Rhizospher fungi Identified from Poaceae and Cyperaceae Family Grass in North and East Shewa: Ethiopia
Birhanu Gizaw*, Zerihun Tsegay, Genene Tefera, Endegena Aynalem and Abebe Tesfaye
Department of Microbial Biodiversity, Ethiopian Biodiversity institute, Ethiopia
Submission: August 13, 2018; Published: September 27, 2018
*Corresponding author: Birhanu Gizaw, Department of Microbial Biodiversity, Ethiopian Biodiversity institute, P.O. Box 30726 Addis Ababa, Ethiopia, Tel: +251911862561/0933578070; Email: gizachewbirhan@gmail.com
How to cite this article: Birhanu G, Zerihun T, Genene T, Endegena A, Abebe T. Rhizospher fungi Identified from Poaceae and Cyperaceae Family Grass in North and East Shewa: Ethiopia. Adv Biotech & Micro. 2018; 11(4): 555818. DOI:10.19080/AIBM.2018.11.555818
Abstract
The rhizosphere is a hotspot of plant-microbe interactions with profound influence on plant productivity and ecosystem function. The aim of this research is to isolate and identify Rhizospher fungi from Poaceae and Cyperaceae family grass for agricultural application to use as biofertilizer, biocontrol and biopesticide. Rhisospher soil and root sample were collected from 18 Poaceae and Cyperaceae famiy grass. Malt and potato dextrose agar were used to primary and secondary fungiculture. Pure fungiisolates were transferred to biolog universal yeast agar media. Pure yeast cells and filamentous fungiwere suspended in sterile water and Filamentous fungi(FF) inoculum fluid at 49±2 and 75±2 turbidity measured by biolog turbidimeter, respectively. 100 μL transferred from each suspension into 96 wells of the biolog yeast microplate and filamentous fungimicroplate tagged with different carbon source and incubated at 26 °C for 24 to 72 h for yeast and 24-240 h for filamentous fungiand read by micro station at a single wavelength of 590 nm, The Biolog software micro log3 ver. 4.20.05 compared the results obtained with the test strain to the database and provided identification based on distance value of match and separation score produces similarity index value and probability results were recorded and processed for identification by micro log3 software ver. 4.20.05. Biolog microstation read 27 fungispecies. The result revealed that Fellomyces fuzhounsis, Schizoblastosporon starkeyi-henricii, Cryptococcus luteolus, Rhodotrula aurantica A, Cryptococcus terreus A, Trichosporon beigeliiB, Rhodotrula aurantica B, Cryptococcus albidus var aerius, were fully identified above 75% probability and 0.5 simlarity index. In conclusion these rhizopspher- microbes interaction study useful for agricultural application for biofertilizer after further evaluation.
Keywords: Filamentous fungi; Fellomyces fuzhounsis; Schizoblastosporon Starkeyi-henricii; Cryptococcus luteolus; Rhodotrula aurantica A; Cryptococcus terreus A; Trichosporon beigeliib; Rhodotrula aurantica B; Cryptococcus albidus ; Aerius; Oryza l; Zea l; Poaceae; Cyperaceae; Biofertilizer; Rhizosphere; Plant growth; Agriculture; Population
Introduction
Poaceae are the fifth largest family of flowering plants following the Asteraceae, Orchidaceae, Fabaceae and Rubiaceae. Globally, about 12,000 grass species in about 771 genera that are classified into 12 subfamilies and the family is economically important because it includes Teff (Eragrottis teff), wheat (Triticum L.), rice (Oryza L.) and corn (Zea L.), as well as numerous forage, bamboo and biofuel grass species [1]. Grasses grow on all continents in tropical, temperate and Arctic zones and are absent only from Antarctica [2]. Grasses have long had significance in human society for feed and fodder for people and domesticated animals for thousands of years. During seed germination and seedling growth of this grass, there is great interaction with a range of microorganisms present in the surrounding soil. Root exudates from this grass are mainly composed of water-soluble sugars, organic acids, and amino acids, hormones, vitamins, amino compounds, phenolics and sugar phosphate esters [3]. Broadly, there are three distinct components recognized in the rhizosphere; the rhizosphere per se (soil), the rhizoplane, and the root itself.
