Bioinformatic Algorithm and Prediction to Control SARS-Cov-2 Spike Protein Variant on COVID-19 Pandemic Progression
Hyunjo Kim1* and Jae Hoon Song2
1School of Pharmacy, Yonsei University, South Korea
2Chairman of Asian-Pacific Research Foundation for Infectious and ANSORP, South Korea
Submission:February 05, 2021;Published: March 02, 2021
*Corresponding author: Hyunjo Kim, School of Pharmacy, Yonsei University, South Korea
How to cite this article: Hyunjo K, Jae Hoon S . Bioinformatic Algorithm and Prediction to Control SARS-Cov-2 Spike Protein Variant on COVID-19 0010 Pandemic Progression. Adv Biotech & Micro. 2021; 16(2): 555932.DOI:10.19080/AIBM.2021.15.555932
Abstract
Recently severe SARS-CoV-2 variant carrying the Spike protein amino acid change D614G has become the most prevalent form in the global COVID-19 pandemic. Systemic tracking algorithms of variant frequencies revealed a recurrent pattern of G614 increase at multiple geographical levels, the shift occurred even in narrow scope of local epidemics. The consistency of this pattern was highly statistically significant, suggesting that the G614 variant may have fitness advantage. Based on findings that the G614 variant grows to a higher titer as pseudo-typed virions. These findings illuminate changes important for mechanistic understanding of the virus and support continuing surveillance of Spike mutations to aid with development of immunological interventions. Furthermore, predictive stewardship would be conducted for protection and limitation of transmissibility.
Keywords: COVID-19; SARS-CoV-2 variant/mutant; Spike protein (S); D614G; Tracking algorithms; Predictive stewardship
Introduction
Coronavirus family has caused several human illnesses, the latest caused by SARS-CoV-2, has led to COVID-19 pandemic posing serious threat to global health [1-3]. A SARS-CoV-2 variant encoding a D614G mutation in the viral spike (S) protein has now become the most prevalent form of the virus worldwide, suggesting fitness advantage for the mutant [4-7]. The G614 variant is associated with higher upper respiratory tract viral load, higher infectivity, increased total S protein incorporation into the virion, reduced S1 shedding and a conformational change leading to a more ACE2- binding and fusion- competent state. However, it does not seem to be correlated to increased disease severity or escape neutralizing antibodies [8-12]. The SARS-CoV-2 D614G S protein variant supplanted the ancestral virus worldwide, reaching near fixation in a matter of months. Here it is shown that D614G was more infectious than the ancestral form on human lung cells, colon cells, and on cells rendered permissive by ectopic expression of human ACE2 or of ACE2 orthologs from various mammals. D614G did not alter S protein synthesis, processing, or incorporation into SARS-CoV-2 particles, but D614G affinity for ACE2 was reduced due to a faster dissociation rate [13, 14]. Consistent with this more open conformation, neutralization potency of antibodies targeting the S protein receptor-binding domain was not attenuated. Over the course of the SARS-CoV-2 pandemic, the identified SNPs were seen only once in the dataset, and only four SNPs rose to high frequency reported in GISAID [15,16]. This suggests that D614G confers replication advantage to SARSCoV-2 increases the likelihood of human-to-human transmission, which would support this hypothesis. Future prospective comparisons of D614G transmission to that of D614 seem unlikely given that D614G has gone to near fixation worldwide. However, the SARS-CoV-2 genomes that have been sequenced are only a narrow snapshot of the pandemic and additional sequencing of archived samples might pinpoint the origin of D614G or better resolve the variant’s trajectory. This D614G is associated with increased viral load in people with COVID-19 although these studies quantitated SARS-CoV-2 RNA and did not measure infectious virus.
When the SARS-CoV-2 S protein RBD is in its closed conformation, the binding site for ACE2 is physically blocked [17]. Models of coronavirus S-mediated membrane fusion describe ACE2 binding to all three RBD domains in the open conformation as destabilizing the pre-fusion S trimer, leading to dissociation of S1 from S2 and promoting transition to the post-fusion conformation. According to these models, the wellpopulated all- open conformation of D614G would reflect an intermediate that is on-pathway to S-mediated membrane fusion [18]. Therefore, structural biology provides key insights into 3D structures, critical residues/mutations in SARS-CoV-2 proteins, implicated in infectivity, molecular recognition and susceptibility to a broad range of host species [19]. The detailed understanding of viral proteins and their complexes with host receptors and candidate epitope/lead compounds is the key to developing a structure-guided therapeutic design. Since the discovery of SARS-CoV-2, several structures of its proteins have been determined experimentally at an unprecedented speed and deposited in the Protein Data Bank [20, 21]. Further, specialized structural bioinformatics tools and resources have been developed for theoretical models, data on protein dynamics from computer simulations, impact of variants/mutations and molecular therapeutics. Here, we provide an overview of ongoing efforts on developing structural bioinformatics algorithms and predictions for COVID-19 research. We also discuss the impact of these resources and structure-based studies, to understand various aspects of SARS-CoV-2 variant infection and therapeutic development. These include understanding differences between SARS-CoV-2 and SARSCoV-2 variant, leading to increased infectivity of SARS-CoV-2, and deciphering key residues in the SARS-CoV-2 involved in variant as well.
Methods
Coronavirus sequences and structures analysis
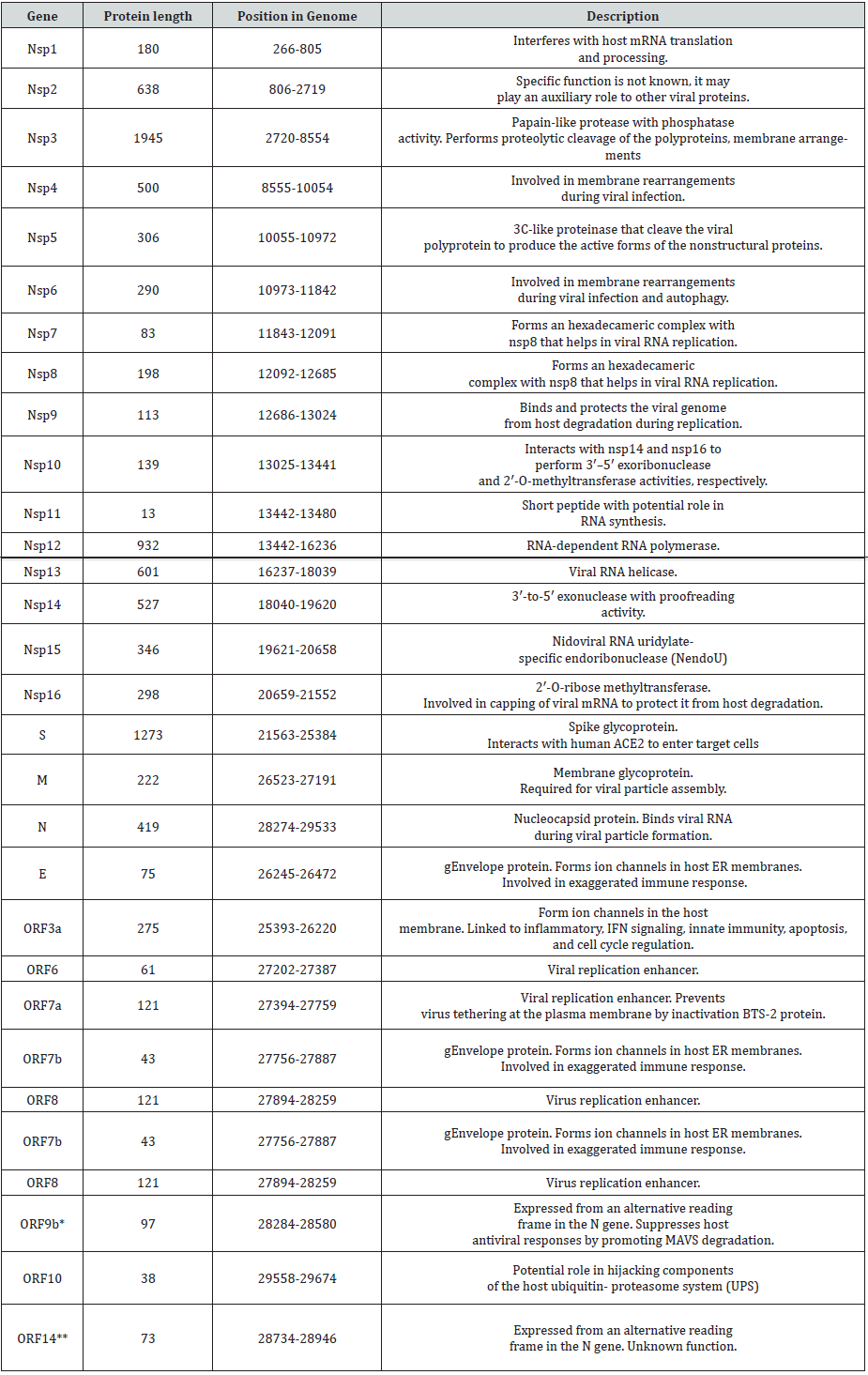
Amino acid sequences and mutant data of the S protein used in the analysis were obtained from NCBI GenBank and GISAID. After the first complete genome sequence of SARS-CoV-2 released on NCBI GenBank (accession number: NC 045512.2), there has been a large number of genome sequences as proceeding information is referred in Table 1. The mutant information of whole genome sequences of S protein with high coverage of SARS-CoV-2 strains from the infected individuals around the world was obtained from the GISAID database (https://www.gisaid.org). Sequence analysis and k-means clustering were described in detail elsewhere. For structural analyses, visualization, analysis and in silico mutations of protein structures were done (see Table 3). First, we downloaded the molecular structure of the spike protein from the Protein Data Bank (PDB). This structure corresponds to that resolved by colleagues and deposited in PDB.
