great creature such as this insect for a clean, healthy, and rich environment.
References
- van Huis A (2016) Edible insects are the future? Proceedings of the Nutrition Society 75(3): 294-305.
- Sheppard DC, G Larry Newton b, Sidney A. Thompson, Stan Savage (1994) A value added manure management system using the black soldier fly. Bioresource technology 50(3): 275-279.
- Tomberlin J, A Van Huis (2020) Black soldier fly from pest to ‘crown jewel’of the insects as feed industry: an historical perspective. Journal of Insects as Food and Feed 6(1): 1-4.
- Kaya C, Tomas N Generalovic, Gunilla Ståhls, Martin Hauser, Ana C Samayoa et al. (201) Global population genetic structure and demographic trajectories of the black soldier fly, Hermetia illucens. BMC Biology 19(1): 94.
- Smetana S, Megala Palanisamy, Alexander Mathys, Volker Heinz (2016) Sustainability of insect use for feed and food: Life Cycle Assessment perspective. Journal of cleaner production 137: 741-751.
- Gold M, TomberlinJK , Diener S , Zurbrügg C, Mathys A (2018) Decomposition of biowaste macronutrients, microbes, and chemicals in black soldier fly larval treatment: A review. Waste Management 82: 302-318.
- Erickson MC, Islam M, Sheppard C, Jean Liao, Doyle MP (2004) Reduction of Escherichia coli O157: H7 and Salmonella enterica serovar Enteritidis in chicken manure by larvae of the black soldier fly. Journal of food protection 67(4): 685-690.
- Liu, Q, TomberlinJK, Brady JA, Sanford MR, Ziniu Yu (2008) Black soldier fly (Diptera: Stratiomyidae) larvae reduce Escherichia coli in dairy manure. Environmental entomology 37(6): 1525-1530.
- Jeon H, Soyoung Park, Jiyoung Choi, Gilsang Jeong, Sang-Beom Lee, et al. (2011) The intestinal bacterial community in the food waste-reducing larvae of Hermetia illucens. Current microbiology 62(5): 1390-1399.
- Zheng L, Tawni L Crippen, Baneshwar Singh, Aaron M Tarone, Scot Dowd, et al. (2013) A survey of bacterial diversity from successive life stages of black soldier fly (Diptera: Stratiomyidae) by using 16S rDNA pyrosequencing. Journal of medical entomology 50(3): 647-658.
- Zheng L, Tawni L Crippen, Leslie Holmes, Baneshwar Singh, Meaghan L Pimsler, et al. (2013) Bacteria mediate oviposition by the black soldier fly, Hermetia illucens (L.), (Diptera: Stratiomyidae). Scientific reports 3(1): 1-8.
- Zhang X, Junzhe Zhang, inlin Jiang, Xin Yu, Hongwei Zhu, et al. (2021) Black soldier fly (Hermetia illucens) larvae significantly change the microbial community in chicken manure. Current Microbiology78(1): p. 303-315.
- Kuznetsova TA, Maksim V Vecherskii, David R Khayrullin , Aleksandr A Stepankov , Irina A Maximova, et al (2022) Dramatic effect of black soldier fly larvae on fungal community in a compost. Journal of the Science of Food and Agriculture 102(6): 2598-2603.
- Generalovic TN, Shane A McCarthy, Ian A Warren, Jonathan M D Wood, James Torrance, et al. (2021) A high-quality, chromosome-level genome assembly of the Black Soldier Fly (Hermetia illucens). G3 (Bethesda) 11(5): p. jkab085.
- Schlee,D (1975) Numerical taxonomy. The principles and practice of numerical classification JSTOR.
- Tamura K, M Nei, S Kumar (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. Proceedings of the National Academy of Sciences 101(30): 11030-11035.
- Kumar S, G Stecher, K Tamura (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Molecular biology and evolution 33(7): 1870-1874.
- Kovendan K, Kadarkarai Murugan, Savariar Vincent, Siva Kamalakannan (2011) Larvicidal efficacy of Jatropha curcas and bacterial insecticide, Bacillus thuringiensis, against lymphatic filarial vector, Culex quinquefasciatus Say (Diptera: Culicidae). Parasitology research 109: 1251-1257.