The rhizosphere is thus the zone of soil influenced by roots through the release of substrates that affect microbial activity. The rhizoplane is the root surface, including the strongly adhering root particles. The root itself is a part of the system, because certain endophytic microorganisms are able to colonize inner root tissues [4]. Microorganisms present in the rhizosphere play important roles in ecological fitness of their plant host. The rhizosphere is a hotspot of plant-microbe interactions with profound influence on plant productivity and ecosystem function [5]. Shaped by the release of labile carbon (C) from plant roots and root uptake of nutrients and water, the physiochemical environment of the rhizosphere supports a microbial community compositionally and metabolically distinct from that found in bulk soil [6]. The resulting rhizosphere microbiome performs critical functions, modulating plant growth and development [7], plant health, and plant nutrient acquisition [5] as well as the production of antibiotics, geochemical cycling of minerals and plant colonization [8].
Plant-microbe interactions may thus be considered beneficial, neutral, or harmful to the plant, depending on the specific microorganisms and plants involved and on the prevailing environmental conditions [9]. Exploring these microorganisms by unravelling their possible relationships with plants has launched a new and fascinating area of investigations in the rhizosphere research. A better understanding of the basic principles of the rhizosphere ecology, including the function and diversity of inhabiting microorganisms is one ways to improve agriculture and to reduce improper use of chemical pesticides and fertilizers creating a long list of environmental and health problems, Thus, the exploration of rhizospher microorganisms from grass family as one of the best options to increase biomass yield of the Teff crops through developing biofertilizer, biopesticide, biocontrol, In general understanding microbial diversity in rhizospher of grass family useful for compost making, bioremediation and vast agricultural as well as industrial application. There for the aim of this research is isolation and identification of rhizospheric fungi from different grass species at different agro ecology in later for use as agriculture input.
Materials and Methods
Study area
The study was conducted in North and East Shewa in 10 selected districts, in North Shewa zone particularly in Kewot, Tarmaber, Efratana gidim, Siadeberna wayu and Ankober. North shewa zone is one of the 10 zones of Amhara regional state. The elevation ranges from 1100 to 3009 m above sea level. Geographic coordinate latitude: 9°46’8.4” and longitude: 39°40’4.8”. The zone is located in approximately average200 km far from Addis Ababa. East shewa is found in, Oromia regional state, It has 12 districts according to the zonal statistics and information center, study in east shewa comprise Adama, Bost, Adaa, Gimbichu and Lomme. East Shewa is found between 38° 57’and 39° 32’ E and 7° 12’ and 9° 14’N. 93% of the district falls in the lowlands in rift valley. The average altitude is 1600 m, but rises up to 2300 m at the north western and western mountain fringes of the rift on one hand, and it falls to 900-1000m towards north east. Agro ecology of the zone is divided in to three, 0.2% area of the zone is found in the high land, 61.1% midland and the rest 38.7% is found in the low land (Figure 1).
Sample collection
Six hundred forty rhizospher soil through drillings at 5, 10, and 15 cm depth and 640 root samples from Poaceae and Cyperaceae family grass in sterile sample tube were collected during August2017 to March 2018G.C (Table 1, Figure 2&3). Approximately 5 g of soil were taken from each depth of sampling point for a total of 15g composite soil stored in sterile sample tube via icebox and transported to microbial directorate laboratory in Ethiopian biodiversity institute to Addis Ababa and kept in +4 °C until processed.
