A Sensitive and Cost-Effective Method for Isolation of Staphylococcus aureus from Agricultural Soils Suitable for Antimicrobial Resistance Surveillance Programs
Omega Y Amoafo, Ravi Gooneratne and Stephen L W On*
Department of Wine, Food and Molecular Biosciences, Faculty of Agriculture, Lincoln University, New Zealand
Submission: March 26, 2020; Published: April 06, 2020
*Corresponding author:Stephen LW On, Department of Wine, Food and Molecular Biosciences Faculty of Agriculture, Lincoln University, P. O Box 85084, Lincoln 7647, Canterbury New Zealand
How to cite this article: Omega Y Amoafo, Ravi Gooneratne, Stephen L W On. A Sensitive and Cost-Effective Method for Isolation of Staphylococcus Aureus from Agricultural Soils Suitable for Antimicrobial Resistance Surveillance Programs. Adv Biotech & Micro. 2020; 15(4): 555918. DOI:10.19080/AIBM.2020.15.555918
Significance and Impact of Study
The proliferation of antimicrobial resistance among bacteria is a major public health concern. Staphylococcus aureus is widespread in animals, the environment and humans, in which infections can be fatal. Certain agricultural practices have been linked to the proliferation of antimicrobial resistance traits among S. aureus strains, yet existing isolation methods use selective antibiotics, making investigations of the origins of resistance problematic. We describe a sensitive, antibiotic-free and cost-effective procedure for S. aureus isolation from agricultural soils, suited for surveillance and/or evolutionary studies with an interest in elucidating the origin of antimicrobial resistance.
Abstract
The inclusion of antimicrobials in selective media introduces culture bias that may confound surveillance studies aimed at exploring the origins of Antimicrobial Resistance (AMR). We describe a cost-effective procedure to isolate Staphylococcus aureus, a pathogen susceptible to acquisition of AMR, from agricultural soil without the use of selective antibiotics. Environmental stress was modelled using heat and ultraviolet light and S. aureus recoveries assessed using seeded sterile soils with several combinations of enrichment and isolation media. Suitable combinations were then applied to native soils. Using Modified Vogel-Johnson enrichment Broth (MVJB) and Baird-Parker Agar (BPA), as few as 3 cfu ml-1 of stressed S. aureus ATCC 25923 cells seeded into 25 g of sterile soil were retrieved. This combination detected S. aureus in 100% of 40 dairy farm soil samples and enabled differentiation from non-target organisms, with all isolates selected confirmed by species-specific PCR. We believe this cost-effective method could assist long-term surveillance studies aimed at exploring the origin and evolution of antimicrobial resistance in S. aureus.
Keywords: Staphylococcus aureus; Antimicrobial resistance; Isolation; Detection; Surveillance
Introduction
Staphylococcus aureus is an important, toxigenic foodborne pathogen [1,2], naturally found in milk and other dairy products [3,4] and is a major cause of mastitis and other diseases in the dairy industry [5,6]. It is also the cause of serious, life-threatening diseases in humans including toxic shock syndrome, meningitis and septicaemia [2,7]. In the context of antibiotic treatment for all of the above conditions, the emergence of Antimicrobial Resistance (AMR) in S. aureus (and many other bacteria) has now become a major global health threat [8,9]. As a consequence, considerable attention is now given to the surveillance of many microbes to help elucidate their prevalence, significance and indeed evolution of AMR. There is a concern about agricultural practices that could promote the development of AMR in zoonotic pathogens [10,11]. Furthermore, many clinically relevant antibiotics have their origins as secondary metabolites of fungi, thus naturally present in soil environments [12,13]. Low-level exposure to antimicrobials is an established mechanism for bacterial AMR development, and the transfer of AMR traits via mobile genetic traits is well recognized [10,13]. It seems prudent to understand the origins of AMR, including the potential of transfer from donor strains to recipients in natural agricultural environments, in order to inform and improve agricultural practices.