- Park Y, J Kyo Jung, Y Kim (2016) A mixture of Bacillus thuringiensis subsp. israelensis with Xenorhabdus nematophila-cultured broth enhances toxicity against mosquitoes Aedes albopictus and Culex pipiens pallens (Diptera: Culicidae). Journal of Economic Entomology 109(3): 1086-1093.
- Park Y (2015) Entomopathogenic bacterium, Xenorhabdus nematophila and Photorhabdus luminescens, enhances Bacillus thuringiensis Cry4Ba toxicity against yellow fever mosquito, Aedes aegypti (Diptera: Culicidae). Journal of Asia-Pacific Entomology 18(3): 459-463.
- da Silva OS, Geronimo Rodrigues Prado, João Luiz Rosa da Silva, Carlos Eugenio Silva, Marisa da Costa, et al. (2013) Oral toxicity of Photorhabdus luminescens and Xenorhabdus nematophila (Enterobacteriaceae) against Aedes aegypti (Diptera: Culicidae). Parasitology research 112: 2891-2896.
- Ruiu L, A Satta, I Floris (2013) Emerging entomopathogenic bacteria for insect pest management. Bull Insectol 66(2): 181-186.
- Wells CL, T D Wilkins (1966) Clostridia: spore forming anaerobic bacilli. Medical Microbiology (4th ).
- Brown K (200) Control of bacterial spores. British Medical Bulletin 56(1): 158-171.
- Uzal FA, John C Freedman, Archana Shrestha, James R Theoret, Jorge Garcia, et al. (2014) Towards an understanding of the role of Clostridium perfringens toxins in human and animal disease. Future microbiology 9(3): 361-377.
- Iversen C, Niall Mullane, Barbara McCardell, Ben D Tall, Angelika Lehner, et al (2008) Cronobacter gen. nov., a new genus to accommodate the biogroups of Enterobacter sakazakii, and proposal of Cronobacter sakazakii gen. nov., comb. nov., Cronobacter malonaticus sp. nov., Cronobacter turicensis sp. nov., Cronobacter muytjensii sp. nov., Cronobacter dublinensis sp. nov., Cronobacter genomospecies 1, and of three subspecies, Cronobacter dublinensis subsp. dublinensis subsp. nov., Cronobacter dublinensis subsp. lausannensis subsp. nov. and Cronobacter dublinensis subsp. lactaridi subsp. nov. International journal of systematic and evolutionary microbiology 58(6): 1442-1447.
- Joseph S, Esin Cetinkaya , Hana Drahovska , Arturo Levican , Maria J Figueras , et al. (2012) Cronobacter condimenti sp. nov., isolated from spiced meat and Cronobacter universalis sp. nov., a novel species designation for 2 Cronobacter sp. genomospecies 1, recovered from a leg infection, water, and food ingredients Int J Syst Evol Microbiol 62(Pt 6): 1277-1283
- PFENNIG N, HG TRÜPER (1971) Higher taxa of the phototrophic bacteria. International Journal of Systematic and Evolutionary Microbiology 21(1): pp. 17-18.
- Ferrario C, Rosario Statello , Luca Carnevali , Leonardo Mancabelli , Christian Milani, et al. (2017) How to feed the mammalian gut microbiota: bacterial and metabolic modulation by dietary fibers. Frontiers in microbiology 8: 1749.
- Dutkiewicz J, Barbara Mackiewicz, Marta Kinga Lemieszek , Marcin Golec , Janusz Milanowski (2016) Pantoea agglomerans: a mysterious bacterium of evil and good. Part IV. Beneficial effects. Annals of Agricultural and Environmental Medicine 23(2): 206-222.
- Zhu D, Guangce Wang, Hongjin Qiao, Jinling Cai (2008) Fermentative hydrogen production by the new marine Pantoea agglomerans isolated from the mangrove sludge. international journal of hydrogen energy 33(21): 6116-6123.