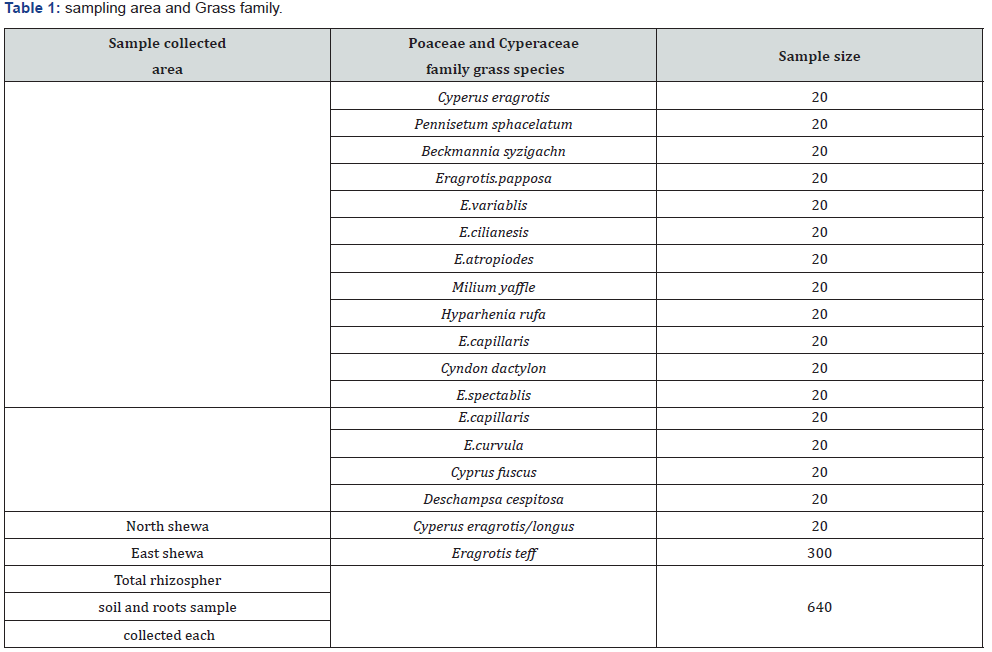


Screening and isolation of rhizospher fungi from Poaceae and Cyperaceae family grass
Six hundred forty soil samples were clustered according to altitude, soil type and grass varieties and merged into 40 composite samples. From each soil samples 1 g was taken and diluted in distilled water serially up to 10-6 mL. In the same way the root samples were clustered and washed by distilled water. Finally, roots were disinfected with 3% Hypochlorate, 70% ethanol alcohol for 3 to 5 minutes and rinse 6 times by distilled water. Finally surface sterilized root were crushed and diluted by distilled water the same way to soil sample. About 0.1 mL inoculum sample was transferred by swab through and streaked by nichrom loop on Malt extract agar and potato dextrose agar. Primary cultures were incubated for 26 °C in digital incubator for 48 h. Isolates were subculture twice until pure colony obtained for morphological identification. A single yeast colony and pure filamentous fungi was streaked to Biolog universal yeast agar (BUY agar plate, (60 g / 1 L) and incubated for 48 h at26 °C for micro plate inoculum preparation.
Identification of rhizospher fungi from Poaceae and Cyperaceae family grass
Based on primary and secondary colony morphology fungi isolates were screened and transferred to biolog universal yeast agar media. Pure fungi cell were suspended in 9 mL sterile water for yeast and 15 mL Filamentous fungi(FF) inoculum fluid for filamentous fungi at 49±2 and 75±2 turbidity measured by biolog turbidimeter, respectively. 100 μL transferred using digital pipettor from each suspension into 96 wells of the biolog yeast microplate (YT) and Filamentous fungi microplate (FF) tagged with different carbon source and incubated at 26 °C for 24 to 72 h for yeast and 24-240 h for filamentous fungi and read by micro station readear at a single wavelength of 590 nm. The Biolog software micro log3 ver. 4.20.05 compared the results obtained with the test strain to the database and provided identification based on distance value of match and separation score produces similarity index value and probability. Acceptable species identification must have similarity index value above or equal to 0.5 or probability above or equal to 75% were chosen only for species identification and characterization [10].
