Epigenetics and Epigenomics for Crop Improvement: Current Opinion
Suresh Kumar*
Division of Biochemistry, ICAR-Indian Agricultural Research Institute, India
Submission: May 13,2019; Published: May 28, 2019
*Corresponding author: Suresh Kumar, Division of Biochemistry, ICAR-Indian Agricultural Research Institute, USDA Norman E Borlaug Fellow, IUSSTF Fellow, INSA V Fellow, New Delhi-110012, India
How to cite this article: Suresh Kumar. Epigenetics and Epigenomics for Crop Improvement: Current Opinion. Adv Biotech & Micro. 2019; 14(1): 555879. DOI: 10.19080/AIBM.2019.14.555879
Abstract
Epigenetics refers to the studies on heritable changes in gene expression and cellular functions due to DNA methylation, histone modifications and biogenesis of non-coding RNAs without involving any alteration in the underlying DNA sequence. For the past two decades, extensive investigations have been made which provided a wealth of information on epigenetic regulation of gene expression both in animals and plants, and the effect it might have on plant growth, development and efficiency to cope with the environmental stresses. The rapid progress in high-throughput technologies has resulted in the unveiling of epigenetic landscapes at the genome level (the epigenome), deciphering the dynamics of methylome at a single-base resolution of plants including rice and tomato. Transgenerational nature of epigenetic variations can be a newer source of diversity, especially for the species showing low genetic diversity. The dynamic nature of epigenome, however, puts limitations on exploiting it without identifying the heritable epigenetic marks. Epigenome-Wide Association Studies (EWAS) using different genotypes/tissues/cell types/environmental conditions can provide valuable inputs for the development of epimarks for their possible use in crop improvement. The present opinion article provides a comprehensive view of the progress made in the field of epigenetics, and current opinion of the scientists to exploit them for crop improvement to meet the challenges of sustainable food security for the burgeoning global population.
Keywords: Crop improvement; DNA modification; Epigenetics; Epigenetic engineering; Epigenetic marks; Epigenetic variation; Gene regulation; Histone modification; Stress tolerance; Transgenerational inheritance
Introduction
Crop husbandry plays a key role in human society. We mainly depend on the domesticated plants like rice, wheat, maize, soybean, cotton, etc. for the supply of food, feed, fiber, and fuel. Ever since the domestication of plants, considerable progress has been made in agriculture because of the behavioral changes in human from food gathering to farming. While breeders aim at developing newer crop varieties with better tolerance to the environmental stresses, understanding the adaptation process to the changing climatic conditions is essential. Therefore, scientists have been interested in deciphering the underlying mechanisms that plants have evolved to adapt under diverse environment, particularly different types of biotic and abiotic stresses [1]. The concept of evolution is mainly based on the genetics and mutation deploying random changes in the DNA sequence that affect the phenotype and trait. The proposed models in evolutionary biology include the variation in the nucleotide sequence of DNA as a primary molecular mechanism behind the heritable phenotypic changes. Recent studies suggest that genetic variations might be sufficient for evolution, but genetic theory alone fails to explain some of the evolutionary features [2].
Explaining genotypic variations based on classical genetics and the rapid evolutionary changes under environmental pressure has become difficult. Additional mechanisms such as epigenetics can help to explain some of the enigmas [3]. If epigenetics is considered as a complementary molecular mechanism, many of the phenotypic variations, such as dissimilarity between the clones, can be easily explained. Plants being sessile in nature face multiple environmental stresses throughout their life [1]. Until the last century, it was thought that isolation of the gene(s) associated with a trait of interest was sufficient enough to transfer the trait to a plant and to achieve the expected phenotype [4,5]. However, the recent evidence suggests that the DNA provide only part of the genetic information for a trait while the chromatin structure also contributes to the expression of the trait.
