Recent Advances in Genetics of Bacterial Arsenic Resistance Mechanisms and their Applications in Bioremediation
Srilakshmi Sunitha M, Prashant S, Rajasheker G and Kavi Kishor PB*
Department of Genetics, Osmania University, India
Submission: February 18,2019; Published: March 15, 2019
*Corresponding author: Kavi Kishor PB, Department of Genetics, Osmania University, Hyderabad - 500 007, India
How to cite this article: Srilakshmi Sunitha M, Prashant S, Rajasheker G, Kavi Kishor PB. Recent Advances in Genetics of Bacterial Arsenic Resistance Mechanisms and Their Applications in Bioremediation. Adv Biotech & Micro. 2019; 13(1): 555855. DOI: 10.19080/AIBM.2019.13.555855
Abstract
Arsenic is a toxic metalloid that occurs naturally in both aquatic and terrestrial environments. The wide spread nature of arsenic derivatives converts this element into a serious problem of public health worldwide. Considering its carcinogenicity and toxicity, remediation of arsenic-contaminated waters and soils have become a high priority. Bioremediation technology offers great potential for clean-up of arsenic polluted sites. In the global arsenic geocycle, microbes play a relevant role in its transformation. Numerous ars operons, with a variety of genes and different combinations were identified in diverse species. Besides the ars operon, which confers resistance to inorganic arsenic, recently additional genes were reported, which broaden the spectrum of arsenic resistance by detoxifying organic arsenic derivatives. This review briefly discusses the currently available bacterial genes involved in arsenic resistance mechanisms and their deployment in developing efficient bioremediation strategies.
Keywords: Arsenic metalloid; Arsenic resistant bacteria; Ars operons; Bioremediation
Introduction
Arsenic (As), the 20th most abundant element in earth’s crust is a deadly, major environmental pollutant. It is categorized as group 1 carcinogen by International Agency of Research on Cancer (IARC). It is released into the environment on a global scale as a result of geogenic, anthropogenic and biogenic activities and on local scale as a result of industrialization. Arsenic toxicity has become a global concern and high concentrations of it in groundwater have been reported from several countries including India. Arsenic toxicity, bioavailability and mobility vary depending on its state of oxidation. To combat the toxic consequences of arsenic mobilization, it is crucial to remove arsenic from soils and aquatic systems. In the present scenario, in situ bioremediation appears to be the most efficient, cost effective, environmentally friendly and a safe mechanism to detoxify arsenic. This involves the successful exploitation of native arsenic resistant bacteria and their genes associated with arsenic detoxification. This review focuses on the knowledge of currently available bacterial genes involved in arsenic resistant mechanisms and their use in developing efficient bioremediation strategies.
Advances in ars genes of arsenic resistant bacteria
In arsenic resistant bacteria, ars operon comprising two regulatory genes, ars R and ars D and three structural genes, ars A, ars B and ars C are the most well characterized genetic systems [1]. The operons are found either on chromosomes or on the plasmids in bacteria. The presence of arsenic resistance genes on plasmids offers an opportunity for microbes to disseminate these adaptive genetic traits by horizontal gene transfer to maintain the fitness among bacteria inhabiting polluted habitats [2]. Research reports on complex ars gene clusters with a wide variety of gene configurations were also reported [3]. Many bacteria with multiple, redundant ars genes appear to be frequent in arsenic contaminated environments [4]. However, the resistance levels to arsenic may not directly correlate with the number of ars operons. Genes like ars O and ars T were reported to be a part of ars operon, but their exact functions have not been deciphered yet [5]. Inorganic arsenic efflux pumps including Acr3, AqpS and Major Facilitator Superfamily (MFS) transporters, commonly linked to ars gene clusters are also reported to confer arsenical resistance. Over and above, arsenic resistance is also conferred by the paired gapdh and ars J genes [6], ars N gene [7]. Recently, a genetic mechanism for bacterial arsenic resistance was characterized which includes genes encoding Ars P permease, Ars H, Ars M and Ars I enzymes [8]. Figure 1 illustrates the arsenic transformation and resistance mechanisms in bacteria.