Result
From Poaceae and Cyperaceae family grass a total of 476 rhizospher fungal colony were grown on Malt extract agar and Potato dextrose agar. The fungal isolates were screened, and morphologically similar colonies were clustered. Representative clusters were transferred in to YT/FF Microplate for BiOLOG Micro Station reading. Finally, 27 fungal species (21 yeast and 6 filamentous fungus) were read by MicroStation reader and the result revealed that 8 yeast species >0.5 similarity index and >75%probability were identified. These are Fellomyces fuzhounsis, Schizoblastosporon starkeyi-henricii, Cryptococcus luteolus, Rhodotrula aurantica A, Cryptococcus terreus A, Trichosporon beigeliiB, Rhodotrula aurantica B, Cryptococcus albidus var aerius. Where as19 fungal species its similarity index were below 0.5. There for eight rhizospher, rhizoplan and endophyte yeast were fully identified from five Grass species in the study area (Figure 3 & Table2). Their percentage frequency on MEA media was recorded, the result revealed Trichosporon beigelii B (24.5%), Rhodotrula. aurantiac A (21.5%), Cryptococcus terreus A (20%), Rhodotrula aurantiac B (17.5%), Cryptococcus luteolus (9%), Cryptococcus albidus var aerius (7.5%). The Cryptococcus genera (36.5%) yeast was the dominant in this study. Filmentous fungi also confirmed by cellular morphology using Lacophenol cotton blue staining, the result revealed that Penicillium genera were dominant (Table 2).
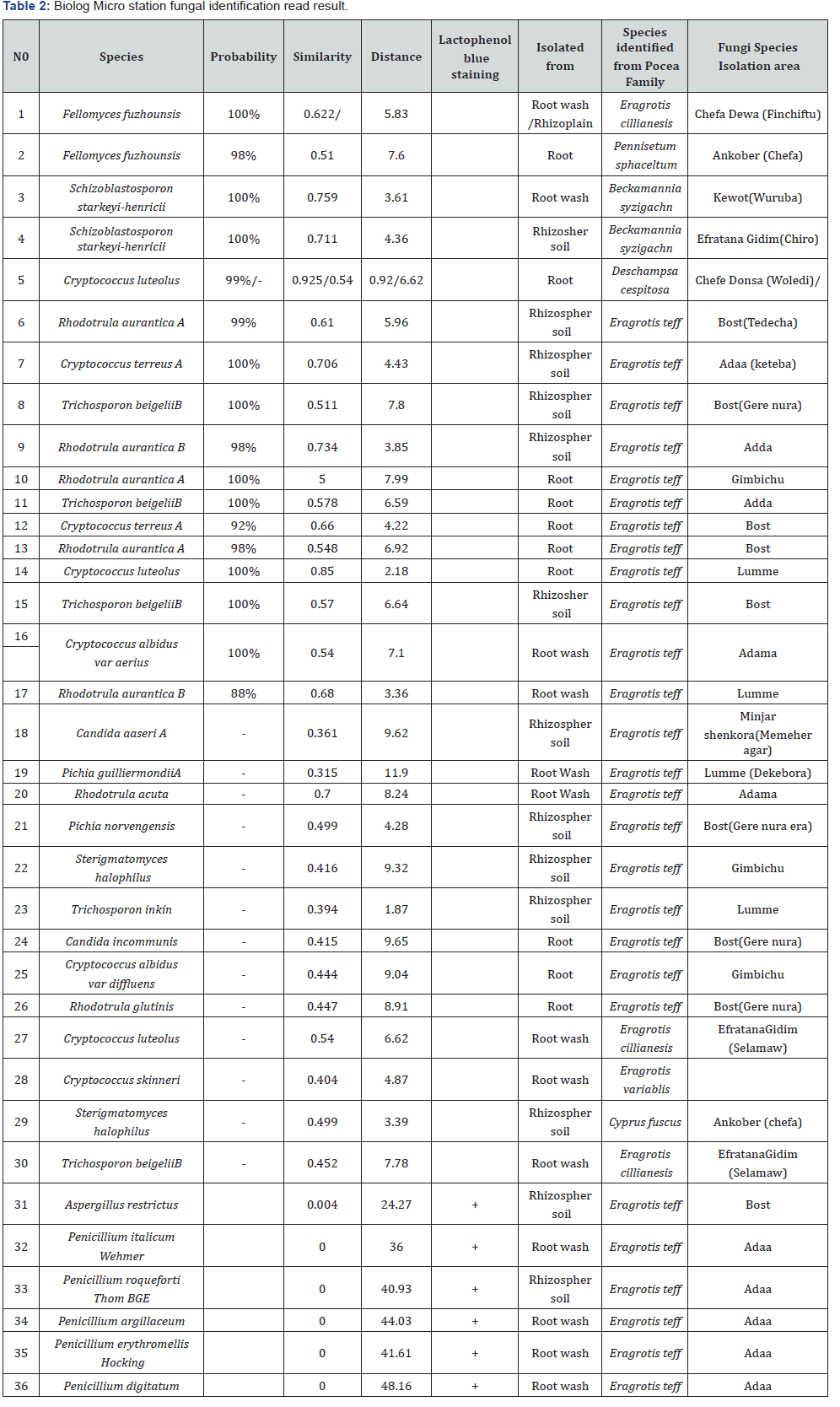
Discussion
Rhizodeposition of carbon compounds dramatically increases microbial activity and biomass in soil closely associated with the roots [11]. Integrated interactions between plants and microbes have superior importance for improving plant growth as well as maintaining proper soil conditions. Rhizosphere interactions that are based on complex exchange are more complicated than those occurring above soil surface or non-rhizosphere soil. Among diverse microbial population, plant growth Promoting rhizobacteria (PGPR) gain special attention owing to their multifarious functional characters like effective root colonization, nitrogen fixation, hormone production, solubilization of nutrients, and production of certain enzymes that are beneficial for sustainable agriculture. An understanding