The role of the environment as a reservoir for AMR, and more specifically resistant strains of S. aureus has received considerable attention, including suggestions for improved surveillance systems [14,15]. Some authors favor the use of selective media for this purpose [15]; however such approaches inevitably run the risk of being too specific, and unable to recover strains that may possess traits conferring resistance to antibiotics other than those being selected for [16]. Inevitably, recovery rates using selective agents are less sensitive than those which are not [17], and also hinder attempts to fully evaluate the evolution of AMR in the environment over time or under different selective pressures such as divergent (for example, organic compared to conventional) agricultural practices. However, isolation of S. aureus from microbiologically complex media is far from trivial [18]. The aim of this study was to develop and evaluate a sensitive, simple and cost-effective isolation procedure for S. aureus without the use of selective antibiotics, for potential use in long-term evolutionary and epidemiological studies of AMR trends in S. aureus in the environment.
Materials and Methods
Preparation of heat- and ultraviolet light (UV)- treated S. aureus inoculant
Staphylococcus aureus strain ATCC 25923 cells stored at -80 °C were streak plated onto Mueller-Hinton (MH) agar and incubated at 37 °C for 24 h. A single colony was propagated under the same conditions and a cell suspension of 0.110-0.130 OD600 (Bio Rad Smartspec300™) (equivalent to ca. 106 CFU ml-1 [data not shown]) was prepared in sterile phosphate buffered saline. One ml aliquots were heat stressed by incubating for five min in a 48 °C or 50 °C water bath, pipetted as droplets to form ~ 3mm diameter into a sterile petri dish and exposed for 1 min at 60 cm from UV light lamp at 400 mW m-2 in a Biological Safety Cabinet class II. Treated and untreated aliquots were serially diluted and plated on MH agar to determine the percentage of original cells that were viable after each temperature-time exposure.
Evaluation of enrichment broth and isolation media combinations
Four Shott bottles (500 ml) each containing 25 g of dairy farm soil were sterilized at 121 °C for 15 min by autoclaving. Cooled soil samples were inoculated with one ml of the stressed S. aureus ATCC 25923 inoculum and incubated at room temperature for 2 h to allow for permeation into the soil material. Subsequently, 225 ml of the test enrichment (Baird-Parker broth, Vogel-Johnson Broth, modified nutrient broth) or control (phosphate buffered saline) was added and incubated while shaking at 37 °C for 8 h. After incubation, a 1:10 serial dilution was made from each enrichment using 0.1% peptone diluent (Fort Richards, Auckland, NZ), and 100 μl from the 10-3 and 10-6 dilution from each of the soil slurries of the various broths were spread-plated onto Baird-Parker, Mannitol Salt (MS), Mueller-Hinton and VJ selective agar plates. Plates were examined and enumerated after 24 h incubation at 37 °C. Experiments were conducted in triplicate.
Evaluation of isolation protocols on native agricultural soil samples
Farm soil samples from each of four different locations in Canterbury, New Zealand were processed using the candidate enrichment broths (BPB, VJB, modified BPB, modified VJB) to retrieve S. aureus cells and plated on the candidate agar media (BPA, VJA). Soil samples were processed as described above, without the seeding step. Presumptive S. aureus isolates were characterized by morphology on the agar medium, Gram stain and species-specific PCR [19] (see below).
Modification of enrichment broths
Following initial experiments, candidate enrichment broths BPB and VJB were each modified by adjusting the total NaCl content to 7.5% w v-1, prior to autoclaving. The efficacy of these media were then evaluated on sterilized soil samples, to which 1 ml suspensions each of stressed S. aureus (as above) and Proteus spp. (soil isolate strain number), adjusted to an OD of 0.110-0.130 OD600 (BioRad Smartspec300™) were added and incubated as described above.
Evaluation of antimicrobial resistance in S. aureus soil isolates
Isolates of S. aureus from each of the 40 dairy soil samples recovered with the modified protocol were examined for their resistance to nalidixic acid (30μg), ampicillin (10 μg), tetracycline (30 μg), penicillin G (5IU) and kanamycin (30 μg ) using disc diffusion procedures described by the European Committee on Antimicrobial Susceptibility Testing [20], whose guidelines for the description of intermediate resistance among strains were also used.
