Degradation of Phenol, an Innovative Biological Approach
Arutchelvan V1, Atun Roy Choudhury2*
1Department of Civil Engineering, Annamalai University, India
2Ramky Advanced Centre for Environmental Research, Ramky Enviro Engineers Limited, India
Submission: November 02,2018;Published: January 11, 2019
*Corresponding author: Atun Roy Choudhury, Department of Civil Engineering, Annamalai University, Annamalainagar-608002, Tamil Nadu, India
How to cite this article: Arutchelvan V, Atun R C. Degradation of Phenol, an Innovative Biological Approach. Adv Biotech & Micro. 2019; 12(2): 555835. DOI: 10.19080/AIBM.2019.12.555835
Abstract
A bacterial strain capable of utilizing phenol as a sole carbon source was isolated from the soil samples, collected at a vicinity of industrial wastewater treatment plant of phenol manufacturing unit. Based on the biochemical tests and 16S rRNA sequencing results the organism was identified as Serratia marcescens. The organism is capable of degrading the highly toxic phenolic compound to an optimum concentration of 2500 mgL-1 in 120 h at approx. a neutral pH of 8. The organism was also efficient with wide pH and temperature tolerance with an extremely stipulated lag phase in the case of higher influent concentrations. Amid two different models incorporated to justify the growth kinetics of the organism the Haldane’s model fits (R2=0.930) very satisfactorily with kinetic constants in the range of Umax = 0.05 - 0.095 h-1; Ks = 8.49 - 16.1 mgL-1; Kj - 1154.75 - 1700.68 mgL-1. Despite limitations in terms of inhibition of microbial growth subjected to an extremely elevated concentration of phenol the study claims the feasibility of Serratia marcescens to an utmost concentration of 2500 mg/L and suggests the method as best possible alternative to the existing chemical treatment procedures.
Keywords: Biodegradation; Phenol; Serratia marcescens; Kinetic modeling; Phenolic compounds
Introduction
Phenol and its higher homologues are aromatic molecules containing hydroxyl group attached to the benzene ring. Phenolic compounds are widely distributed in the environment from various industrial as well as natural sources [1-3]. Phenol pollution is associated with petroleum refineries, pharmaceuticals, pulp mills, gas and coke oven industries, explosive manufacture, phenol- formaldehyde resin manufacture, plastic and varnish industries, wood processing industries etc [4-7].
Natural sources of phenol include forest fire, natural run-off from urban area where asphalt is used as the binding material and natural decay of ligno-cellulosic materials. The presence of phenol in water imparts carbolic odor to receiving water bodies and can cause toxic effects on aquatic flora and fauna [8]. Phenolic compounds are hazardous to human health and resistant to biodegradation [9,10]. It acts as a substrate inhibitor in the biotransformation [11]. Phenol is also a priority pollutant and is included in the list of EPA [12].
Several physico-chemical treatments have been suggested in the last two decades to remove phenol efficiently: adsorption with bone char or zeolites, stripping with air or stream [13] wet air oxidation etc. [14]. On the other hand, biological treatments with pure or mixed cultures of microorganisms have also been used due to its eco-friendly, cost effective and complete bio-mineralization [15]. Bacteria, yeast and fungi are capable of utilizing Phenolic compounds. Although phenol is a toxic and substrate inhibitory, it is also a carbon and energy source for several bacterial strains belonging to the species of Pseudomonas, Bacilli, Klebsiella, Ochrobactrum, Rhodococcus, Aureobasidium etc [16-21]. In the present investigation, a novel high strength phenol degrading bacterial strain was isolated, identified and its phenol utilization was studied under varied experimental conditions like the effect of pH, initial phenol concentration and inoculum size. Also, the results were validated in Monod and Haldane’s models.
Material and methods
All fine chemicals used were of analytical grade from Merck, India. The chemicals used for media preparations were of Bacteriological grade from Hi Media, India. Glass wares were sterilized in hot air oven at 180 °C for 3 h. All growth media, water blanks were steam sterilized m autoclave at 121 °C, 15 lb/inch for 20 min. pH of the growth media was adjusted to 8.01 using phosphate buffers.