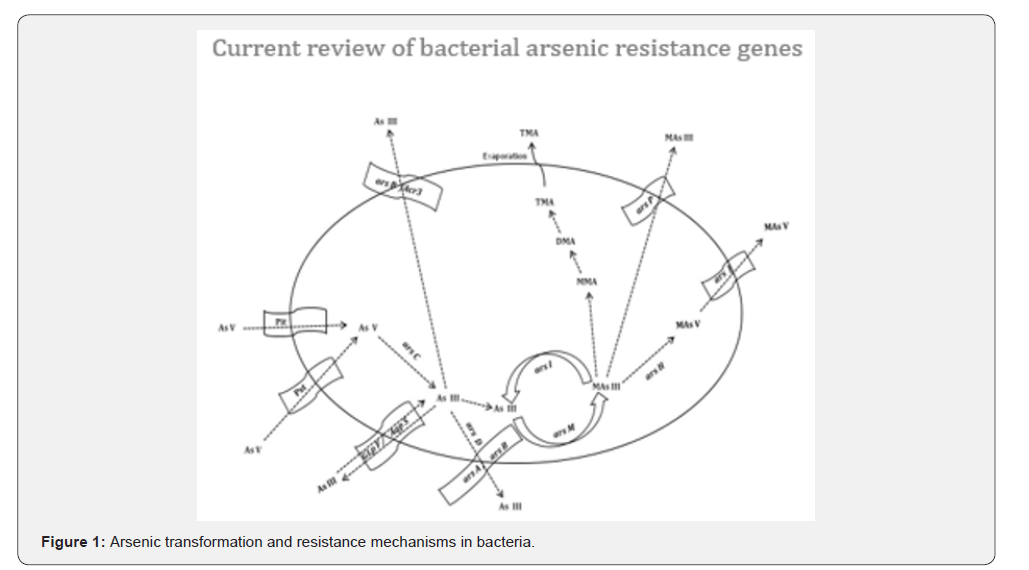
While As(V) enters the cells through Pit and Pst transporters, and As(III) enters through GlpF and AqpS. As(V) is reduced to As(III) by arsC. As(III) extrusion is carried out by arsB/Acr3 or arsA/arsB mediated by arsD. As(III) methylation occurs by arsM. MAs(III) extrusion takes place by arsP permease or oxidation by arsH to MAs(V). MAs(V) extrusion is carried out by arsJ. Demethylation of MAs(III) to As(III) is modulated by arsI.
Potential use of arsenic resistant bacteria in remediation
Many reports indicate the possible role of native microbial communities in in situ bioremediation of the arsenic contaminated sites [9, 10, 11]. The estuarine waters of Goa situated on the west coast of India, are known to be highly contaminated with arsenic besides other states like West Bengal and Bihar [12]. Earlier studies reported the arsenite tolerance and its biotransformation potential in estuarine bacteria isolated from waters of Goa, India [13]. The identification and characterization of ars genes may serve as potential molecular biomarkers to monitor the level of arsenic pollution in that environment [14]. However, studies related to the identification of ars genotype in arsenicresistant bacteria inhabiting these estuarine waters were not available. This prompted us to isolate diverse arsenic-resistant bacteria from the Mandovi and Zuari estuarine systems and characterize the ars genotype among the isolates. In our earlier study, 44 arsenic-resistant bacteria were isolated and identified. They belonged to the genera Brevibacterium, Acinetobacter, Providencia, Pseudomonas, Halomonas, Vibrio, Exiguobacterium and Staphylococcus.
The ars genotype characterization revealed the arsA, B and C genes were individually detected using PCR in 16, 9 and 13 bacterial isolates respectively. Besides, ars B and ars C genes were isolated from the genera Halomonas and Acinetobacter. The arsC gene isolated from Acinetobacter species complemented arsenate resistance in E. coli WC3110 and JM109 validating its function [15]. Hence, these bacteria with ars genes may play significant role in controlling the mobility of arsenic in environment and its subsequent detoxification by complexation.
Recent studies reported the use of genetically modified bacteria as biosensors to detect the inorganic arsenic pollution levels in the contaminated sites [16]. These biosensors are based on the gene constructs in which an ars R operator and promoter sequence is joined with the sequence of the reporter gene such as luciferase, β-galactosidase, an autofluorescent protein or cytochrome c peroxidase. Bioaccumulation of arsenic by indigenous bacteria and genetically engineered bacteria provide great opportunities for their use in bioremediation. Recently, Lysinibacillus strain B1-CDA and Corynebacterium glutamicum mutants [developed by removal of ars C and Acr3 efflux systems and also by over-expression of aquaglyceroporin genes (Glp F)] were reported to accumulate arsenic within the cell. Such a system contributes to clean up of waters polluted with arsenic metalloid [17,18].
Several microbial-assisted arsenic removal technologies incorporating biological transformation of arsenic and its subsequent adsorption by different compounds have also been developed [19]. Recently identified genes such as ars P, ars H, ars M and arsI can also be engineered to develop genetic models that could play crucial role in combating the toxic effects of organic
arsenic species in the environment. Another promising way of improving bioremediation processes is to deploy biofilters by expressing novel genes like phytochelatin synthase and metallothioneins in soil bacteria, which may scavenge inorganic arsenic species from polluted sites.
References
- Rosen BP (1999) Families of arsenic transporters. Trends Microbiol 7(5): 207-212.