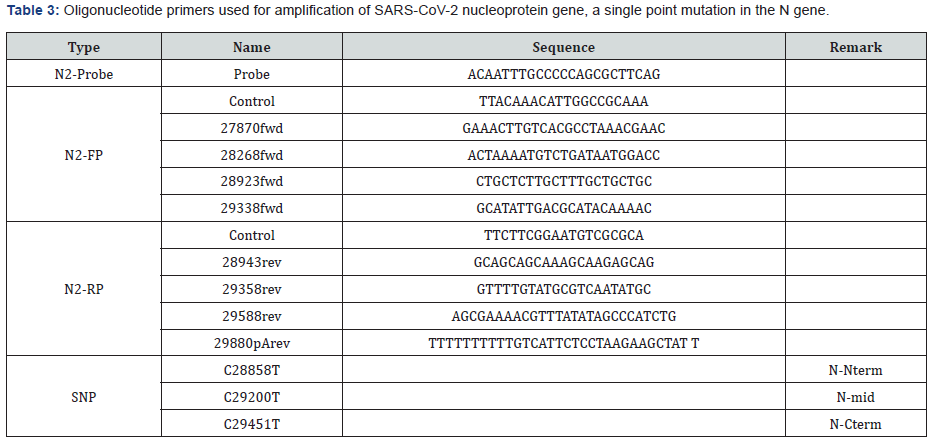
*Note: Primer names reflect the 5’ end of the respective oligonucleotide on the reference genome NC_045512.2.
Genetic Algorithm (GA) application
A population with defined fitness undergoes random mutation, crossover, and selection, and those with high fitness are retained. The individuals in the population are RNA sequences in our case and the fitness function is the number of residues that have the same 2D structure in both the target (provided for fitness calculation) and the predicted structure as determined by the Hamming distance, a metric for comparing two binary data strings. Based on prediction performance on the residue FSE (Frame shifting RNA Element) and computational complexity (see below result section), we choose NUPACK [22] for our GA [23,24]. The Nucleic Acid Package web server (http:// www.nupack.org) currently enables and GAs mimic evolution in nature. The initial population is obtained by randomly assigning nucleotides to the mutation region in the RNA sequence. This population is then subject to iterations of random mutation, crossover, selection, and nomination.
Nucleotide and amino acid variant detection
We first aligned each of these SARS-CoV-2 sequences using BLAT software [25, 26]. After the alignment, we extracted nucleotide sequences corresponding to individual proteins of SARS-CoV-2, translated them to amino acid sequences, and then compared them to reference amino acid sequences. Using the nucleotide mutations, the resulting amino acid mutations throughout the proteome of SARS‐CoV‐2 were determined. The amino acid changes were automatically annotated using the bioinformatics tool and a subsequent run, the resulting proteome from each SARS‐CoV‐2 genome was created and edited using CLC Genomics Workbench 20.0.3 [27]. This bioinformatics software is used by hundreds of microbiology and virology labs around the world for basic research and infectious disease epidemiology. The whole proteome was then aligned for phylogenetic analysis, and for identification of the resulting amino acid mutations.
Energy calculation
The effects of mutations on protein stability of S and binding affinity of RBD with hACE2 were estimated [28] by the folding energy change (G) and the binding energy change (︎ G) between the ︎ mutant structure (MUT) and Wild-Type (WT) structure, respectively [29]. FoldX was used for energy calculations [30,31]. The performance of the FoldX compares favorably with other random based approaches for protein engineering research including therapeutic antibody design. Particularly, FoldX is widely used for computational saturation mutagenesis in biomedical studies. All protein structures were repaired, and stability analysis was performed. The folding energy change was calculated using a Equation:
ΔΔG ︎stability︎ = ΔG ︎folding︎ MUT − ΔG ︎folding︎ WT ︎
A negative ΔΔG value suggests that the mutation can stabilize the protein and a positive value indicates that it makes the protein unstable. The structure-based tools DUET and CUPSAT were applied to check the reliability of FoldX for protein stability predictions [32, 33]. Additionally, interaction analysis carried out and the binding energy change was computed by a Equation [34]:
ΔΔΔG ︎binding︎ = ΔΔG ︎binding︎ MUT − ΔΔG ︎binding︎ WT
A negative ΔΔΔG value suggests that the mutation strengthens the binding affinity, whereas a positive value indicates that the mutation weakens the RBD–ACE2 interaction.
TopNetTree model for PPI BFE changes upon mutation
TopNetTree is a recently developed deep learning algorithm that integrates the advantages of convolutional neural networks and gradient-boosting trees [35]. The topology-based network tree (TopNetTree) was constructed by an innovative integration between the topological representation and NetTree for predicting PPI BFE changes following mutation ΔΔG [36,37]. In this work, TopNetTree is applied to predict the BFE changes of mutations that happened on the RBD with ACE2 of SARS-CoV-2. The topology-based feature generation is the first step followed by a convolutional neural network- assisted model as described in more details below result section. The topological representation uses element- and site-specific persistent homology to simplify the structural complexity of protein–protein complexes and encode vital biological information into topological invariants.
LSTM Network Based on NLP and Infection Rate
We propose a deep learning model based on Long Short-Term Memory (LSTM) for sentiment classification of COVID-19–related comments, which produces better results compared with several other well-known machine-learning methods [38,39]. Deep neural networks have the capacity to fit complex distributions but tend to overfit without sufficient supervision. As infection rate features are based on the growing percentage of each factor, they are stable across time. However, epidemic models based on the infection rate cannot predict policy changes and emergency conditions nor adjust the prediction with short-term influence. Therefore, we introduce the LSTM network based on Neuro- Linguistic programming (NLP) features to model the current policy and social media. Then, the short-term flexibility and longterm stability are both ensured.
Statistical Data Analysis
Fisher’s exact test was used to analyze the enrichment of epitopes and differences of mutation rates of SARS-CoV-2 isolated from different areas. Statistical analysis was carried out using the R statistical environment version 3.6.1. Mutation hotspots were identified as genome sites with two or more occurring mutations; on the other hand, mutation cold spots are those with no occurring mutations. The characterization of nucleotide mutations was done in terms of the nature of the nucleotide substitution (transition or transversion) and insertion and deletions (indel). The mutation densities (see a Equation) in the genome and proteome of SARSCoV‐ 2 were determined [40].
Mutation density = number of mutations ÷ size of genomic (nt length) or proteomic (aa length) region
Bioinformatic analysis: Data quality control and processing
Read quality control was carried out using FAST-QC and the default parameters [41]. Adapter sequences and low-quality bases were removed. Low-complexity reads, those with a length shorter than 40 bases, and duplicates were excluded using CDHIT- DUP v.4.6.8, where CD-HIT is a widely used program for clustering biological sequences to reduce sequence redundancy and improve the performance of other sequence analyses. Offtarget reads were then filtered out using Bowtie2 v2.3.4.3 with the default parameters against human genome version GRCh38.p13 (Genome Reference Consortium Human Build 38 patch release 13), and the SILVA (a ribosomal RNA) database as a reference to filter out human DNA and ribosomal sequences [42].