Species Identification by PCR
Suspensions of 24 h old bacterial culture in 0.1% peptone water (Fort Richards, Auckland New Zealand) standardized to 0.5 -2.0 O.D600 (Bio-Rad Smartspec300™) were used as the template in PCR reactions. Cell suspensions (5 μl) were added to a reaction mixture containing 2 μl PCR buffer, 2 μl Q, 0.4 μl MgCl2, 0.2 μl forward and reverse primers described by Brakstad et al [19] (forward, 5’to3’- GCG ATT GAT GGT GAT ACG GTT, reverse: 5’to3’ – AGC CAA GCC TTG ACG AAC TAA AGC: Invitrogen™ Auckland, New Zealand), 0.2 μl dNTPs, 0.2 μl Taq polymerase, made up to 20 μl with de-ionized water. The PCR was run in a Multigene Thermocycler (Labnet International Inc., Edison, NJ, USA) based on the following protocol: Initial heating at 95 °C for 15 min, then 1 min at 94 °C, annealing for 30 sec at 55 OC, extension at 72 OC for one min and the reaction mix was held at 4 OC. The PCR product was analysed with a horizontal 3% (wt vol-1) agarose gel electrophoresis in TBE buffer (pH 8.3) as described by Brakstad et al. [19] using sib safe (Invitrogen™, Auckland, New Zealand). Visualisation was done with a Biorad Gel doc. Reference strain ATCC 25923 was used as a positive control.
Results and Discussion
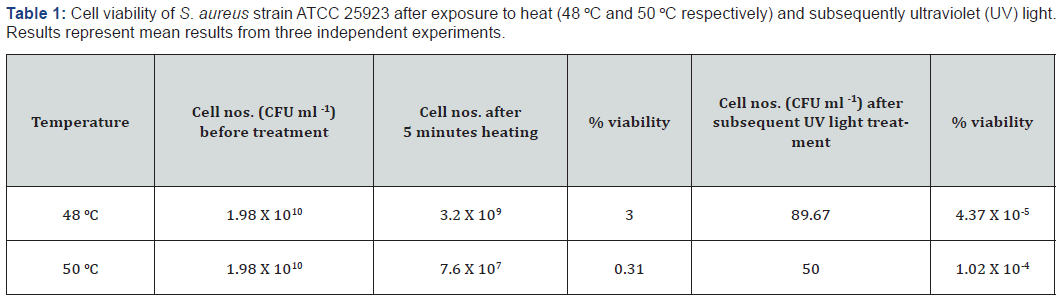
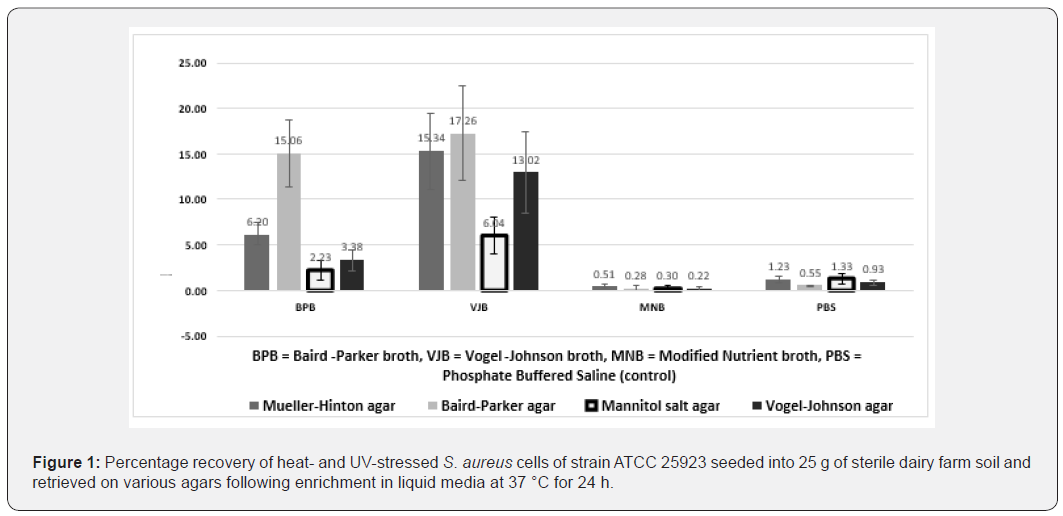
The stress of heat and UV light that S. aureus present in agricultural soils (also containing other organisms) are likely to experience were mimicked, as with other organisms in this environment [21,22]. Previous studies have used similar stress conditions to develop isolation media for S. aureus (reviewed by Baird and Lee [18]). Heat treatments reduced viable populations by up to 99% and UV treatments reduced the populations by a further 99% (Table 1), clearly indicating that inoculate produced in this manner had been subjected to significant environmental stresses. We hypothesised that an isolation protocol efficacious for such cells would also be effective for recovery of environmental S. aureus strains. We assessed combinations of previously described selective enrichment and solid isolation media that did not incorporate antibiotic agents, to enable application in AMR surveillance studies in which identifying the origins of resistance was important. Vogel-Johnson broth (VJB) yielded the highest and second highest mean percentage recovery rates of stressed S. aureus cells when used in combination with Baird-Parker (BPA) - and Mueller-Hinton agar respectively; Baird-Parker broth combined with Baird-Parker agar was also an effective recovery method (Figure 1). The high recovery rate and diagnostic features of the VJB-BPA combination (in which S. aureus colonies yield a characteristic halo on the agar, facilitated their identification [23], making this an ideal method for initial trials on native soil samples. However, our initial examination of 20 soil samples from four local dairy farms revealed that, although isolates positively identified as S. aureus were recovered, each of these samples also harbored bacteria that morphologically resembled S. aureus on BPA, yet yielded negative results in the species-specific PCR test used [19]. Microscopic examination demonstrated such isolates to be Gram-negative rods, of which a typical exemplar was identified as Proteus spp. by comparative analysis of the 16S rRNA gene using the BLAST algorithm (data not shown).