DNA base modifications, post-translational modifications (acetylation, methylation, phosphorylation, etc.) of histone proteins, and regulatory RNAs (non-coding RNAs) outline distinct chromatin/epigenetic states of the genome (epigenome), which vary with the changing environmental conditions. Unlike the genome, which is largely invariable within an individual throughout its life, the epigenome is dynamically altered by the environmental factors. Since the epigenetic state of chromatin is variable, transfer of a trait from one species to another not only requires the transfer of the gene(s) associated with the trait but also the appropriate chromatin/epigenetic states so as to enable the trait to express [1]. It is, therefore, essential to study the epigenetic states in the donor species and to ensure proper re-establishment of the epigenetic state of the gene in the recipient species for the appropriate expression of the gene. However, epigenetic mechanisms of gene regulation are yet to be fully understood and utilized as epialleles (the alleles that are genetically identical but epigenetically different due to the epigenetic modifications, showing variable expression) in crop improvement programs.
An epigenome is defined as the sum total of all the changes in nuclear DNA, histone proteins and non-coding RNAs (ncRNAs) biogenesis in a cell [6]. Studies of the epigenetic changes in and around DNA that regulate genome activity have been defined as epigenetics. The area of epigenomics is broadening continuously with the identification of newer epigenetic/epitranscriptomic changes [6,7]. Chemical modifications of nitrogenous bases of DNA play a vital role in the regulation of gene expression [8]. Methylated cytosine (5-mC), also known as the fifth base of DNA, was recognized long before the DNA was identified as the genetic material. While more attention is given on the conventional modified DNA base (5-methylcytosine, 5-mC), the recent discovery of additional base modifications has resulted in an increased interest in epigenomics studies. With the identification of supplementary DNA base modifications [viz. 5-hydroxymethylcytosine (5-hmC) and N6-methyladenine (6- mA)], having known epigenetic regulatory functions at least in the animal system, the significance of epigenomic study has increased considerably. DNA base modifications have been detected in all the kingdoms of living organisms, including eukaryotes, prokaryotes, and viruses. Advanced studies in epigenetics, particularly in the area of cancer research, are being reported in the animal system, while the basic epigenomic study on the plant is still in the infancy and only little is known about the functional consequences of epigenetic/epigenomic changes in plants [9].
The newly discovered diversity in epigenetic modifications and the potential for their combinatorial interaction indicate that the epigenetic codons are considerably more complicated than it is considered today. Most of the proteins involved in DNA (de) methylation in Arabidopsis have been identified; however, the interaction between DNA methylation and other epigenetic or chromatin features remains unclear [10].
While relatively fewer modifications in DNA bases are known, more than 160 base modifications in RNAs have been identified so far. Among the modified nucleosides in DNA, 5-mC (also known as the 5th base of DNA) is a well-studied epigenetic mark [7,10]. Emerging evidence indicates that post-transcriptional modifications of nucleobases (e.g., N6-methyladenosine, 5-mC, 5-hmC, etc.) in RNA are promising players in the area of posttranscriptional regulation of gene expression. This is leading to the emergence of a newer branch of functional genomics known as epitranscriptomics [6].
Applications in crop improvement
The epigenetic phenotypes are being explained based on the fundamental discoveries such as activation, excision, and translocation of TEs, allelic interactions, transgene silencing and epialleles of the endogenous genes. Recent studies on epiRIL in Arabidopsis demonstrate that epigenetics of QTL can explain the heritability of the complex traits. Since epigenetic variations can affect important traits in crop plants, manipulation of stably inherited epigenetica variation could be a powerful tool in plant breeding [1]. It can enable modification of traits in the plant without altering the nucleotide sequence of the gene. Similarly, understanding the basis of phenotypic plasticity (the ability of a single genotype to express multiple phenotypes in response to environment) is crucial for crop breeding [11]. Reports on phenotypic plasticity have increased considerably in recent years, particularly on abiotic and biotic stresses. A growing body of evidence suggests that the processes of phenotypic plasticity involve epigenetic mechanisms. Therefore, epigenetic mechanisms, particularly DNA methylation, a generator of epialleles, would have important implications for plant breeders. Once it is established that an epigenetic mark is having causal effects on gene expression, its effect can be epigenetically manipulated with the help of targeted DNA/chromatin-modifying enzyme. A number of chromatin-modifying enzymes have been used to demonstrate successful manipulation of chromatin/ epigenetic marks at the target site. In a recent example, dCas9-p300 histone acetyltransferase fusion was used to activate transcription of MYOD and OCT4 genes from proximal promoters and distal enhancers [12]. Thus, chromatin modifiers can be used for up- or down-regulating gene transcription. However, it is important to note that effects on transcription are observed on the modification of some, but not all, targeted sites [13].