Results
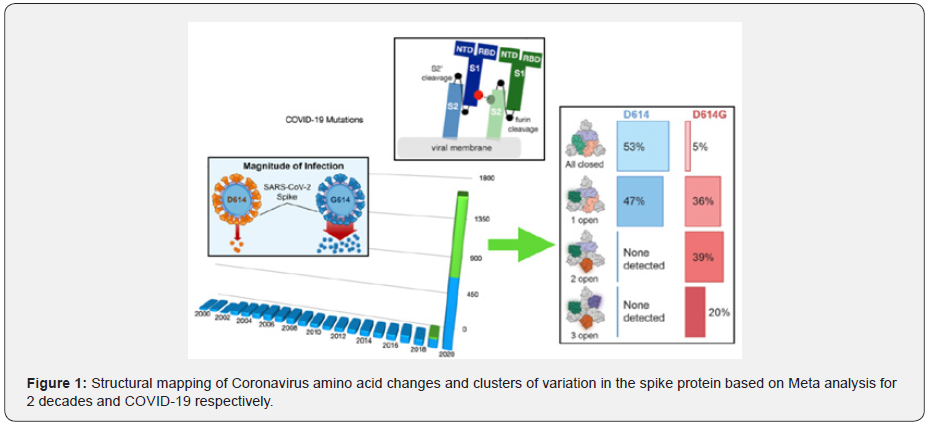
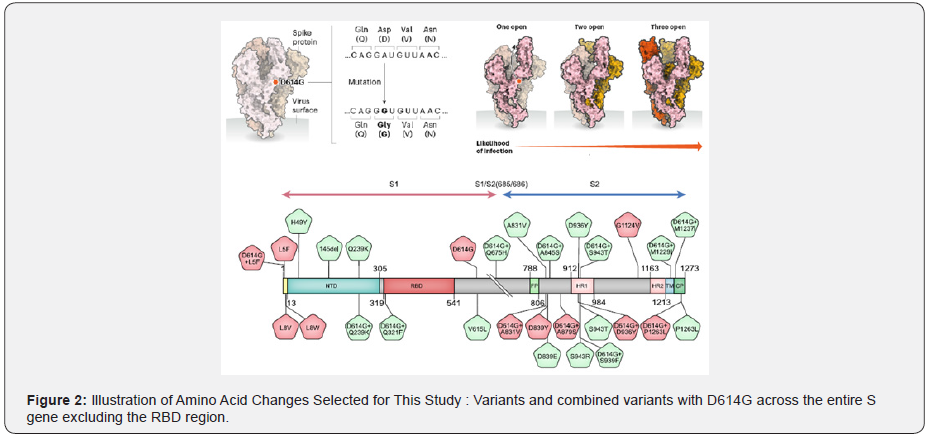
It is reported that increased COVID-19 pandemic severity has not been detected in association with D614G infection [15,43- 45]. Perhaps there are fitness tradeoffs for D614G in vivo due to the more open conformation of its RBD, which potentially renders D614G more immunogenic [44,46-48]. As shown in Figure 1, coronavirus mutants are prevalent since the end of 2019 and the location of D614G within the S protein is remote from the receptor-binding domain in keeping with the fact, that D614G affinity for ACE2 is less than that of D614, and that the relatively better-concealed D614 receptor-binding domain is likely to be advantageous for immune evasion, the D614G and D614 variants are equally sensitive to neutralization by human monoclonal antibodies targeting the S protein RBD. If SARS-CoV-2 Spike D614G is an adaptive variant that was selected for increased human-to-human transmission after spillover from an animal reservoir, one might expect that increased infectivity would only be evident on cells bearing ACE2 orthologs similar to that in humans. The increased infectivity of D614G was equally evident on cells bearing ACE2 orthologs from a range of mammalian species. Among these viruses, only SARS-CoV-2 possesses a polybasic furin cleavage site at the S1-S2 junction in the S protein, which is required for SARS- CoV-2 to infect human lung cells but not other cell types (see Figure 1). The mutation that loosens the spike protein is illustrated in Figure 2. Spike proteins on SARACoV- 2 bind to receptors on human cells helping the virus to enter. A spike protein is made up of three smaller peptides in open or closed orientations, when more are open. It’s easier for the protein to bind. The D614G mutation, the results of a single letter change to the viral RNA code, seems to relax connections between peptides. This makes open conformations more likely and might increase the chance of infection as shown in the top of Figure 2. Further, variants and combined variants with D614G across the entire S gene excluding the RBD region are more detailed in the bottom of Figure 2.
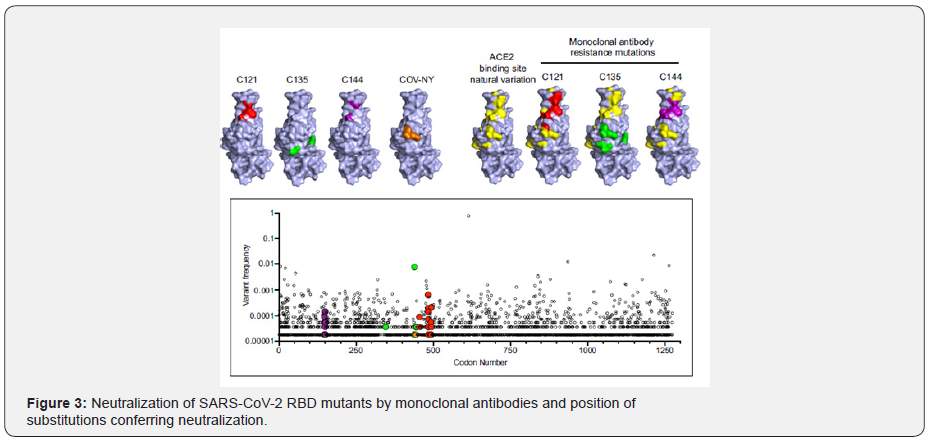
Presence of a novel mutation and a high frequency mutation in SARS‐CoV‐2
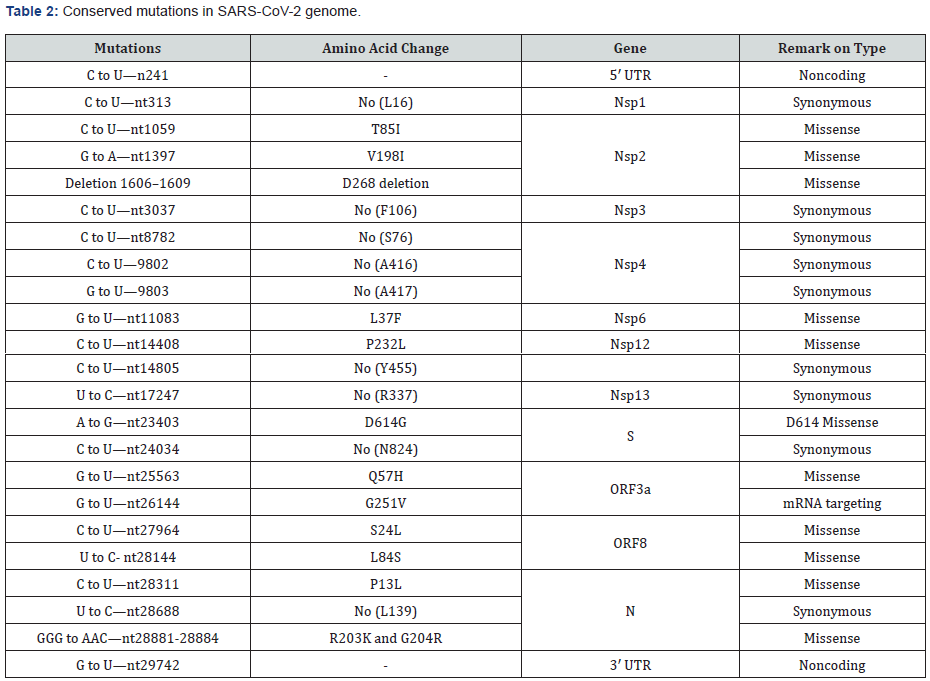
SARS-CoV-2 is a RNA coronavirus responsible for the COVID-19 pandemic of the Severe Acute Respiratory Syndrome. RNA viruses are characterized by a high mutation rate, up to a million times higher than that of their hosts. The virus mutagenic capability depends upon several factors, including the fidelity of viral enzymes that replicate nucleic acids, as SARS-CoV-2 RNA dependent RNA polymerase (RdRp). Mutation rate drives viral evolution and genome variability, thereby enabling viruses to escape host immunity and to develop drug resistance (see Figure 3). The morbidity of SARS-CoV-2 (COVID-19) is a serious public health concern globally and it is enigmatic how several antiviral and antibody treatments were not effective in the different period across the globe [49]. With the drastic increasing number of positive cases around the world WHO raised the importance in the assessment of the risk of spread and understanding genetic modifications that could have occurred in the SARS-CoV-2. Using all available deep sequencing data of complete genome from all over the world (NCBI repository). In the present work we have compared the SARS-CoV-2 reference genome to those exported from the GISAID database with the aim of gaining important insights into virus mutations, their occurrence over time and within different geographic areas. A total of SARS-CoV-2 sequences were collected from NCBI and GISAID databases and aligned against SARS-CoV-2 reference sequence NC_045512.2 as presented in Table1& 2. Additionally, Table 3 listed the used primmer for sequencing with the related descriptions.