- Zhu D, Yingchao Ma, Guangce Wang, Guanghua Pan (2015) Identification of Candida tropicalis BH-6 and synergistic effect with Pantoea agglomerans BH-18 on hydrogen production in marine culture. Applied biochemistry and biotechnology 175(5): 2677-2688.
- SHOJI JI R Sakazaki, T Hattori, K Matsumoto, N Uotani, T Yoshida (1989) Isolation and characterization of agglomerins A, B, C and D. The Journal of Antibiotics 42(12): 1729-1733.
- Pidot SJ, Sébastien Coyne, Florian Kloss, Christian Hertweck (2014) Antibiotics from neglected bacterial sources. International Journal of Medical Microbiology 304(1): pp.14-22.
- Giddens SR, Y Feng, HK Mahanty (2002) Characterization of a novel phenazine antibiotic gene cluster in Erwinia herbicola Eh1087. Molecular microbiology 45(3): 769-783.
- Giddens SR, DC Bean (2007) Investigations into the in vitro antimicrobial activity and mode of action of the phenazine antibiotic D-alanylgriseoluteic acid. International journal of antimicrobial agents 29(1): pp. 93-97.
- Lim J A, Dong Hwan Lee, Byoung-Young Kim, Sunggi Heu (2014) Draft genome sequence of Pantoea agglomerans R190, a producer of antibiotics against phytopathogens and foodborne pathogens. Journal of biotechnology 188: pp.7-8.
- Manos J, R Belas (2006) The genera proteus, providencia, and morganella. Prokaryotes 6: 245-269.
- Liu D (2011) Molecular detection of human bacterial pathogens CRC press.
- Drzewiecka D (2016) Significance and roles of Proteus spp. bacteria in natural environments. Microbial ecology 72(4): 741-758.
- BohovaJ, Juraj Majtan, Viktor Majtan, Peter Takac (2014) Selective antibiofilm effects of Lucilia sericata larvae secretions/excretions against wound pathogens. Evidence-Based Complementary and Alternative Medicine.
- Barker H S, James W Snyder, Adam B Hicks, Stephen P Yanoviak, Paul Southern, et al. (2014) First case reports of Ignatzschineria (Schineria) indica associated with myiasis. Journal of Clinical Microbiology, 2014. 52(12): 4432-4434.
- Le Brun C, Martin Gombert, Sylvie Robert, Emmanuelle Mercier, Philippe Lanotte (2015) Association of necrotizing wounds colonized by maggots with Ignatzschineria–associated septicemia. Emerging Infectious Diseases 21(10): 1881-1883.
- Lee JK, Youn Yeop Lee, Kwan Ho Park, Jeonggu Sim, Youngcheol Choi, et al. (2014) Wohlfahrtiimonas larvae sp. nov., isolated from the larval gut of Hermetia illucens (Diptera: Stratiomyidae). Antonie Van Leeuwenhoek105(1): PP. 15-21.
- Čeřovský, V R Bém (2014) Lucifensins, the Insect Defensins of Biomedical Importance: The Story behind Maggot Therapy. Pharmaceuticals 7(3): 251-264.
- Klammsteiner T, Andreas Walter , Tajda Bogataj , Carina D Heussler , Blaž Stres, et al. (2021) Impact of processed food (canteen and oil wastes) on the development of black soldier fly (Hermetia illucens) larvae and their gut microbiome functions. Frontiers in microbiology 12: 619112.
- Cifuentes Y Stefanie P Glaeser, Jacques Movie, Jens-Ole Bartz, Ariane Müller, et al. (2020) The gut and feed residue microbiota changing during the rearing of Hermetia illucens Antonie Van Leeuwenhoek 113: 1323-1344.
- Li, XY, Cheng Mei, Xing-Yu Luo, Dilinuer Wulamu, Shuai Zhan, Yong-Ping Huang, et al. (2022) Dynamics of the intestinal bacterial community in black soldier fly larval guts and its influence on insect growth and development. Insect Science.