Zheng et al. [14] reported an accumulation of transgenerational epimutations in rice due to drought stress over 11 successive generations. They also reported that the genes of stressresponsive pathways showed accumulation of transgenerational epimutations, and the DNA methylation patterns in the droughtresponsive genes were affected by multi-generational drought. They reported 30% of the changes in methylation to be stable and heritable, which corroborated with the earlier findings of Wang et al. [15] who reported 29% of the drought-induced DNA methylation to be maintained even after recovery to the normal condition, and that of Kumar & Singh [16] who reported 25% of the increased methylation to be retained in a rice genotype IR-64- DTY1.1 even after removal of drought stress. Thus, epigenetics can be considered as an important regulatory mechanism in plant’s long-term adaptation and evolution under adverse environments. In Arabidopsis, DNA demethylases target promoter TEs to regulate stress-responsive genes [17]. The TEs (accounting for ~35% of the genome in rice) are usually suppressed by DNA methylation, and contribute to the activation of plant responses to abiotic stress [18]. Therefore, manipulating DNA methylation of TEs in the promoter region (by recruiting DRM2 to the target loci) could be considered for epigenetic manipulation of stress tolerance in plants [19,20].
Further evidence for the potential role of differentially methylated regions (DMRs) in influencing plant phenotype has been generated through the analysis of epigenetic recombinant inbred line (epiRIL) populations in Arabidopsis. The epiRILs are generated by crossing two genetically identical plants that differ in DNA methylation levels, owing to one parent being a homozygous mutant for a gene required for the proper maintenance of DNA methylation. The selection of offspring with the wild-type copy of this gene followed by multiple generations of self-pollination results in a population with very similar genomes that vary only in methylation levels. Due to the stripping of methylation at particular chromosomal regions of some of these epRILs, they have the potential to express information that is typically silenced by DNA methylation in natural variants.
Inheritance of epigenetic variation
One of the challenges ahead is to improve our understanding of the stability, reversibility, and heritability of epialleles. We require to devise strategies to ensure stable retention of the desirable epialleles within a breeding material, and to develop techniques for targeted epigenetic manipulation [21]. Many plants possess complex genome because of the repetitive elements and their polyploid nature, which makes gene interaction networks complicated and difficult to modulate. How epigenetic changes are superimposed on the multiple copies of a gene may provide a clue towards developing epigenetically engineered crop variety tailored to cope with multiple stresses. Stable inheritance of the adaptive epiallele is very important for increased fitness of the plant to changing climatic conditions. The question about the stability of epigenetic mark over the generations is crucial to decide the value of epigenomic information for crop improvement. If inheritance of DNA methylation is stable, then an epiallele would be faithfully inherited and new epialleles will rarely appear. However, if DNA methylation patterns are unstable then we can expect rapid formation/loss of epialleles within a population. Hagmann et al. [22] studied methylome of Arabidopsis grown in natural environments for about 100 years and found only a few differences, suggesting that methylomes can be stably transmitted for many generations even in a varying environment. However, Arabidopsis subjected to salt stress [23] and rice plants subjected to drought stress for several generations [14], showed the evidence for increased rates of changes in DNA methylation, suggesting that environmental stresses may alter methylation stability. Analysis of inheritance of DNA methylation in recombinant inbred populations of maize and soybean revealed stable inheritance with rare examples of unexpected patterns [24,25]. A careful analysis of epiRIL populations provides evidence for such inheritance. Epialleles that are created because of genetic variation, such as nearby transposon insertions, may be fairly stable. While the epialleles that are created because of spontaneous variation may be less stable [26].
Epigenome engineering for crop improvement
Epigenetic engineering provides a promising new way to achieve adaptive advantages without altering the underlying DNA sequence. Researchers have uncovered several epialleles that can provide major morphological/phenotypic variation. However, identification of epialleles that are linked to phenotypic variation is a slow process which often depends on spontaneous/natural epimutations. However, the studies on epiRILs of Arabidopsis have shown the ways by which epialleles can be generated and associated with phenotypic traits [26]. The ability to manipulate DNA methylomes and then to create individuals with a mixture of methylomes reveals that there are sources of allelic variation present in plant genomes. Accessing this allelic variation in plants other than Arabidopsis has been difficult because other plants are generally more sensitive to genome-wide alteration in DNA methylation [27,28]. Therefore, novel approaches that moderately disturb the methylomes of crop plant or the methods that precisely manipulate DNA methylation at the selected site will need to be developed to create novel epiallelic variation(s) in the plant.