Collection of samples
The soil samples rich in phenol were collected from six different points from the vicinity of the effluent treatment plant of phenol-formaldehyde resin manufacturing industry, Chennai India and stored in plastic containers at 4°C as per the standard procedure [22]. The industry is discharging phenol concentration of about 4000 mgL-1 in the effluent [23].
Isolation of bacteria
The medium used for isolation of bacteria contains glucose 5.0 gL-1, peptone 5.0 gL-1, NaCl 5.0 gL-1, beef extract 3.0 gL-1, phenol 100 m gL-1 and 10 gL-1 of soil sample. The broth was inoculated with soil and kept for 48 h at 25 °C under aerobic conditions in a mechanical shaker at 100 rpm. Aliquot from the enrichment flask was plated on an agar plate containing glucose 5.0 g L-1, peptone 5.0 gL-1, NaCl 5.0 gL-1, beef extract 3.0 gL-1, agar 20 gL-1 and phenol 200 mgL-1. The plates were incubated at 25 °C for 36 h under aerobic condition. Colonies developed were carefully observed for the uniformity and differences. Four morphologically distinct colonies were obtained. The colonies were transferred to new plates and further screened by sub-culturing in a basal minimal medium containing (per liter): NH4C1 10 g, NH4NO3 4 g, K2HPO4 0.2 g, KH2PO4 0.8 g, MgSO4.7H2O 0.2 g and 200 mgL-1 of phenol as the sole carbon and energy source.
The morphological, physiological, biochemical tests were carried out to identify the selected efficient isolate as per Bergy’s Manual of systematic Bacteriology [24]. The bacterium was further confirmed by 16S rRNA sequencing from Microbial Type Culture Collection and Gene Bank, Institute of Microbial Technology, Chandigarh, India.
Phenol degradation
The experimental studies were performed in the shake flask as batch reactor. Each flask contains sample of 100 ml basal minimal medium with varying concentrations of phenol as sole carbon source. The shake flasks were maintained at 34 °C±0.1°C at 150 rpm. The samples were regularly analyzed for phenol concentration and cell growth.
Growth of the organism was recorded by monitoring the Optical Density (OD) of the culture in a Hitachi-U-2001 UVvis Spectrophotometer at 600 nm. Phenol was estimated spectrophotometricaly using 4-aminoantipyrine as per standard procedure [22].
Results and Discussion
Isolation and Identification
The soil samples were collected from the different locations of effluent treatment plant of phenol-formaldehyde resin manufacturing industry. The samples were inoculated in the basal minimal medium containing phenol as a sole carbon source. Eighteen morphologically different colonies were developed, out of which four were distinct and one of the isolate exhibits more growth in high phenol concentration.
The morphological, physiological, biochemical and carbon source utilization characteristic of the isolate is given. The biochemical test results expressed the organism was a Gramnegative with brown pigment producing, having a proficient growth between 10 and 37°C, pH tolerance to a wide range of 5.0 - 11.0 and growth on NaCl upto 7%. The metabolic activity of the isolate was dynamic both in aerobic and anaerobic conditions, showing the organism as a facultative. The organism has positive results on MacConkey agar and Voges Proskauer test. It utilizes citrate, hydrolyze casein and reduces nitrate. It produces acid from the tested carbohydrates viz. adonitol, arabinose, dextrose, melibiose, salicin and xylose. Based on these characteristics and 16S rRNA sequencing results, the isolate has been identified as Serratia marcescens.
Influence of Initial concentration
Shake flask experiments were conducted to examine the effect of various initial concentrations of phenol on the degradation behavior of S. marcescens at 34±0.1 °C. The results of batch studies for high strength phenol degradation in minimal media by S. marcescens are given. The organism degrades phenol with an initial concentration of 1000, 1500, 2000 and 2500 mgL-1 at about 98.5, 99.1, 98.9 and 98.2 % in 72, 96, 108 and 120 h respectively. At higher the initial concentration of phenol more time required for the organism to degrade.