Structural and function impact of mutations
Upon viral infection, the viral proteins express in the infected cells and are processed into small peptides by proteosomes [50]. These peptides are then presented by HLA molecules on the surface of the infected cells and recognized by T cells through their T cell receptors. Thus, the potential T cell epitopes can be derived from any of the viral structural and nonstructural proteins. Nucleotide conservation Shannon entropy is a measure of the amount of information (measure of uncertainty). Conservation of each of the four nucleotides has been determined using Shannon entropy. Note that it is assumed log (0) = 0 for smooth calculation of the SE. For a given sequence of length l, the conservation SE (converse) is calculated as follows using a Equation:
Conv_SE = - Sum i=1-4 pNilog (pNi)
where pNi = fi /i; fi represents the of represents the occurrence frequency of a nucleotide Ni in the given sequence [51].
Functional importance of the D614G mutation in the SARS-CoV-2 spike protein
Point mutations in SARS‐CoV‐2 variants
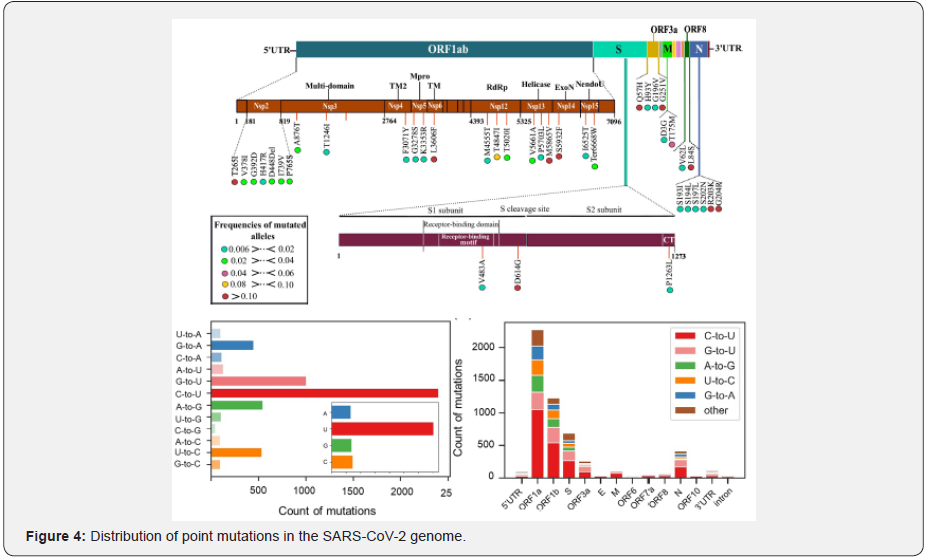
We performed phylogenetic network analysis using the sequences published in GISAID to investigate the frequency of point mutations in SARS- CoV-2 variants [52-55]. These sequences were collected and point mutation were calculated by the phylogenetic network analysis. We also analyzed the locations of these point mutations and observed a higher frequency of point mutations in several locations. In addition to it, we further counted the number of point mutations per gene in order to further analyze the polarization of point mutations in each gene and found more point mutations in ORF-1a and ORF-1b. However, as shown in Figure 4, Open Reading Frame (ORF)-1a and ORF-1b are much longer than other regions, which may result in more mutations; hence, we estimated the rate of point mutations per 100 bases in each gene ( see Figure 4). When normalized by gene length, the highest frequency of point mutations occurred in the 5′-untranslated Region (UTR) and 3′-UTR. These results indicate that point mutations are present in SARS- CoV-2 variants but they do not cluster within the gene coding regions.
The Impact on viral infectivity and antigenicity
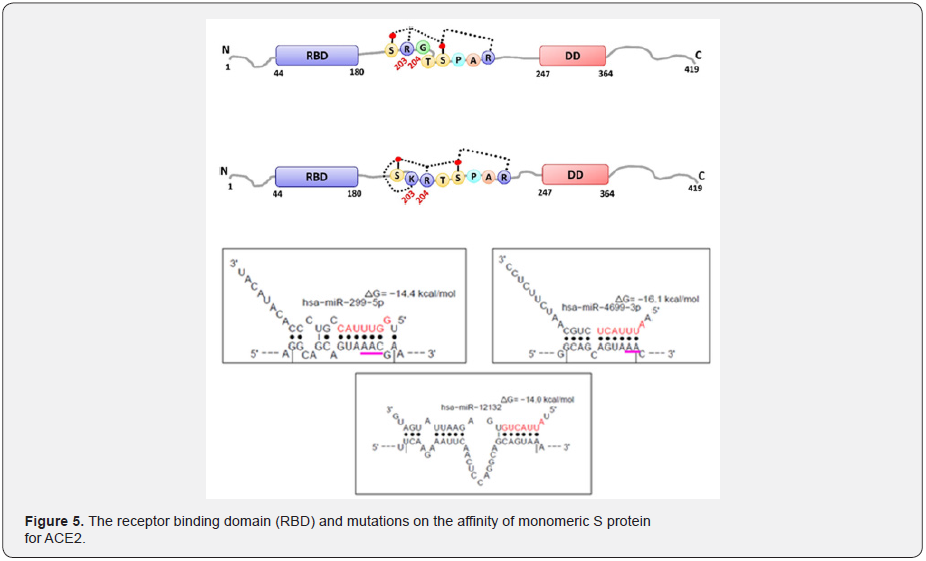
Severe acute respiratory syndrome coronavirus 2 (SARSCoV- 2) is an enveloped virus which binds its cellular receptor angiotensin-converting enzyme 2 (ACE2) and enters hosts cells through the action of its spike (S) glycoprotein displayed on the surface of the SARS-CoV-2. Compared to the reference strain of SARS- CoV-2, the majority of currently circulating isolates possess an S protein variant characterized by an aspartic acid-to-glycine substitution at amino acid position 614 (D614G). Residue 614 lies outside the receptor binding domain (RBD, see Figure 5) and the mutation does not alter the affinity of monomeric S protein for ACE2. However, S(G614), compared to S(D614), mediates more efficient ACE2-mediated transduction of cells by S-pseudotyped vectors and more efficient infection of cells by live SARSCoV- 2. This review article summarizes and synthesizes the epidemiological and functional observations of the D614G spike mutation, with focus on the biochemical and cell-biological impact of this mutation and its consequences for S protein function. We further discuss the significance of these recent findings in the context of the current global pandemic. The spike protein of SARSCoV- 2 has been undergoing mutations and is highly glycosylated. It is critically important to investigate the biological significance of these mutations. Here, we investigated 80 variants and 26 glycosylation site modifications for the infectivity and reactivity to a panel of neutralizing antibodies from convalescent patients [56-59]. D614G, along with several variants containing both D614G and another amino acid change were significantly more infectious. Most variants with amino acid change at receptor binding domain were less infectious, but some variants became resistant to some neutralizing antibodies. These findings could be of value in the development of incrementally modified vaccine and future therapeutic antibodies to quench COVID-19 pandemic eventually.