Genetic engineering technology offers a new method of stress management having several advantages over conventional methods; however, having drawbacks like gene silencing due to the epigenetic changes. Plant-incorporated protectants (e.g. Bt gene) have been one of the modern crop biotechnology applications with several products in the world market [29,30]. In plants, transcriptional and post-transcriptional transgene silencing have been correlated with methylation of the gene. Generally, methylation of the promoter attached with the transgene correlates with the transcriptional gene silencing, while methylation of the coding region of the gene has been associated with post-transcriptional gene silencing. With the increasing knowledge of epigenetic transgene silencing, it is expected that it would be possible for us to solve this problem in the near future [1]. One of the strategies suggested to avoid transgene silencing has been the careful designing of the transgene and thorough analyses of the transformants at the molecular level.
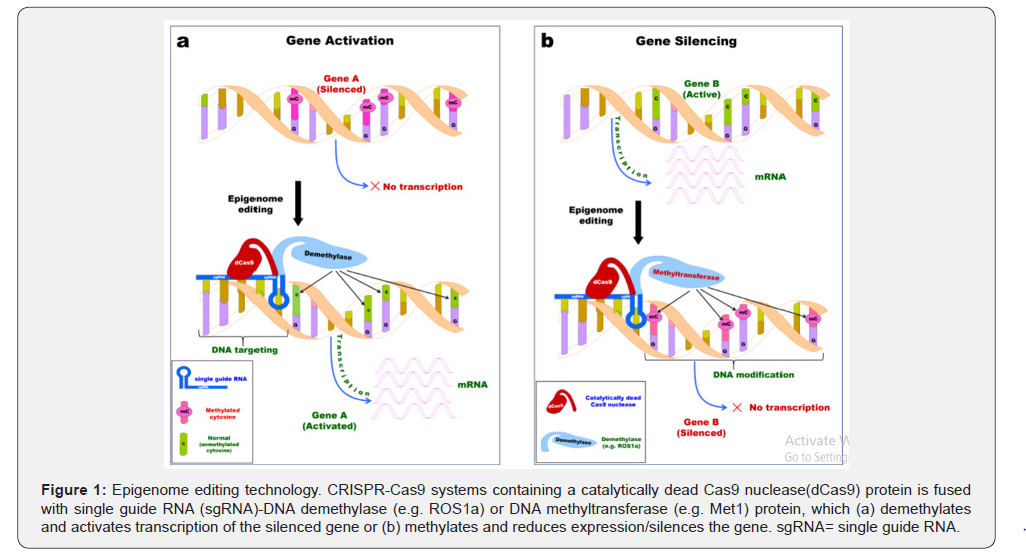
A promising method of epigenome editing for DNA methylation in a site-specific manner in plant genomes would require fusion of enzyme(s) that can add/remove DNA methylation with a guide (e.g. single guide-RNA) that will recruit the enzyme to the targeted DNA sequence (Figure 1). Nucleases like Zinc Finger Nucleases (ZFNs), Transcription Activator-Like Effector Nucleases (TALENs) and Clustered Regulatory Interspaced Small Palindromic Repeat (CRISPR)–Cas9 system are rapidly being developed to edit genome sequences.
CRISPR–Cas9 system uses two components to edit the target locus
a. a guide RNA (gRNA) and
b. a Cas nuclease (Cas9 being the most common).
