Actinobacteria: A Promising Source of Enzymes Involved in Lignocellulosic Biomass Conversion
Chadlia Hamdi1, Fatma Arous1 and Atef Jaouani1,2*
1Laboratory of Microorganisms and Active Biomolecules, Faculty of Sciences of Tunis, Institut Supérieur des Sciences, Tunisia
2Sciences Biologiques Appliquées de Tunis, University of Tunis El Manar, Tunisia
Submission: February 26, 2019; Published: May 09, 2019
*Corresponding author: Atef Jaouani, Laboratory of Microorganisms and Active Biomolecules, Faculty of Sciences of Tunis, Institut Supérieur des Sciences Biologiques Appliquées de Tunis, University of Tunis El Manar, 9, Rue Zouhair Essafi, 1007 Tunis, Tunisia
How to cite this article: Chadlia H, Fatma A, Atef J. Actinobacteria: A Promising Source of Enzymes Involved in Lignocellulosic Biomass Conversion. Adv Biotech & Micro. 2019; 13(5): 555874. DOI: 10.19080/AIBM.2019.13.5558584/a>
Abstract
Lignocellulosic materials are the most abundant and bio-renewable biomass on Earth, However, the breakdown of their major components (Cellulose, hemicellulose and lignin) requires the actions of different types of lignocellulolytic enzymes. Actually, several studies report that bacterial species belonging to Actinobacteria are able to produce a wide array of valuable enzymes implicated in biomass decomposition. In this mini-review, we attempt to provide an overview of the actinobacterial enzymes mainly involved in biomass conversion.
Keywords: Actinobaceria; lignocellulosic biomass; Enzymes; Conversion; Breakdown
Introduction
Lignocellulosic materials (plant cell walls) are the most abundant and bio-renewable biomass on Earth with more than 150 billons of tons produced annually [1,2]. Lignocellulose in nature derives from agricultural residues, forest-based woody materials and municipal and industrial solid wastes. However, the natural recalcitrance of these materials is the main obstacle for the separation of the major components [3]. Among the pretreatment methods used to break biomass recalcitrance, biological methods based on microbial enzymes, appear to be a promising technique without risks, low energy input and low capital costs [4]. Currently, actinobacerial enzymes are extensively studied due to their high efficiency in the bioconversion of biomass into interesting derived products [5,6]. In the same context, the present mini-review mainly contemplates on the actinobacterial enzymes involved in lignocellulosic biomass conversion.
Composition, structure and degradation of lignocellulosic biomass
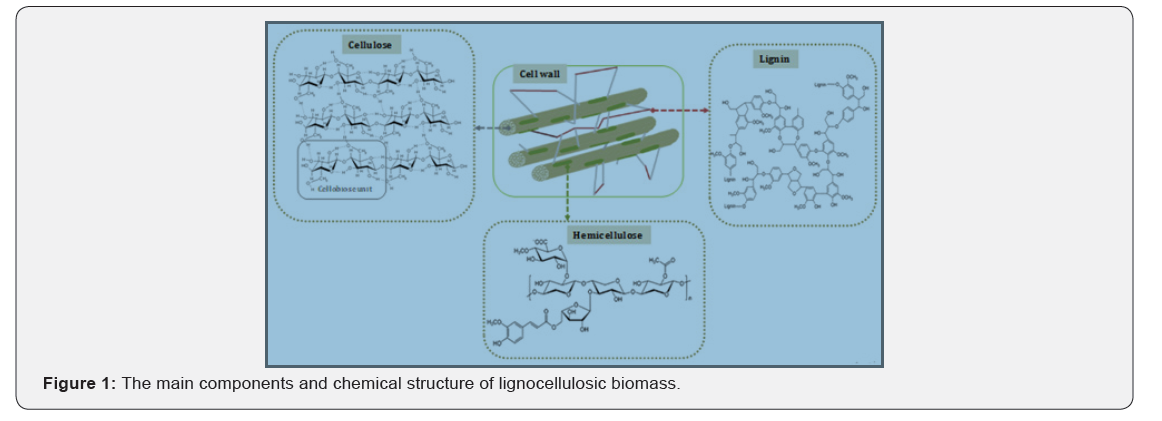
Lignocellulosic biomass is mainly composed of three major polymeric components namely cellulose (30-50%), hemicellulose (20-35%) and lignin (10-25%) that are strongly interlinked [1,7-9] (Figure 1). Cellulose is the major component of fibrous lignocellulosic material consisting of glucose and arabinose units linked by β-1,4 glycosidic bonds forming cellobiose molecules connected together by hydrogen bonds. The complete breakdown of cellulose requires the synergistic action of three types of cellulolytic enzymes (Endoglucanases, Exoglucanases and β-glucosidases) [6]. Hemicellulose is the second constituent of lignocellulose composed of hexoses (mannose), pentoses (xylose) and sugars acids. The complete degradation of Xylan the most important carbohydrate component found in hemicelluloses requires the cooperative action of a variety of hydrolytic enzymes such as endo-1,4-b-xylanase (endo-xylanases, exo-xylanases), xylan 1,4-b-xylosidase and xylan esterases [10]. Lignin is the last major part of lignocelluloses, basically composed of phenylpropane units linked to each other by the irregular coupling of C-C and C-O. Lignin degradation is mediated by three principal enzymes namely laccases, manganese peroxidases, and lignin peroxidases [11].