The growth of organism increases with increase in phenol concentration as, but the lag phase of the organism was extended at higher concentration of phenol. A positive correlation between cell biomass and phenol degradation was noted. In the batch studies, the biomass concentration of S. marcescens increases after 24 h of inoculation. During initial stages of biodegradation, the corresponding biomass was less, subsequently in stages the biodegradation was much pronounced with less increase in biomass and this result was in conformity with the low conversion efficiency to biomass with high removal of phenol [25]. This organism more effectively degrades 2500 mgL-1 of phenol in 120 h. The results of cell biomass concentration versus time indicate that the lag phase increase with increase in phenol concentration. This organism has a very short lag phase and thus overall degrading period was reduced. The short lag phase character of the organism may also help to optimize the design of bioreactor.
During the exponential growth phase of the organism the phenol degradation was maximum. Further, the substrate removal rate was relatively slow towards the end of the substrate consumption. This may be due to deficit in availability of oxygen as well as the substrate for the whole biomass, which in turn the fall in pH of the solution during the degradation period being noticed. The fall in the oxygen and pH of the solution during degradation has been reported when the phenol was metabolized by mixed culture composed Pseudomonadaceae, Vibrionaceae, etc. [26-33].
Inoculum size
The inoculum concentration in the shake flasks were varied from 2 to 6 % (v/v). Six percent (v/v) of biomass concentration provided the best degradation in short duration and 5 % (v/v) concentrations also have an efficiency of degradation equivalent as 6 % concentration. Since the results show that the lag phase for all concentrations of phenol is almost same, the optimum size of inoculum (5%) can be adopted for S. marcescens. Hill and Robinson (1975) concluded that not only the inhibitory effect of the substrate but also the size of the inoculum might affect the duration of lag phase.
Growth Kinetics of the isolated organism in presence of phenol
After isolation and identification of organism, determination of growth and degradation kinetics of the same organism has also been considered in this study. The study of growth kinetics is vital to know more about the capacities of the organisms in degradation and to design an appropriate treatment unit. The observation made on phenol inhibition on the growth of culture can be modeled using proper substrate inhibition model described in literature [7,10,28].
By considering phenol as non-inhibitory compound for an adapted population, the microbial growth can be represented by a simple Monod’s non-inhibitory kinetic Eq. [29]
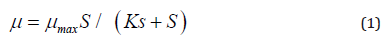
Where, p is the specific growth rate (h-1), S is the limiting substrate concentration (mg L-1), μ max is the maximum specific growth rate (h-1), Ks is the half saturation co-efficient (mg L-1). However, this Eq. becomes unsatisfactory for explaining inhibitory growth of microorganism at higher substrate concentrations. In such a case to represent the growth kinetics of inhibitory compounds, several kinetic models were fitted to the experimental data for selecting the most suitable models. Amongst Haldane’s model was adopted due to its mathematically simplicity and wide acceptance for representing the growth kinetics of inhibitory substrates. The Haldane or Andrews’s model has the same form (Kennes and Veiga, 2001) as:
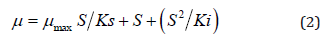
This is the linearized Haldane’s Eq.
The experimental data on specific growth rate (μ) obtained at various initial phenol concentrations (So) were used for estimating the kinetic parameters from the above two models. the experimental specific growth rate and model predicted ones. From the fig., it could be evident that Haldane model was found to fit the data well. the values of the growth kinetic parameters for various species obtained from different researchers for phenol and were found to be higher compared to the literature report. The Haldane model is more appropriate to explain the substrate inhibition on growth of a microbial culture. Therefore, Haldane’s model fitted the experimental data obtained in the study reasonably well. The inhibition constant estimated by Haldane model was high as compared to the values reported in literature [7,30,31].
Endogenous or decay coefficient
The endogenous growth phase curve in a batch reactor has been modeled using the
Eq. dx/xy = − kd X (4)
Where, Kd is the first-order death rate constant
And X is cell mass concentration
In order to determine the value of kd, the growth of culture was continued, and the cell mass concentration was observed for few days even after the complete consumption of phenol. The selection of the particular growth run was arbitrary, assuming that the kd is not dependent on initial concentration. The values of the decay rate coefficients obtained were in the range of 0.002 - 0.042 h-1 for phenol. Kumaran and Paruchuri [2] have reported the value of decay coefficient as 0.005 h-1 for phenol degradation by a mixed culture. The values obtained in this study are comparable with these values of decay coefficients. The decay coefficient affects the growth kinetics. The growth rate will be reduced by as much the value of decay coefficient so that the wash out condition would occur at lower dilution rates.