The gRNA forms a complex with Cas9 and helps in the identification of the targeted genomic sequence followed by generation of a double-stranded break by the nuclease [6,31]. A modified version of it, namely CRISPR–dCas9, can be used in RNA-guided, dCas9-mediated (de)methylation of targeted loci in the plant genome for epigenetic engineering. Recently, a catalytically inactive SpdCas9 was fused with (de)methylases (SpdCas9-Tet1 or SpdCas9-Dnmt3a) to manipulate DNA (de) methylation in a site-specific manner in a mammalian cell line [19]. Subsequently, Liu et al. [32] used this technique using dCas9- Tet1-sgRNA to reverse hypermethylation at CGG expansion to convert the heterochromatin status of upstream FMR1 promoter into an active chromatin state, restoring a persistent expression of FMR1. The first demonstration of site-specific epigenome editing in plants was accomplished in Arabidopsis using a ZFN fused to SUVH9, a protein that is an integral part of the RdDM pathway [33]. Similarly, gluten content in wheat, for example, can be manipulated using epigenome editing technique by fusion of dCas9 with sgRNA-TaDME (a DNA demethylase) [34]. The genome-editing technologies are rapidly being improved, and it is expected that plant epigenome editing will be a reality in the near future (Figure 1).
In view of the biosafety concerns of genetically modified organisms currently being adopted for genetic manipulation of crop plants [4,35,36], the epigenetic engineering (which is supposed to have limited biosafety issues, if any) would be a preferred approach [6]. However, appropriate safety guidelines framed by the regulatory agencies of the country must be followed for personnel, laboratory and environmental safety [37,38]. Thus, epigenome engineering provides unprecedented opportunities for the manipulation of biological systems in an efficient manner to improve stress tolerance against the changing climatic conditions. This would allow functional integration of epigenetic marks and their usage towards stable improvement in the agriculturally important traits in the crop plants [13,26].
Future perspectives
Tremendous progress has been witnessed in recent years towards understanding the epigenetic regulation of gene expression in plants, particularly in Arabidopsis. The enzymes involved in DNA (de)methylation, histone modifications and roles of ncRNA in regulating developmental processes in plants are becoming more and more clear [34]. However, many areas of epigenetics are remaining to be explored. Does DNA (de) methylation interplay with other epigenetic or chromatin features, is not clear yet. Future epigenetic research should aim at identifying the developmental processes that involve epigenetic regulation. Evaluating the contribution of heritable epigenetic marks to the phenotypic variation has been a major challenge as many of the chromatin changes and variations in gene expression co-segregate with DNA sequence polymorphisms. Nevertheless, evidence suggests that plants possess heritable epiallelic variations that can be associated with a trait of interest and utilized for crop improvement. Although it is yet difficult to alter DNA methylation and chromatin states in a locus-specific manner, the situation is changing rapidly with the advances in genome editing tools. Thus, we can anticipate that soon epigenome editing will provide a means to assess the role of a QTL in epiallelic variations which might provide an interesting new route for the improvement of crop plants [1]. However, this will need a deeper understanding of the interactions between crop genomes, and the contribution of the epigenomic regulatory networks to the phenotypic plasticity.
Conclusion
Human being continues to improve understanding about the foods, their production and uses in maintaining and optimizing health. Domestication of plants followed by the selection based on desirable traits, breeding varieties for higher yield, tolerance to abiotic/biotic stress, better quality/nutrition, and the technological advancements have enabled a significant increase in food production [39]. The global population is expected to reach 9 billion by 2050. These 2-3 billion upsurges in the global population would require to increase food production by 70% [40]. Moreover, to feed the ever-growing population, we need to produce more food and feed from the day-by-day limiting arable lands, water, and other natural resources. Providing ample food to the global population is only the preliminary challenge; the major challenges would be to produce the food in a safe and sustainable manner [41,42] under the increasingly unfavorable environmental conditions [43]. Plants frequently face a variety of environmental stresses and cannot move away to avoid them; hence, they have developed strategies to produce a variety of protective compounds against the stressful conditions [44]. Although pants have the innate capability to survive under adverse climatic conditions, yet crop plants need improvement in their efficiency to produce more and more nutritious food even under unfavorable climatic conditions.
Properly harnessing the epigenetic variation is must to provide new opportunities for crop improvement and boost the production. Recent technological advances have provided new insights into the sources and inheritance of epigenetic variations. The coming years are likely to realize increased opportunities for monitoring and manipulating crop epigenomes. It would be critical to develop strategies to predict and stabilize epigenetic variations so that we can use epigenetics for stable improvement of agronomic traits. Because gene expression profile provides the primary account of the epigenotype to phenotype effect, it becomes essential to dissect the relative contributions of genetic and epigenetic variations on gene expression. The knowledge of epigenetic variation might allow exploitation of different epigenetic marks towards the development/selection of superior genotype at the early stage of plant growth. The views expressed herein are those of the author only. These may not necessarily be the views of the institution/organization the author is associated with.