- Jiang CL, Wei-Zheng Jin, Xin-Hua Tao, Qian Zhang, Jun Zhu et al. (2019) Black soldier fly larvae (Hermetia illucens) strengthen the metabolic function of food waste biodegradation by gut microbiome. Microbial biotechnology 12(3): 528-543.
- Bruno D, Marco Bonelli , Francesca De Filippis , Ilaria Di Lelio , Gianluca Tettamanti, et al. (2019) The intestinal microbiota of Hermetia i llucens larvae is affected by diet and shows a diverse composition in the different midgut regions. Applied and environmental microbiology 85(2): e01864-18.
- Ao Y, Chongrui Yang, Shengchen Wang, Qingyi Hu, Li Yi, et al. (2021) Characteristics and nutrient function of intestinal bacterial communities in black soldier fly (Hermetia illucens) larvae in livestock manure conversion. Microbial biotechnology 14(3): 886-896.
- Rhishipal R, R Philip (1998) Selection of marine yeasts for the generation of single cell protein from prawn-shell waste. Bioresource technology 65(3): 255-256.
- Soares CA, Márcio Maury, Fernando C. Pagnocca, Fábio V. Araujo, et al. (1997) Ascomycetous yeasts from tropical intertidal dark mud of southeast Brazilian estuaries. The Journal of general and applied microbiology 43(5): 265-272.
- Limtong S, Wichien Yongmanitchai , Hiroko Kawasaki , Tatsuji Seki (2007) Candida thaimueangensis sp. nov., an anamorphic yeast species from estuarine water in a mangrove forest in Thailand. International Journal of Systematic and Evolutionary Microbiology 57(3): 650-653.
- Pfaller M A D Diekema (2007) Epidemiology of invasive candidiasis: a persistent public health problem. Clinical microbiology reviews 20(1): 133-163.
- Dang Xl, Jin-Huan Tian, Hui-Yu Yi, Wen-Xian Wang, Min Zheng, et al. (2006) Inducing and isolation of antibacterial peptides from oriental fruit fly, Bactrocera dorsalis Hendel. Insect Science 13(4): 257-262.
- Choi W, Ji-Hye Yun, Jong-Phil Chu, Ki-Back Chu, et al. (2012) Antibacterial effect of extracts of Hermetia illucens (Diptera: Stratiomyidae) larvae against Gram-negative bacteria. Entomol. Res 42(5): 219–226
- Park SI, BS Chang, SM Yoe, Detection of antimicrobial substances from larvae of the black soldier fly, H ermetia illucens (D iptera: S tratiomyidae). Entomological Research 44(2): 58-64.
- Jiang C L, Wei-Zheng Jin, Xin-Hua Tao, Qian Zhang, Jun Zhu et al (2019) Black soldier fly larvae (Hermetia illucens) strengthen the metabolic function of food waste biodegradation by gut microbiome. Microbial Biotechnology 12(3): 528-543.
- Varotto Boccazzi I, Matteo Ottoboni, Elena Martin, Francesco Comandatore, Lisa Vallone, et al. (2017) A survey of the mycobiota associated with larvae of the black soldier fly (Hermetia illucens) reared for feed production. PLoS one 12(8): e0182533.
- Tanga CM, Jacqueline Wahura Waweru, Yosef Hamba Tola, Abel Anyega Onyoni, Fathiya M Khamis, et al. (2021) Organic waste substrates induce important shifts in gut microbiota of black soldier fly (Hermetia illucens): coexistence of conserved, variable, and potential pathogenic microbes. Frontiers in Microbiology 12: 635881.
- Klammsteiner T, Andreas Walter, Tajda Bogataj , Carina D Heussler, Blaž Stres et al. (2020) The core gut microbiome of black soldier fly (Hermetia illucens) larvae raised on low-bioburden diets. Frontiers in Microbiology 11: 993.
- Greenwood MP, Kelvin L Hull, Marissa Brink-Hull, Melissa Lloyd, Clint Rhode (2021) Feed and host genetics drive microbiome diversity with resultant consequences for production traits in mass-reared black soldier fly (Hermetia illucens) larvae. Insects 12(12): 1082.






