Predicting the affinity of ACE2 mutants to SARS-CoV-2 S1 protein using fast methods
We predicted the effect of the detected mutations in ACE2 on the affinity of ACE2 variants to S1 by using different computational methods in this section. There are many bioinformatic methods to predict the stability of the protein complex and the affinity between subunits by using various approaches as shown in Figure 6. The possible source of the variation between the prediction results of thermodynamic-based and other descriptor-based affinity predictors is the interface issue.
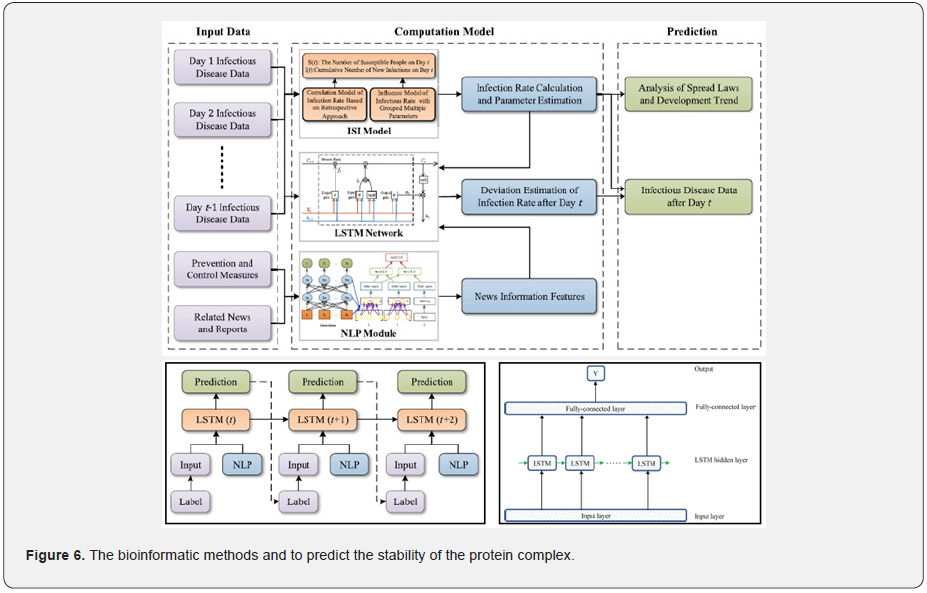
Long short‐term memory networks
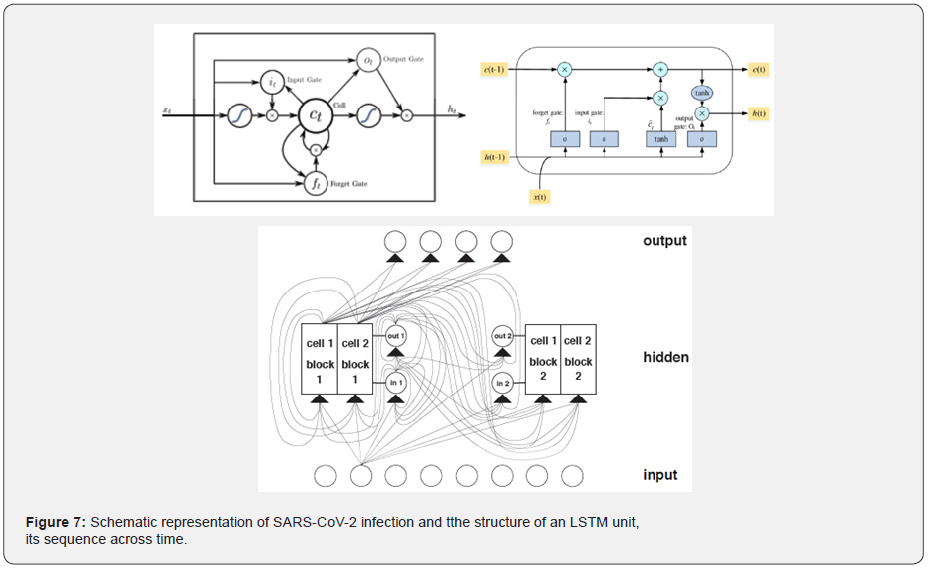
An LSTM network is a subclass of RNNs, trying to circumvent RNNs’ inability to learn to recognize long-term dependencies in the data sequences [38, 39, 60, 61]. It is addressed the latter by presenting the LSTM unit, whereas LSTM networks are constructed by combining several layers of LSTM units [60]. Specially, Figure 7 shows the structure of an LSTM unit and its sequence across time. Every and each single LSTM unit consists of three gates that operate on the input vector, xt, to generate the cell state, Ct, and the hidden state, ht. From a physical interpretation, the cell state can be viewed as the memory of the cell, while the gates control the flow of information in and out of the memory. In addition to it, the input gate determines the incorporation of new information, the forget gate determines which information should be discarded, and the output gate controls the information that passes along to the next layer. Following the interconnections presented in Figure 7, the following formulas per category of the variables hold:
ft =σ (Wfxt+Ufht-1+bf) (1) it =σ (Wixt+Uiht-1+bi) (2) ot =σ (Woxt+Uoht-1+bo) (3)
2. Candidate (memory) cell state variable:
C t =tanh(Wcxt+Ucht-1+bc) (4)
3. Cell and hidden state variables:
Ct =ft Ct-1+it Ct (5) ht = ot tanh(Ct) (6)
where {W, U} and b are the learnable weights and bias of the LSTM layer, respectively, for the input and the recurrent connections for the input/output/forget gates and cell state; ◦ is the elementwise product of two vectors; σ is a sigmoid function given by σ (x) = (1 + e−x )−1 to compute the gate activation function, whereas the hyperbolic tangent function (tanh) is used to compute the state activation function [62].
Potential ORF3a protein of SARS-CoV-2 possibility on viral immune-pathogenicity
Since none of the accessible SARS-CoV-2 genomes are streamlined in protein databases like STRING we could not directly get ORF3a (SARS-CoV-2)−human protein interactome. Our data have significantly established the structural resemblance of ORF3a protein between SARS-CoV and SARS-CoV-2 thus conceding further functional prediction. Thus, we deduce the functional pertinence of SARS-CoV-2 putative ORF3a protein from interactome and pathway enrichment analysis of SARS-CoV. The aforementioned neutralizing antibody escape mutations were artificially generated during in vitro replication of a recombinant virus. However, as monoclonal antibodies are developed for therapeutic and prophylactic applications, and vaccine candidates are deployed, and the possibility of SARS-CoV-2 reinfection becomes greater, it is important both to understand pathways of antibody resistance and to monitor the prevalence of resistance-conferring mutations in naturally circulating SARS-CoV-2 populations [63-65]. We used the GISAID and CoV-Glue, SARS-CoV-2 databases to survey the natural occurrence of mutations that might confer resistance to the monoclonal and plasma antibodies used in our experiments. Among the SARS-CoV-2 sequences in the CoV2-Glue database at the time of writing, different non-synonymous mutations were present in natural populations of SARS-CoV-2 S protein sequences.