References
- Kumar S, Singh AK, Mohapatra T (2017) Epigenetics: history, present status and future perspective. Indian J Genet Plant Breeding 77(4): 445-463.
- Ho WC, Zhang J (2014) The genotype-phenotype map of yeast complex traits: basic parameters and the role of natural selection. Mol Biol Evol 31(6): 1568-1580.
- Kumar S (2017) Epigenetic control of apomixis: a new perspective of an old enigma. Adv Plants Agric Res 7: 00243.
- Kumar S, Arul L, Talwar D, Raina SK (2006) PCR amplification of minimal gene expres-sion cassette: an alternative, low cost and easy approach to clean DNA transformation. Curr Sci 91: 930-934.
- Kumar S, Arul, L, Talwar D (2010) Generation of marker-free Bt transgenic indica rice and evaluation of its yellow stem borer resistance. J Appl Genet 51(3): 243-257.
- Kumar S (2018) Epigenomics of plant responses to environmental stress. Epigenomes 2(1) :6.
- Kumar S, Chinnusamy V, Mohapatra T (2018) Epigenetics of modified DNA bases: 5-methylcytosine and beyond. Front Genet 9: 640.
- Wang X, Li Q, Yuan W, Cao Z, Qi B, et al. (2016) The cytosolic Fe-S cluster assembly component MET18 is required for the full enzymatic activity of ROS1 in active DNA demethylation. Sci Rep 6: 26443.
- Kumar S (2018) Environmental stress, food safety, and global health: biochemical, genet-ic and epigenetic perspectives. Med Safety Global Health 7: e145.
- Singh A, Krishnan V, Vinutha T, Kumar S (2018) Biochemical, physiological, and mo-lecular approaches for improving salt tolerance in crop plants-a review. In: Goyal MR, Gupta SK, Singh A, (Eds) Engineering Practices for Management of Soil Salinity. Apple Academic Press pp. 159-208.
- Kooke R, Johannes F, Wardenaar R, Becker F, Etcheverry M, et al. (2015) Epigenetic basis of morphological variation and phenotypic plasticity in Arabidopsis thaliana. Plant Cell 27(2): 337-348.
- Hilton IB, D Ippolito AM, Vockley CM, Thakore PI, Crawford GE, et al. (2015) Epigenome editing by a CRISPR– Cas9 based acetyltransferase acti-vates genes from promoters and enhancers. Nat Biotechnol 33(5): 510-517.
- Stricker SH, Köferle A, Beck S (2017) From profiles to function in epigenomics. Nat Rev Genet 18(1): 51-66.
- Zheng X, Chen L, Xia H, Wei H, Lou Q, et al. (2017) Transgenerational epimutations induced by multi-generation drought imposition mediate rice plant’s adaptation to drought condition. Sci Rep 7: 39843.
- Wang WS, Pan YJ, Zhao XQ, Dwivedi D, Zhu LH, et al. (2011) Droughtinduced site-specific DNA methylation and its association with drought tolerance in rice (Oryza sativa L.). J Exp Bot 62(6): 1951-1960.
- Kumar S, Singh A (2016) Epigenetic regulation of abiotic stress tolerance in plants. Adv Plants Agric Res 5: 00179.
- Le TN, Schumann U, Smith NA, Tiwari S, Au PCK, et al. (2014) DNA demethylases target promoter transposable elements to positively regulate stress responsive genes in Arabidopsis. Genome Biol 15(9): 458.
- Jiao Y, Deng XW (2007) A Genome-wide transcriptional activity survey of rice transposable element-related genes. Genome Biol 8: R28.
- Liu XS, Wu H, Ji X, Stelzer Y, Wu X, et al. (2016) Editing DNA methylation in the mammalian genome. Cell 167(1): 233-247.
- Xu R, Qin R, Li H, Li D, Li L, et al. (2017) Generation of targeted mutant rice using a CRISPRCpf1 system. Plant Biotechnol J 15(6): 713-717.
- Kumar S (2018) Epigenetic memory of stress responses in plants. J Phytochemistry Bichem 2: e102.