- Martinez RJ, Wang Y, Raimondo MA, Coombs JM, Barkay T, et al. (2006) Horizontal gene transfer of PIB-type ATPases among bacteria isolated from radionuclide- and metal-contaminated subsurface soils. Appl Environ Microbiol 72(5): 3111-3118.
- Muller, D, Medigue C, Koechler S, Barbe V, Barakat M, et al. (2007) A tale of two oxidation states: bacterial colonization of arsenic-rich environments. PLoS Genet 3(4): e53.
- Páez-Espino AD, Durante-Rodriguez G, de Lorenzo V (2015) Functional coexistence of twin arsenic resistance systems in Pseudomonas putida KT2440. Environ. Microbiol 17(1): 229-238.
- Wang L, Chen S, Xiao X, Huang X, You D, et al. (2006) arsRBOCT Arsenic resistance system encoded by linear plasmid pHZ227 in Streptomyces sp. strain FR-008. Appl. Environ. Microbiol 72 (5): 3738-3742.
- Chen J, Yoshinaga M, Garbinski LD, Rosen BP (2016) Synergistic interaction of glyceraldehydes-3-phosphate dehydrogenase and ArsJ, a novel organoarsenical efflux permease, confers arsenate resistance. Mol Microbiol 100(6): 945-953.
- Chauhan NS, Ranjan R, Purohit HJ, Kalia VC, Sharma R (2009) Identification of genes conferring arsenic resistance to Escherichia coli from an effluent treatment plant sludge metagenomic library. FEMS Microbiol Ecol 67(1): 130-139.
- Yang HC, Rosen BP (2016) New mechanisms of bacterial arsenic resistance. Biomed J 39(1): 5-13.
- Dey U, Chatterjee S, Mondal NK (2016) Isolation and characterization of arsenic-resistant bacteria and possible application in bioremediation. Biotechnol. Rep 10: 1-7.
- Satyapal GK, Mishra SK, Srivastava A, Ranjan RK, Prakash K, et al. (2018) Possible bioremediation of arsenic toxicity by isolating indigenous bacteria from the middle gangetic plain of Bihar, India. Biotechnol Rep 17: 117-125.
- Biswas R, Vivekanand V, Saha A, Ghosh A, Sarkar A (2019) Arsenite oxidation by a facultative chemolithotrophic Delftia spp. BAs29 for its potential application in groundwater arsenic bioremediation. International Biodeteriorat & Biodegrad 136: 55-62.
- Nair M, Joseph T, Balachandran KK, Nair KKC, Paimpillii JS (2003) Arsenic enrichment in estuarine sediments-impact of iron and manganese mining. Fate of arsenic in the environment. In: Ahmed, MF, Ali MA, Adeel Z (eds.) (Int. Symp. on Fate of Arsenic in the Environment, Dhaka; Bangladesh; Feb 2003). International Training Network (ITN), Dhaka, Bangladesh, pp 57-67.
- Nagvenkar GS, Ramaiah N (2010) Arsenite tolerance and biotransformation potential in estuarine bacteria. Ecotoxicol 19(4): 604-613.
- Stocker J, Balluch D, Gsell M, Harms H, Feliciano J, et al. (2003) Development of a set of simple bacterial biosensors for quantitative and rapid measurements of arsenite and arsenate in potable water. Environ Sci Technol 37: 4743-4750.
- Sri Lakshmi Sunitha M, Prashant S, Chari PVB, Rao SN, Balaravi P, et al. (2011) Molecular identification of arsenic-resistant estuarine bacteria and characterization of their ars genotype. Ecotoxicol 21(1): 202-212.
- Kruger MC, Bertin PN, Heipieper HJ, Arsene-Ploetze F (2013) Bacterial metabolism of environmental arsenic-mechanisms and biotechnological applications. Appl. Microbiol Biotech 97(9): 3827- 3841.
- Rahman A, Nahar N, Nawani NN, Jass J, Desale P, et al. (2014) Isolation and characterization of a Lysinibacillus strain B1-CDA showing potential for bioremediation of arsenics from contaminated water. J Env Sci Health 49(12): 1349-1360.
- Villadangos AF, Ordóñez E, Pedre B, Messens J, Gil JA, et al. (2014) Engineered coryneform bacteria as a bio-tool for arsenic remediation. Appl. Microbiol. Biotechnol 98(24): 10143-10152.
- Wan J, Klein J, Simon S, Joulian C, Dictor MC, et al. (2010) AsIII oxidation by Thiomonas arsenivorans in up flow fixed-bed reactors coupled to As sequestration onto zerovalent iron-coated sand. Water Res 44(17): 5098-5108.






