Yield coefficient
The values of the yield coefficient for various phenol concentration ranges from 0.243 to 1.157 mg VSS/mg bs COD. These values are comparably smaller than the values obtained by other researchers, this may be due to high substrate concentrations.
Conclusion
Biodegradation of highly toxic inorganic compounds provoked the idea of isolation and utilization of a micro-organic pure strain. In this context, a pure culture of S. marcescens was isolated from the soil samples collected near the effluent treatment plant of phenol-formaldehyde resin manufacturing industry. The same found to be extremely effective in absolute degradation of high strength phenolic compounds, concentration ranging from 1000 to 2500 mg L-1. The duration taken up by the species to degrade the maximum concentration of 2500 mg L-1 was 120 h. This favorable degradation was obtained with pH 8.0, temperature 34±0.1°C and biomass concentration 5% (v/v). The organism has a very short lag phase at high substrate concentration. The substrate inhibition due to phenol and the growth kinetics of the culture was explained using the Monod non- inhibitory model and Haldane’s substrate inhibitory model. Between the two models, the Haldane model gave a better fit (R2=0.930) to the experimental data. The bio-kinetic constants estimated shows the organism is efficient in phenol degradation.
References
- Hirayama KK, Tobita S, Hyrayama K (1994) Biodegradation of phenol and mono chlorophenols by yeast Rhodotorula glutinis. Water Science Technology 30(9): 59-66.
- Kumaran P, Paruchuri YL (1997) Kinetics of phenol biotransformation. Water Research 31(1): 11-22.
- Bobdziewicy J (1998) Biodegradation of phenol by enzyme from Pseudomonas sp. Immobilized onto ultrafiltration membranes. Process Biochemistry 33(8): 811-818.
- Ahmed AM, Nakhla GF, Farooq SJ (1995) Phenol degradation by Pseudomonas aeruginosa. Journal of Environmental Science and Health 30(1): 99-107.
- Van Schei PM, Young LY (1998) Isolation and Characterization of phenol degrading denitrifying bacteria. Appl Environ Microbiol 64(7): 2432-2438.
- Bandhyopadhyay K, Das D, Bhattacharya P, Maiti BR (2001) Reaction engineering studies on biodegradation of phenol by Pseudomonas putida MTCC1194 immobilized on calcium alginate. Biochemical Engineering Journal 8(3): 179-186.
- Kumar A, Kumar S, Kumar S (2005) Biodegradtion kinetics of phenol and catechol using Pseudomonas putida MTCC1194. Biochemical Engineering Journal 22(2): 151-159.
- Ghadhi SC, Sangodkar UMX (1995) Potentials of Pseudomonas cepacia PAA in bioremediation of aquatic wastes containing phenol. Proceedings of National symposium frontiers in applied and environmental microbiology 11-13.
- Sufit DJ (1978) Some effects of exposing rainbow trout in (Salmo gairdneri Richardson) phenol solution. Journal of Fish Biology 13(1): 7- 17.
- Nuhoglu A, Yalcin B (2005) Modeling of phenol removal in a batch reactor. Process Biochem 40(3-4): 233-239.
- Prieto MB, Hidalgo A, Rodriguez FC, Serra JL, Llama MJ (2002) Biodegradation of phenol in synthetic and industrial wastewater by Rhodococcus erythropitics. UPV-1 immobilized in air-stirred reactor with clarifier. Appl Microbiol Biotechnol 58(6): 583- 859.
- EPA (1979) Phenol ambient water quality criteria. Office of the planning and standards. Environ Protect Agency, Washington pp.296- 786.
- Zilli M, Converti A, Lodi A, Borghi MD, Ferraiolo G (1993) Phenol removal from waste gases with a biological filter by Pseudomonas putida. Biotechnol Bioeng 41(7): 693-699.