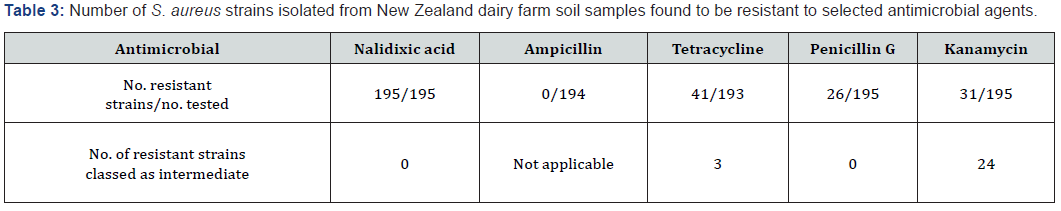
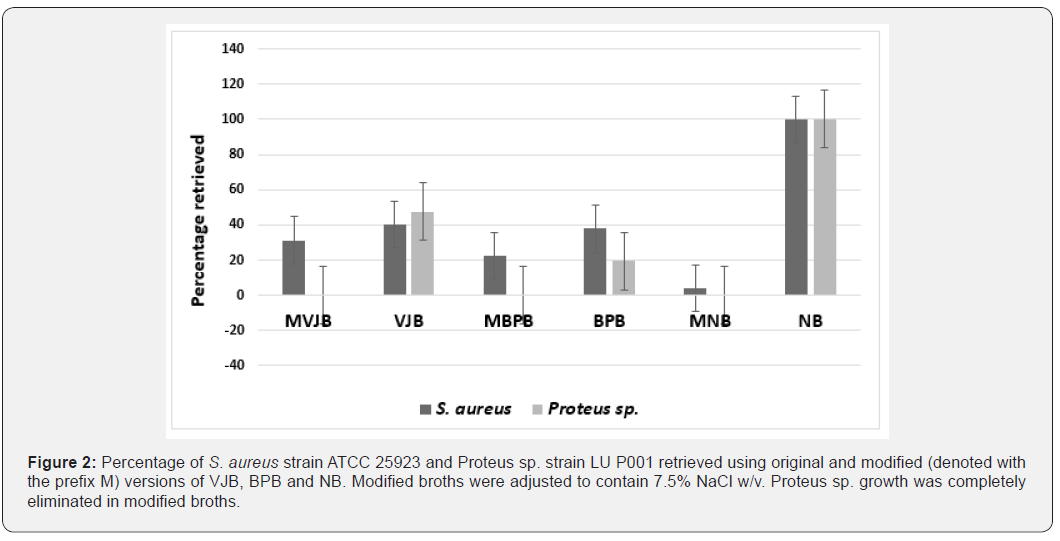
To improve selectivity without the use of antibiotics such as sulphamezathine [24], we adapted a candidate enrichment broth method by incorporation of 7.5% NaCl. Hypersaline conditions are hostile to many bacteria, yet S. aureus isolates can grow in media containing up to 25% NaCl w v-1 [25]. Although NaCl concentrations exceeding 4% w v-1 have been suggested as inhibiting the recovery of sub lethally damaged S. aureus cells [18], the inclusion of 7.5% NaCl w v-1 in selective media has been found to be effective in repressing a variety of Gram-negative bacteria in clinical samples [26]. This 7.5% NaCl concentration is lower than the 12.0% used to induce osmotic-stress induced cross-resistance to antibiotics [27], supporting its suitability in environmental surveillance studies of AMR in S. aureus. Supplementation of each of the enrichment broths with 7.5% NaCl (Figure 2) inhibited the growth of the Proteus strain recovered in our preliminary experiments. Experiments to determine the Limit of Detection (LOD) of NaCl-supplemented enrichment broths showed that the modified VJB was capable of recovering as few as 3 cfu ml-1 (Table 2). The use of the mVJB-BPA combination to examine 40 soil samples from four dairy farms (five paddocks from each farm, in each time period) sampled in September 2017 and September 2018 resulted in 100% of samples testing positive for S. aureus. In every positive sample, although a range of bacterial colonial morphologies were seen, only those with the characteristic black convex and round appearance yielding a halo of clearance of the otherwise opaque egg-yolk containing BPA medium were identified as S. aureus by PCR. Although our method does not exclusively select for S. aureus, the unique morphological features for each colony allow strains of interest to be readily identified for further study. Of the 195 S. aureus strains we recovered, all were found to be resistant to nalidixic acid, 72 were resistant to at least one other antimicrobial and 13 resistant to two others (Table 3), indicating the utility of the isolation method.


Conclusion