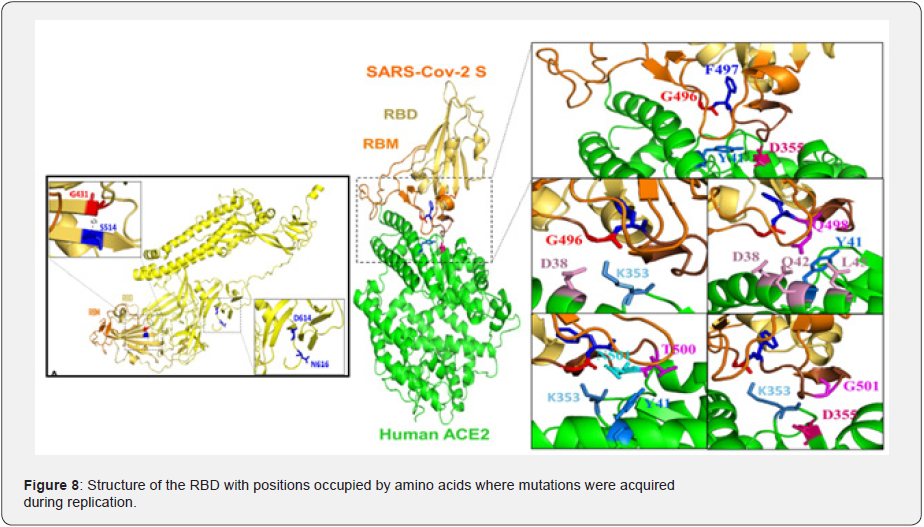
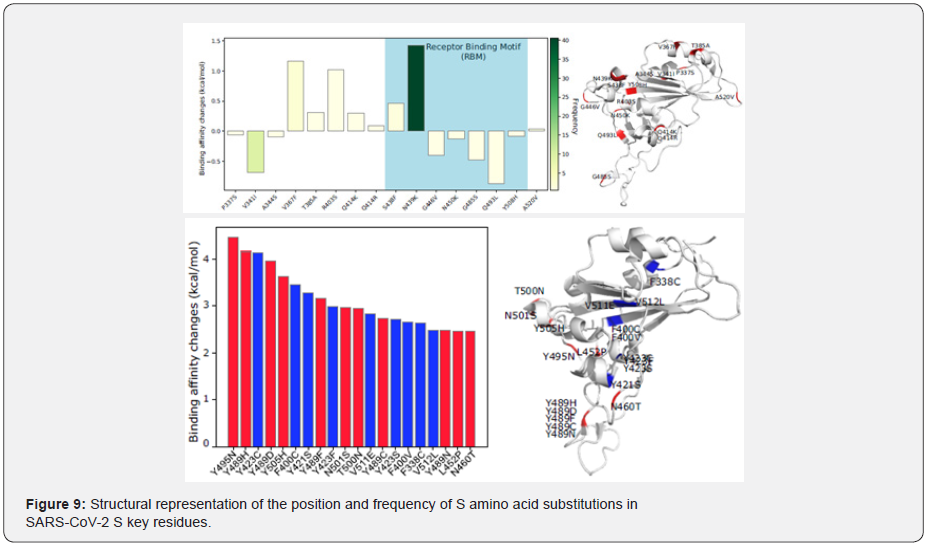
Consistent with the finding that none of the mutations that arose in our selection experiments gave an obvious fitness deficit, most were also present in natural viral populations. CoV-GLUE is an online web application for the interpretation and analysis of SARS- CoV-2 virus genome sequences, with a focus on amino acid sequence variation [66]. It is based on the GLUE data-centric bioinformatics environment and provides a browsable database of amino acid replacements and coding region indels that have been observed in sequences from the pandemic. Users may also analyse their own SARS-CoV-2 sequences by submitting them to the web application to receive an interactive report containing visualisations of phylogenetic classification and highlighting genomic variation of potentially high impact, for example linked to primer mismatches (see Table 3 as reference). Position of neutralization resistance-conferring substitutions. Structure of the RBD with positions that are occupied by amino acids where mutations were acquired during replication in the presence of each monoclonal antibody indicated (see Figure 8). Additionally, Figure 9 showed the position and frequency of S amino-acid substitutions in SARS-CoV-2 S. We identified and analysed the amino acid mutations that gained prominence worldwide from the early months of the pandemic. Eight mutations have been identified along the viral genome, mostly located in conserved segments of the structural proteins and showing low variability among coronavirus, which indicated that they might have a functional impact. At the moment of writing this paper, these mutations present a varied success in the SARS-CoV-2 virus population; ranging from a change in the spike protein that becomes absolutely prevalent.
Future selection of combinations of monoclonal antibodies for therapeutic and prophylactic applications
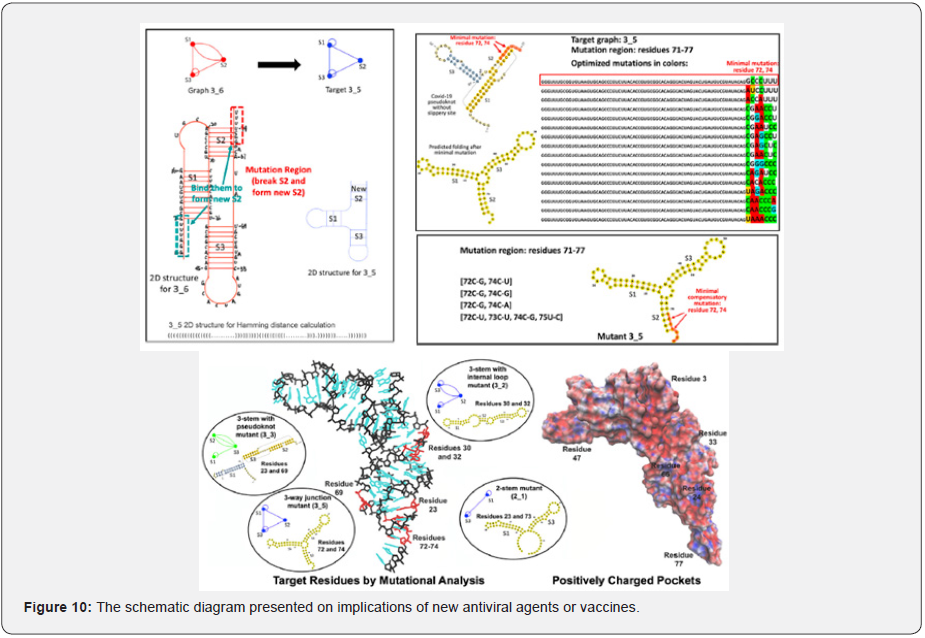
The ability of SARS-CoV-2 monoclonal antibodies and plasma to select variants that are apparently fit and that naturally occur at low frequencies in circulating viral populations suggests that therapeutic use of single antibodies might select for escape mutants. As shown in Figure 10, the schematic diagram presented on implications of new antiviral agents or vaccines. We tested whether combinations of monoclonal antibodies could suppress the emergence of resistant variants to mitigate against the emergence or selection of escape mutations during therapy, or during population-based prophylaxis [67].
Miscellaneous comment on recent spike protein changes
As seen on many occasions before, mutations are naturally expected for viruses and are most often simply neutral regional markers useful for contact tracing. The changes seen have rarely affected viral fitness and almost never affected clinical outcome, but the detailed effects of these mutations remain to be determined fully. Changes in the spike protein have relevance for potential effects on both host receptor as well as antibody binding with possible consequences for infectivity, transmission potential and antibody and vaccine escape. Actual effects need to be measured and verified experimentally and GISAID reports updates on spike mutations of recent submissions via gisaid. org/ spike and any sequence can be tested for spike mutations via gisaid.org/covsurver and from the internal analysis interface where individual countries/regions and time periods can be selected for custom analysis. This allows highlighting and tracking the rise of mutations like D614G or the currently most common receptor binding mutations as well as combinations of these mutations with deletions altering the spike protein surface [68,69]. As shown in Figure 11, it has become evident, these few S gene mutations and some deletions are found in multiple genomic contexts (different clades in different countries) that may be an early indication for some potential advantage for these viruses but needs to be verified and does not necessarily mean change in clinical severity or transmission efficiency.

Discussion
The severity of COVID-19 greatly varies from patient to patient. Majority of the patients either remain asymptomatic or develop mild to moderate symptoms. However, some COVID-19 patients who develop severe disease die even after hospitalization and intensive care. Why the disease severity differs so much from one person to another is one of the mysteries scientists are still trying to solve. The present study was designed to explore whether genetic variation in SARS-CoV-2 can explain variable severity of COVID-19. Mutation profiles of SARS-CoV-2 isolated from mildly affected and severely affected COVID-19 patients were explored and compared. Among numerous mutations observed in this study, two missense mutations, affecting RdRp and spike protein genes, respectively, were found most predominantly in the severely affected group compared with mildly affected group. Along with these two mutations in the 5′ UTR and a silent mutation in the ORF1ab were predominantly found in severely affected group the later not significantly [70,71] however, these mutations do not alter amino acid sequence in a protein. Many other mutations that were found in low frequency in the present study are unlikely to exert an effect on the severity of COVID-19. Therefore, the ability of spike protein and RdRp mutations on the severity of COVID-19 needs to be considered.
Conclusion
Ongoing efforts by the worldwide variants of SARS-CoV-2 are providing valuable insights into the structural mechanisms of action between bioinformatics algorithms and predictions. Structural biology can help explain the effect of amino acid variations on interactions with other proteins, leading to changes in the infection rate, associated symptoms and so on. This knowledge, when combined with structure-guided efforts to design stable vaccine antigens provides an important foundation for countering the impacts of the disease. These vaccines/drugs should target the regions of the protein which do not mutate fast. Therapeutics against regions with high mutation propensities will make them strain specific. Bioinformatics-guided approaches provide an important framework for understanding the increased virulence of this pathogen and for designing therapeutics and will be important for understanding the emergence of drug resistance and antibody-resistant variants/mutations of SARS-CoV-2.