- Lin SH, Chung TS (1994) Combined treatment of Phenolic wastewater by wet air oxidation and activated sludge. Toxicological & Environmental Chemistry 44(3-4): 243-258.
- Tay ST, Moy BY, Maszenan AM, Tay JH (2005) Comparing activated sludge and aerobic granules as microbial inocula for phenol biodegradation. Appl Microbiol Biotechnol 67(5): 708-713.
- Balasankar T, Nagarajan S (2000) Biodegradation of phenol by Bacillus brevis. Asian Jr Microbio Biotech Env Sci 2: 155-158.
- Neumann G, Teras R, Monson L, Kivisaar M, Schauer F (2004) Simultaneous degradation of atrazine and phenol by Pseudomonas spp. Strain ADP: effect of toxicity and adaptation. Appl Environ Microbiol 70(4): 1907-1912.
- Arutchelvan V, Kanakasabai V, Nagarajan S, Muralikrishnan V (2005) Isolation and identification of novel high strength phenol degrading bacterial strains from phenol- formaldehyde resin manufacturing industrial wastewater. J of Hazard Mater 127(1-3): 238- 243.
- Hsieh F, Huang C, Lin T, Chen Y, Lin J (2008) Study of sodium tripolyphosphate- crosslinked chitosan beads entrapped with Pseudomonas putida for phenol degradation. Process Biochemistry 43(1): 83-92.
- Shourian M, Noghabi KA, Zahiri HS, Bagheri T, Karballei G, et al. (2009) Efficient phenol degradation by a newly characterized Pseudomonas sp. SA01 isolated from pharmaceutical wastewaters. Desalination 246 (1- 3): 577-594.
- Santos VL, Monteiro AS, Braga DT, Santoro MM (2009) Phenol degradation by Aureobasidium pullulans FE13 isolated from industrial effluents. J of Hazard Mat 61(2- 3): 1413-1420.
- APHA (2005) Standard Methods for the Examination of Water and Wastewater 20th ed, Washington, USA.
- Arutchelvan V, Kanakasabai V, Elangovan R, Nagarajan S (2004) Physico-Chemical characteristics of wastewater from Bakelite manufacturing industry. Indian J Environ Ecoplan 8: 757-760.
- Noel RK, John GH (1984) Bergy’s manual of systematic bacteriology, Vol I Eighth ed, Williams & Wilkins, Baltimore, USA.
- Yoong ET, Lant PA, Greenfield PF (1997) The influence of high phenol concentration on microbial growth. Water Science Technology 36(2- 3): 75-79.
- Lallai A, Mura G (1989) pH variation during phenol biodegradation in mixed cultures of microorganisms. Water Research 23(11): 1335- 1338.
- Blanch HW, Clark DS (1996) Biochemical Engineering, Marcel Dekker Inc, New York.
- Okpokwasili GC, Nweke CO (2005) Microbial growth and substrate utilization kinetics. Afr. African Journal of Biotechnology 5(4): 305- 317.
- Kovar KK, Egli T (1998) Growth kinetics of suspended microbial cells: from single - substrate - controlled growth to mixed-substrate kinetics. Microbiology and Molecular Biology Reviews 62: 646-666.
- Rodriguez GV, Youssef CB, Vilanova JW (2006) Two-step modeling of the biodegradation of phenol by acclimated activated sludge. Chem Eng J 117(3): 245-252.
- Arutchelvan V, Kanakasabai V, Elangovan R, Nagarajan S, Muralikrishnan V (2006) Kinetics of high strength phenol degradation using Bacillus brevis. J of Hazard Mat 129(1-3): 216-222.
- Chitra S, Sekeran G, Padmavahti S, Gowri C (1995) Removal Phenolic compounds from wastewater using mutant strain of Pseudomonas pictorium. J Gen Appl Microbio 41: 229-237.
- Hill GA, Robinson CW (1975) Substrate inhibition kinetics: phenol degradation by Pseudomonasputida. Biotechnology Bioengineering 17(11): 1599-1615.






