In summary, we have determined that a combination of mVJB enrichment broth followed by subculture onto BPA is an effective method for the isolation and presumptive identification of S. aureus from soil samples. Our approach accounts for environmental stresses the strains are likely to be exposed to in these environments, yet does not incorporate any antibiotics in the procedure, thus avoiding selective bias. Given the need for better understanding of the role of the environment in the promotion of antimicrobial resistance in S. aureus (an important human pathogen) [10,13,14,28], we hope wider adoption of this method will facilitate studies of the origin and evolution of AMR from farm to fork. The relatively simple and inexpensive nature of the procedure may also lend itself to applications in developing countries where there is currently a dearth of information on the true prevalence of AMR [8].
Acknowledgments
We would like to thank the farmers who allowed the collection of soil samples and provided information about their farm management systems. We thank Lincoln University for payment of the article processing fee.
References
- Haagsma JA, Polinder S, Stein CE, Havelaar AH (2013) Systematic review of foodborne burden of disease studies: Quality assessment of data and methodology. Int J Food Microbiol 166(1): 34-47.
- Castro A, Silva J, Teixeira P (2018) Staphylococcus aureus, a Food Pathogen: Virulence Factors and Antibiotic Resistance. In Foodborne Diseases ed. Holban, AM & Grumezescu AM Academic Press, Massachussetts, pp. 213-238.
- Viçosa GN, Moraes PM, Yamazi AK, Nero LA (2010) Enumeration of coagulase and thermonuclease-positive Staphylococcus spp. in raw milk and fresh soft cheese: an evaluation of Baird-Parker agar, rabbit plasma fibrinogen agar and the Petrifilm™ Staph express count system. Food Microbiol 27(4): 447-452.
- Papadopoulos P, Papadopoulos T, Angelidis AS, Boukouvala E, Zdragas A, et al. (2018) Prevalence of Staphylococcus aureus and of methicillin-resistant S. aureus (MRSA) along the production chain of dairy products in north-western Greece. Food Microbiol 69: 43-50.
- Azara E, Piras MG, Parisi A, Tola S (2017) Antimicrobial susceptibility and genotyping of Staphylococcus aureus isolates collected between 1986 and 2015 from ovine mastitis. Vet Microbiol 205: 53-56.
- Fursova KK, Shchannikova MP, Loskutova IV, Shepelyakovskaya AO, Laman AG, et al. (2018) Exotoxin diversity of Staphylococcus aureus isolated from milk of cows with subclinical mastitis in Central Russia. J Dairy Sci 101(5): 4325-4331.
- Aguilar J, Urday-Cornejo V, Donabedian S, Perri M, Tibbetts R, et al. (2010) Staphylococcus aureus meningitis: case series and literature review. Medicine (Baltimore) 89(2): 117-125.