Acknowledgment
We gratefully acknowledge the authors, originating and submitting laboratories of the sequences from GenBank and GISAID’s hCoV- 19 database on which this research is based.
References
- Rodrigues TS, de Sá KSG, Ishimoto AY, et al. (2021) Inflammasomes are activated in response to SARS-CoV-2 infection and are associated with COVID-19 severity in patients. J Exp Med 218(3): e20201707.
- Pan L, Wang R, Yu N (2021) Clinical characteristics of re-hospitalized COVID-19 patients with recurrent positive SARS-CoV-2 RNA: a retrospective study. Eur J Clin Microbiol Infect Dis.1-8.
- Baek MS, Cha MJ, Kim MC, Jin-Won Chung, Won-Young Kim, et al. (2021) Clinical and radiological findings of adult hospitalized patients with community-acquired pneumonia from SARS-CoV-2 and endemic human coronaviruses. PLoS One 16(1): e0245547.
- Choudhary S, Sreenivasulu K, Mitra P, Misra S, Sharma P (2021) Role of Genetic Variants and Gene Expression in the Susceptibility and Severity of COVID-19. Ann Lab Med 41(2): 129-138.
- Conti P, Caraffa A, Gallenga CES K Kritas, I Frydas, A Younes, et al., (2020) The British variant of the new coronavirus-19 (SarsCov-2) should not create a vaccine problem. J Biol Regul Homeost Agents. 35(1): 10.23812/21-3-E.
- Kim C, Ryu DK, Lee J (2021) A therapeutic neutralizing antibody targeting receptor binding domain of SARS-CoV-2 spike protein. Nat Commun 12(1): 288.
- Ahmadpour D, Ahmadpoor P, Rostaing L (2020) Impact of Circulating SARS-CoV-2 Mutant G614 on the COVID-19 Pandemic. Iran J Kidney Dis. 14(5): 331-334.
- Verkhivker GM, Di Paola L (2021) Dynamic Network Modeling of Allosteric Interactions and Communication Pathways in the SARS-CoV-2 Spike Trimer Mutants: Differential Modulation of Conformational Landscapes and Signal Transmission via Cascades of Regulatory Switches. J Phys Chem B. 125(3): 850-873.
- Mittal A, Verma V (2021) Connections between biomechanics and higher infectivity: a tale of the D614G mutation in the SARS-CoV-2 spike protein. Signal Transduct Target Ther 6(1): 11.
- Gobeil SM, Janowska K, McDowell S, Katayoun Mansouri, Robert Parks, et al. (2021) D614G Mutation Alters SARS-CoV-2 Spike Conformation and Enhances Protease Cleavage at the S1/S2 Junction. Cell Rep 34(2):108630.
- van Doremalen N, Purushotham J, Schulz J, et al. (2021) Intranasal ChAdOx1 nCoV-19/AZD1222 vaccination reduces shedding of SARS-CoV-2 D614G in rhesus macaques. Preprint bioRxiv.
- Klumpp-Thomas C, Kalish H, Hicks J, et al. (2020) D614G Spike Variant Does Not Alter IgG, IgM, or IgA Spike Seroassay Performance. J Infect Dis jiaa743.
- Lanjanian H, Moazzam-Jazi M, Hedayati M, Mahdi Akbarzadeh, Kamran Guity, et al. (2021) SARS-CoV-2 infection susceptibility influenced by ACE2 genetic polymorphisms: insights from Tehran Cardio-Metabolic Genetic Study. Sci Rep 11(1):1529.
- Rodriguez JH, Gupta A (2021) Contact residue contributions to interaction energies between SARSCoV-1 spike proteins and human ACE2 receptors. Sci Rep 11(1): 1156.
- To KK, Hung IF, Ip JD, (2020) COVID-19 re-infection by a phylogenetically distinct SARScoronavirus-2 strain confirmed by whole genome sequencing. Clin Infect Dis Ciaa 1275.
- Zhao Z, Sokhansanj BA, Malhotra C, Zheng K, Rosen GL (2020) Genetic grouping of SARS-CoV-2 coronavirus sequences using informative subtype markers for pandemic spread visualization. PLoS Comput Biol 16(9): e1008269.
- Chi X, Yan R, Zhang J, Guanying Zhang, Yuanyuan Zhang, et al. (2020) A neutralizing human antibody binds to the N-terminal domain of the Spike protein of SARS-CoV-2. Science 369(6504): 650-655.
- Xia S, Liu M, Wang C, et al. (2020) Inhibition of SARS-CoV-2 (previously 2019-nCoV) infection by a highly potent pan-coronavirus fusion inhibitor targeting its spike protein that harbors a high capacity to mediate membrane fusion Cell Res. 30(4): 343-355.
- Sedova M, Jaroszewski L, Alisoltani A, Godzik A (2020) Coronavirus3D: 3D structural visualization of COVID-19 genomic divergence. Bioinformatics. 36(15): 4360-4362.
- Wlodawer A, Dauter Z, Shabalin IG, Miroslaw Gilski, Dariusz Brzezinski, et al. (2020) Ligand-centered assessment of SARS-CoV-2 drug target models in the Protein Data Bank. FEBS J 287(17): 3703-3718.
- Burley SK, Bhikadiya C, Bi C, Chunxiao Bi , Sebastian Bittrich, et al. (2021) RCSB Protein Data Bank: powerful new tools for exploring 3D structures of biological macromolecules for basic and applied research and education in fundamental biology, biomedicine, biotechnology, bioengineering and energy sciences. Nucleic Acids Res 49(D1): D437-D451.
- Zadeh JN, Steenberg CD, Bois JS, et al. (2011) NUPACK: Analysis and design of nucleic acid systems. J Comput Chem 32(1): 170-173.
- Toubiana D, Puzis R, Sadka A, Blumwald E (2019) A Genetic Algorithm to Optimize Weighted Gene Co-Expression Network Analysis. J Comput Biol. 26(12): 1349-1366.
- Sale M, Sherer EA (2015) A genetic algorithm based global search strategy for population pharmacokinetic/pharmacodynamic model selection. Br J Clin Pharmacol 79(1): 28-39.
- Kent WJ (2002) BLAT--the BLAST-like alignment tool. Genome Res 12(4): 656-664.
- Bhagwat M, Young L, Robison RR (2012) Using BLAT to find sequence similarity in closely related genomes. Curr Protoc Bioinformatics. Chapter 10: Unit10.8.
- Liu CH, Di YP (2020) Analysis of RNA Sequencing Data Using CLC Genomics Workbench. Methods Mol Biol. 2102: 61-113.
- Olotu FA, Omolabi KF, Soliman MES (2020) Leaving no stone unturned: Allosteric targeting of SARS-CoV-2 spike protein at putative druggable sites disrupts human angiotensin-converting enzyme interactions at the receptor binding domain. Inform Med Unlocked. 21: 100451.
- Poosapati A, Gregory E, Borcherds WM, Chemes LB, Daughdrill GW (2018) Uncoupling the Folding and Binding of an Intrinsically Disordered Protein. J Mol Biol. 430(16): 2389-2402.
- Christensen NJ, Kepp KP (2012) Accurate stabilities of laccase mutants predicted with a modified FoldX protocol. J Chem Inf Model. 52(11): 3028-3042.
- Nadra AD, Serrano L, Alibés A (2011) DNA-binding specificity prediction with FoldX. Methods Enzymol.498: 3-18.
- Kumar R, Jayaraman M, Ramadas K, Chandrasekaran A (2020) Insight into the structural and functional analysis of the impact of missense mutation on cytochrome P450 oxidoreductase. J Mol Graph Model. 100: 107708.
- Parthiban V, Gromiha MM, Schomburg D (2006) CUPSAT: prediction of protein stability upon point mutations. Nucleic Acids Res. 34(Web Server issue): W239-W242.
- Gyulkhandanyan A, Rezaie AR, Roumenina L, Lagarde N, Fremeaux-Bacchi V, et al. (2020). Analysis of protein missense alterations by combining sequence- and structurebased methods. Mol Genet Genomic Med. 8(4): e1166.