- Hagmann J, Becker C, Müller J, Stegle O, Meyer RC, et al. (2015) Centuryscale methylome stability in a recently diverged Arabidopsis thaliana lineage. PLoS Genet 11(1): e1004920.
- Jiang C, Mithani A, Belfield EJ, Mott R, Hurst LD, et al. (2014) Environmentally responsive genome wide accumulation of de novo Arabidopsis thaliana mutations and epimutations. Genome Res 24(11): 1821-1829.
- Schmitz RJ, He Y, Valdés-López O, Khan SM, Joshi T, et al. (2013) Epigenome-wide inheritance of cytosine methylation variants in a recombinant inbred population. Genome Res 23(10): 1663-1674.
- Li Q, Eichten SR, Hermanson PJ, Springer NM (2014) Inheritance patterns and stability of DNA methylation variation in maize nearisogenic lines. Genetics 196(3): 667-676.
- Springer NM, Schmitz RJ (2017) Exploiting induced and natural epigenetic variation for crop improvement. Nat Rev Genet 18(9): 563- 575.
- Hu L, et al. (2014) Mutation of a major CG methylase in rice causes genome-wide hypo-methylation, dysregulated genome expression, and seedling lethality. Proc Natl Acad Sci 111: 10642-10647.
- Li Q, Eichten SR, Hermanson PJ, Zaunbrecher VM, Song J, et al. (2014) Genetic perturbation of the maize methylome. Plant Cell 26(12): 4602- 4616.
- Kumar S (2011) Biotechnological advancements in alfalfa improvement. J Appl Genet 52(2): 111-124.
- 30. Sharma A, Kumar S, Bhatnagar RK (2011) Bacillus thuringiensis protein Cry6B (BGSC ID 4D8) is toxic to larvae of Hypera postica. Curr Microbiol 62(2): 597-605.
- Xu X, Tao Y, Gao X, Zhang L, Li X, et al. (2016) CRISPR-based approach for targeted DNA demethylation. Cell Discov 2: 1600.
- Liu XS, Wu H, Krzisch M, Wu X, Graef J, et al. (2018) Rescue of fragile X syndrome neurons by DNA methylation editing of the FMR1 gene. Cell 172(5): 979-992.e6.
- Johnson LM, Du J, Hale CJ, Bischof S, Feng S, et al. (2014) SRA- and SETdomain- containing proteins link RNA polymerase V occupancy to DNA methylation. Nature 507 (7490): 124-128.
- Li Y, Kumar S, Qian W (2018) Active DNA demethylation: mechanism and role in plant development. Plant Cell Rep 37(1): 77-85.
- Kumar S, Chandra A, Pandey KC (2008) Bacillus thuringiensis (Bt) transgenic crop: An environment friendly insect-pest management strategy. J Environ Biol 29(5): 641-653.
- Kumar S (2014) Biosafety issues of genetically modified organisms. Biosafety 3: e150.
- Kumar S (2012) Biosafety issues in laboratory research. Biosafety 1: e116.
- Kumar S (2015) Biosafety and biosecurity issues in biotechnology research. Biosafety 4: e153.
- Kumar S, Krishnan V (2017) Phytochemistry and functional food: The needs of healthy life. J Phytochemistry Biochem 1: 103.
- Kumar S (2013) The role of biopesticides in sustainably feeding the nine billion global populations. J Biofertil Biopestici 4: e114.
- Kumar S (2012) Biopesticides: a need for food and environmental safety. J Biofertil Bopestici 3: e107.
- Kumar S (2015) Biopesticide: An environment friendly pest management strategy. J Bifertil Biopesti 6.
- Kumar S, Beena AS, Awana M, Singh A (2017) Salt-induced tissuespecific cytosine methylation downregulates expression of HKT genes in contrasting wheat (Triticum aesivum L.) genotypes. DNA Cell Biol 36: 283-294.
- Kumar S, Beena AS, Awana M, Singh A (2017) Physiological, biochemical, epigenetic and molecular analyses of wheat (Triticum aestivum) genotypes with contrasting salt toerance. Front Plant Sci 8: 1-20.






