Actinobacteria: a source of lignocellulolytic enzymes
Actinobacteria are Gram-positive bacteria with high G+C content. They are ubiquitous in nature, found in different ecological niches [12]. The abundance, the diversity and the ability of Actinobacteria to colonize different ecological niches, have drawn considerable attention of the scientists to search novel enzymes with broader range of tolerance to environmental conditions [13]. Various studies have evaluated the lignocellulolytic ability of Actinobacteria (Table1). In the case of cellulose, the genera Streptomyces, Cellulomonas and Acidothermus were reported as promising source of endocellulases [14-16], whereas exocellulases were found in Thermobifida, Cellulomonas and Cellulosimicrobium genera [17]. On the other hand, species belonging to the genera Sreptomyces, Clavibacter, Terrabacter, Micrococcus and Microbacterium were widely studied for the production of β-Glucosidases [18,19]. Enzymes as xylanases involved in biotransformation of hemicellulose were reported in multiple genera, including Streptomyces, Cellulomonas, Cellulosimicrobium and Kocuria [10,20]. Lignin-degrading enzymes were mainly studied for fungi, but recent reports showed that Actinobacteria are able to breakdown lignin. Indeed, 10 laccase enzymes with industrial importance were reported in Streptomyces genus [21]. Other studies also reported peroxidase secretion by Nocardia, Rhodococcus, Streptomyces and Nonomuraea [22-24]. These studies represent recent examples research attempts interested to potential utility of Actinobacterial enzymes in breakdown of lignocellulosic biomass as well as in different industrial applications.
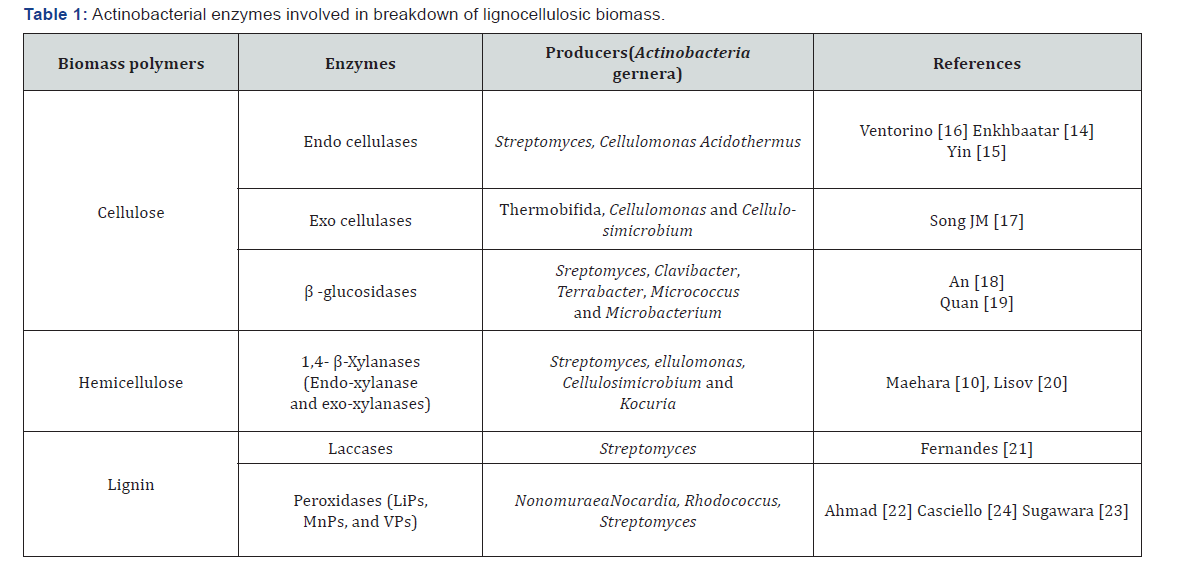
LiPs: lignin peroxidases; MnPs: Manganese peroxidases; VPs: Versatile peroxidases.
Conclusion
It is interesting to note that the screening of Actinobacteria with high ability for producing a wide range of lignocellulolytic enzymes, support their efficient contribution to biomass conversion. This may contribute to promote various biotechnological applications.
References
- Pauly M, Keegstra K (2008) Cell wall carbohydrates and their modification as a resource for biofuels. Plant J 54(4): 559-568.
- Zhang Y, Huang M, Su J, Hu H, Yang M, et al. (2019) Overcoming biomass recalcitrance by synergistic pretreatment of mechanical activation and metal salt for enhancing enzymatic conversion of lignocelluloses. Biotechnol Biofuels 12:12.
- Bomble YJ, Lin CY, Amore A, Wei H, Holwerda EK, et al. (2017) Lignocellulose deconstruction in the biosphere. Curr Opin Chem Biol 41: 61-70.