- World Health Organization (2014) Antimicrobial resistance: global report on surveillance: World Health Organization. ISBN: 978 92 4 156474 8.
- MacGowan A, Macnaughton E (2017) Antibiotic resistance. Medicine (Baltimore) 7(4): 622-628.
- Aarestrup FM, Wegener HC, Collignon P (2008) Resistance in bacteria of the food chain: epidemiology and control strategies. Expert Rev Anti Infect Ther 6(5): 733-750.
- Aires-de-Sousa M (2017) Methicillin-resistant Staphylococcus aureus among animals: current overview. Clin Microbiol Infect 23(6): 373-380.
- Mohr KI (2016) History of Antibiotics Research. Curr Top Microbiol Immunol 398: 237-272.
- Perry J, Waglechner N, Wright G (2016) The Prehistory of Antibiotic Resistance. Cold Spring Harb Perspect Med 6(6): pii: a025197.
- Aarestrup FM (2015) The livestock reservoir for antimicrobial resistance: a personal view on changing patterns of risks, effects of interventions and the way forward. Philos Trans R Soc Lond B Biol Sci 370(1670): 20A140085.
- Huijbers PM, Blaak H, de Jong MC, Graat EA, Vandenbroucke-Grauls CM, et al. (2015) Role of the Environment in the Transmission of Antimicrobial Resistance to Humans: A Review. Environ Sci Technol 49(20): 11993-2004.
- Brown DF, Edwards DI, Hawkey PM, Morrison D, Ridgway GL, et al. (2005) Guidelines for the laboratory diagnosis and susceptibility testing of methicillin-resistant Staphylococcus aureus (MRSA). J Antimicrob Chemother 56(6): 1000-1018.
- Goodwin KD, Pobuda M (2009) Performance of CHROMagar Staph aureus and CHROMagar MRSA for detection of Staphylococcus aureus in seawater and beach sand--comparison of culture, agglutination, and molecular analyses. Water Res 43(19): 4802-4811.
- Baird R, Lee W (1995) Media used in the detection and enumeration of Staphylococcus aureus. Int J Food Microbiol 26(1): 15-24.
- Brakstad OG, Aasbakk K, Maeland JA (1992) Detection of Staphylococcus aureus by polymerase chain reaction amplification of the nuc gene. J Clin Microbiol 30(7): 1654-1660.
- European Committee on Antimicrobial Susceptibility Testing. (2019) Antimicrobial Susceptibility Testing. EUCAST disk diffusion method.
- Ebadi A, Khoshkholgh Sima NA, Olamaee M, Hashemi M, Ghorbani Nasrabadi R (2018) Remediation of saline soils contaminated with crude oil using the halophyte Salicornia persica in conjunction with hydrocarbon-degrading bacteria. J Environ Management 219: 260-268.
- Heděnec P, Singer D, Li J, Yao M, Lin Q, et al. (2018) Effect of dry-rewetting stress on response pattern of soil prokaryotic communities in alpine meadow soil. Appl Soil Ecol 126: 98-106.
- Baird Parker A (1962) An improved diagnostic and selective medium for isolating coagulase positive staphylococci. J Appl Bacteriol 25(1): 12-19.
- Smith B, Baird‐Parker A (1964) The use of sulphamezathine for inhibiting Proteus spp. on Baird‐Parker's isolation medium for Staphylococcus aureus. J Appl Microbiol 27(1): 78-82.
- Parfentjev IA, Catelli AR (1964) Tolerance of Staphylococcus aureus to sodium chloride. J Bacteriol 88: 1-3.
- Merlino J, Gill R, Robertson G (1996) Application of lipovitellin-salt-mannitol agar for screening, isolation, and presumptive identification of Staphylococcus aureus in a teaching hospital. J Clin Microbiol 34(12): 3012-3015.
- McMahon MAS, Xu J, Moore JE, Blair IS, McDowell DA (2007) Environmental stress and antibiotic resistance in food-related pathogens. Appl Environ Microbiol 73(1): 211-217.
- Woolhouse M, Ward M, van Bunnik B, Farrar J (2015) Antimicrobial resistance in humans, livestock and the wider environment. Philos Trans R Soc Lond B Biol Sci 370 (1670): 20140083.






