- Wu K, Wei GW (2018) Quantitative Toxicity Prediction Using Topology Based Multitask Deep Neural Networks. J Chem Inf Model. 58(2): 520-531.
- Chen J, Wang R, Wang M, Wei GW (2020) Mutations Strengthened SARS-CoV-2 Infectivity. J Mol Biol. 432(19): 5212-5226.
- Cheng MH, Zhang S, Porritt RA, Arditi M, Bahar I (2020) An insertion unique to SARS-CoV-2 exhibits superantigenic character strengthened by recent mutations. Preprint bioRxiv.
- Hochreiter S, Schmidhuber J (1997) Long short-term memory. Neural Comput 9(8): 1735-1780.
- Liu X, Liu C, Huang R, Hao Zhu, Qi Liu, et al. (2020) Long short-term memory recurrent neural network for pharmacokinetic-pharmacodynamic modeling. Int J Clin Pharmacol Ther 59(2): 138-146.
- Maragatham G, Devi S (2019) LSTM Model for Prediction of Heart Failure in Big Data. J Med Syst 43(5):111.
- Eskier D, Suner A, Karakülah G, Oktay Y (2020) Mutation density changes in SARS-CoV-2 are related to the pandemic stage but to a lesser extent in the dominant strain with mutations in spike and RdRp PeerJ 8: e9703.
- Brown J, Pirrung M, McCue LA (2017) FQC Dashboard: integrates FastQC results into a web-based, interactive, and extensible FASTQ quality control tool. Bioinformatics. 33(19): 3137-3139.
- Chen H, Li Y, Sun W, Song L, Zuo R, Teng Y (2020) Characterization and source identification of antibiotic resistance genes in the sediments of an interconnected river-lake system. Environ Int 137: 105538.
- Hou YJ, Chiba S, Halfmann P, Camille Ehre, Makoto Kuroda, et al. (2020) SARS-CoV-2 D614G variant exhibits efficient replication ex vivo and transmission in vivo. Science 370(6523): 1464-1468.
- Plante JA, Liu Y, Liu J, Hongjie Xia, Bryan Johnson, et al. (2020) Spike mutation D614G alters SARS-CoV-2 fitness and neutralization susceptibility. bioRxiv.
- Zhang L, Jackson CB, Mou H, Amrita Ojha, Erumbi S Rangarajan, et al. (2020) The D614G mutation in the SARS-CoV-2 spike protein reduces S1 shedding and increases infectivity. Preprint bioRxiv.
- Korber B, Fischer WM, Gnanakaran S, HyejinYoon, JamesTheiler et al. (2020) Tracking Changes in SARS-CoV-2 Spike: Evidence that D614G Increases Infectivity of the COVID-19 Virus Cell 182(4): 812-827.e19.
- Grubaugh ND, Hanage WP, Rasmussen AL (2020) Making Sense of Mutation: What D614G Means for the COVID-19 Pandemic Remains Unclear. Cell 182(4): 794-795.
- Weissman D, Alameh MG, de Silva T, Thushan de Silva, Paul Collini, et al. (2021) D614G Spike Mutation Increases SARS CoV-2 Susceptibility to Neutralization. Cell Host Microbe 29(1): 23-31e4.
- Forni D, Cagliani R, Pontremoli C, Alessandra Mozzi, Uberto Pozzoli, et al. (2020) Antigenic variation of SARS-CoV-2 in response to immune pressure. Mol Ecol.
- Yurkovetskiy L, Wang X, Pascal KE, Christopher Tomkins-Tinch, Thomas P Nyalile, et al. (2020) Structural and Functional Analysis of the D614G SARS-CoV-2 Spike Protein Variant. Cell 183(3): 739-751.e8.
- Poran A, Harjanto D, Malloy M, Christina M Arieta, Daniel A Rothenberg, et al. (2020) Sequence-based prediction of SARS-CoV-2 vaccine targets using a mass spectrometry-based bioinformatics predictor identifies immunogenic T cell epitopes. Genome Med 12(1): 70.
- Kosuge M, Furusawa-Nishii E, Ito K, Saito Y, Ogasawara K (2020) Point mutation bias in SARSCoV-2 variants results in increased ability to stimulate inflammatory responses. Sci Rep 10(1): 17766.
- Pachetti M, Marini B, Benedetti F, et al. (2020) Emerging SARS-CoV-2 mutation hot spots include a novel RNA-dependent-RNA polymerase variant. J Transl Med 18(1):179.
- Lau SY, Wang P, Mok BW, Anna Jinxia Zhang, Hin Chu, et al. (2020) Attenuated SARS-CoV-2 variants with deletions at the S1/S2 junction. Emerg Microbes Infect 9(1): 837-842.
- Daniloski Z, Guo X, Sanjana NE (2020) The D614G mutation in SARS-CoV-2 Spike increases transduction of multiple human cell types. Preprint bioRxiv.
- Li Q, Wu J, Nie J, Li Zhang, Huan Hao, et al. (2020) The Impact of Mutations in SARS-CoV-2 Spike on Viral Infectivity and Antigenicity. Cell.182(5): 1284-1294.e9.
- Shajahan A, Supekar NT, Gleinich AS, Azadi P (2020) Deducing the N- and O-glycosylation profile of the spike protein of novel coronavirus SARS-CoV-2. Glycobiology 30(12): 981-988.
- Allen JD, Watanabe Y, Chawla H, Newby ML, Crispin M (2020) Subtle Influence of ACE2 Glycan Processing on SARS-CoV-2 Recognition J Mol Biol 433(4):166762.
- Pujić I, Perreault H (2021) Recent advancements in glycoproteomic studies: Glycopeptide enrichment and derivatization, characterization of glycosylation in SARS CoV2, and interacting glycoproteins. Mass Spectrom Rev.
- Hochreiter S, Schmidhuber J (1997) Long short-term memory. Neural Comput 9(8):1735-1780.
- Gers FA, Schmidhuber J (2000) Cummins F Learning to forget: continual prediction with LSTM. Neural Comput. 12(10): 2451-2471.
- Guo H, Sung Y (2020) Movement Estimation Using Soft Sensors Based on Bi-LSTM and Two-Layer LSTM for Human Motion Capture. Sensors (Basel) 20(6):1801.
- Oberemok VV, Laikova KV, Yurchenko KA, Fomochkina II, Kubyshkin AV (2020) SARS-CoV-2 will continue to circulate in the human population: an opinion from the point of view of the virushost relationship. Inflamm Res 69(7): 635-640.
- Weisblum Y, Schmidt F, Zhang F, Justin DaSilva, Daniel Poston, et al. (2020) Escape from neutralizing antibodies by SARS-CoV-2 spike protein variants. Elife 9: e61312.
- van Dorp L, Richard D, Tan CCS, Shaw LP, Acman M, Balloux F (2020) No evidence for increased transmissibility from recurrent mutations in SARS-CoV-2. Nat Commun 11(1): 5986.
- Singer J, Gifford R, Cotten M, Robertson D, (2020) CoV-GLUE: A Web Application for Tracking SARS-CoV-2 Genomic Variation. Preprints, 060225.
- Rilinger J, Kern WV, Duerschmied D, Alexander Supady, Christoph Bode, et al. (2020) A prospective, randomised, double blind placebocontrolled trial to evaluate the efficacy and safety of tocilizumab in patients with severe COVID-19 pneumonia (TOC-COVID): A structured summary of a study protocol for a randomised controlled trial Trials 21(1): 470.
- Ortega JT, Serrano ML, Pujol FH, Rangel HR (2020) Role of changes in SARS-CoV-2 spike protein in the interaction with the human ACE2 receptor: An in silico analysis EXCLI J. 19: 410-417.
- Hussain M, Jabeen N, Raza F, Sanya Shabbir, Ayesha A Baig, et al. (2020) Structural variations in human ACE2 may influence its binding with SARS-CoV-2 spike protein. J Med Virol. 92(9):1580-1586.
- Baldassarre A, Paolini A, Bruno SP, Felli C, Tozzi AE, et al. (2020) Potential use of noncoding RNAs and innovative therapeutic strategies to target the 5'UTR of SARS-CoV-2. Epigenomics 12(15): 1349-1361.






