- Zhuo S, Yan X, Liu D, Si M, Zhang K, et al. (2018) Use of bacteria for improving the lignocellulose biorefnery process: importance of preerosion. Biotechnol Biofuels 11:146.
- Saini A, Aggarwal NK, Sharma A, Yadav A (2015) Actinomycetes: A Source of Lignocellulolytic Enzymes. Enzyme Research.
- Vetrovsky T, Steffen KT, Baldrian P (2014) Potential of Cometabolic Transformation of Polysaccharides and Lignin in Lignocellulose by Soil Actinobacteria. PLoS ONE 9(2): e89108.
- Himmel ME, Ding SY, Johnson DK, Adney WS, Nimlos MR, et al. (2007) Biomass recalcitrance: engineering plants and enzymes for biofuels production. Science 315 (5813): 804-807.
- Metzger JO, Huttermann A (2009) Sustainable global energy supply based on lignocellulosic biomass from afforestation of degraded areas. Naturwissenschaften 96 (2): 279-288.
- Mood SH, Golfeshan AH, Tabatabaei M, Jouzani G S, Najafi GH, et al. (2013) Lignocellulosic biomass to bioethanol, a comprehensive review with a focus on pretreatment. Renew Sust Energ Rev 27: 77-93.
- Maehara T, Yagi H, Sato T, Ohnishi-Kameyama M, Fujimoto Z, et al. (2018) GH30 Glucuronoxylan-Specific Xylanase from Streptomyces turgidiscabies C56. Appl Environ Microbiol 84 (4): e01850-17.
- Wang X, Yao B, Su X (2018) Linking Enzymatic Oxidative Degradation of Lignin to Organics Detoxification. Int J Mol Sci 19(11): 3373.
- Arocha-Garza H F, Canales-Del Castillo R, Eguiarte LE, Souza V, Torre Zavala S (2017) High diversity and suggested endemicity of culturable Actinobacteria in an extremely oligotrophic desert oasis. PeerJ 5:e3247.
- Singh NR, Dubey AK (2018) Diversity and Applications of Endophytic Actinobacteria of Plants in Special and Other Ecological. Front Microbiol 9:1767.
- Enkhbaatar B, Temuujin U, Lim JH, Chi WJ, et al. (2012) Identification and Characterization of a Xyloglucan-Specific Family 74 Glycosyl Hydrolase from Streptomyces coelicolor A3(2). Appl Environ Microbiol 78(2): 607-611.
- Yin LJ, Huang PS, Lin HH (2010) Isolation of Cellulase-Producing Bacteria and Characterization of the Cellulase from the Isolated Bacterium Cellulomonas Sp YJ5. J Agric Food Chem 58(17): 9833-9837.
- Ventorino V, Ionata E, Birolo L, Montella S, Marcolongo L, et al. (2016) Lignocellulose-Adapted Endo-Cellulase Producing Streptomyces Strains for Bioconversion of Cellulose-Based Materials. Front Microbiol 7:2061.
- Song JM, Wei DZ (2010) Production and characterization of cellulases and xylanases of Cellulosimicrobium cellulans grown in pretreated and extracted bagasse and minimal nutrient medium M9. Biomass Bioenerg 34(12): 1930-1934.
- An DS, Cui CH, Lee HG, Wang L, Kim SC, et al. (2010) Identification and Characterization of a Novel Terrabacter ginsenosidimutans sp nov beta-Glucosidase That Transforms Ginsenoside Rb1 into the Rare Gypenosides XVII and LXXV. Appl Environ Microbiol 76(17): 5827- 5836.
- Quan LH, Min JW, Jin Y, Wang C, Kim YJ, et al. (2012) Enzymatic Biotransformation of Ginsenoside Rb1 to Compound K by Recombinant beta-Glucosidase from Microbacterium esteraromaticum. J Agric Food Chem 60(14): 3776-3781.
- Lisov AV, Belova OV, Lisova ZA, Vinokurova NG, Nagel AS, et al. (2017) Xylanases of Cellulomonas flavigena: expression, biochemical characterization, and biotechnological potential. AMB Expr 7(1): 5.
- Fernandes AR, da Silveira WB, Passos FML, Zucchi TD (2014) Laccases from Actinobacteria-What We Have and What to Expect Tatiana. Advan Microbiol 4: 285-296.
- Ahmad M, Roberts JN, Hardiman EM, Singh R, Eltis LD, et al. (2011) Identification of DypB from Rhodococcus jostii RHA1 as a lignin peroxidase. Biochem 50(23): 5096-5107.
- Sugawara K, Nishihashi Y, Narioka T, Yoshida T, Morita M, et al. (2017) Characterization of a novel DyP-type peroxidase from Streptomyces avermitilis. J Biosci Bioeng 123(4): 425-430.
- Casciello C, Tonin F, Berini F, Fasoli E, Marinelli F, et al. (2017) A valuable peroxidase activity from the novel species Nonomuraea gerenzanensis growing on alkali lignin. Biotechnol Rep 13: 49-57.






