about their ecology, growth-promoting traits, mechanisms of action, and their application for plant growth stimulation has key importance for maximum utilization of this naturally occurring microbial population [12]. Microbial diversity in soil ecosystem exceeds more than eukaryotic organisms. One gram of soil may harbor up to 10 billion microorganisms of possible thousands of different species [13]. Several implications for the management of plant diseases are derived from knowledge of the basis of the specificity of plant-microbial interactions. New biotechnological products are currently being developed based on stimulation of the plant defense response, and on the use of plant-beneficial microbes for biological control of plant diseases (biopesticides) and for plant growth promotion (biofertilizers) as well as biocontrol.
There for the study of this important plant growth promoting microorganisms are possible through extensive exploration from different plant-microbes association ecology. This study aiming to search rhizospher microbs associated with different grass family at different ecology to know its fungal profile in later to use as biofertilizer, biocontrol, biopesticide for agricultural and bioremediation application. The result revealed that Fellomyces fuzhounsis, Schizoblastosporon starkeyi-henricii, Cryptococcus luteolus, Rhodotrula aurantica A, Cryptococcus terreus A, Trichosporon beigelii B, Rhodotrula aurantica B, Cryptococcus albidus var aerius, yeasts were fully identified using Biolog identification system from Poaceae and Cyperaceae family grass in North and East Shewa, Ethiopia (Table 2). The Cryptococcus genera were dominant in this study and Trichosporon beigelii B was the dominant species identified from different rhizoshper soil of grass family. This is also reported by [14-17]. The basidiomycetous Cryptococcus aerius, Cryptococcus laurentii, Cryptococcus terreus, Cryptococcus terricola, Cryptococcus podzolicus, G. (Trichosporon) pullulans and the ascomycetous B. (Williopsis) californica, B. pratensis, Sc. (Debaryomyces) occidentalis, L. (Williopsis) saturnus, Schizoblastosporon starkeyi-henricii were yeast identified from most soil.
The yeasts, i.e. Cryptococcus tephrensis, Cryptococcus victoriae, Rhodotorula glutinis and Rhodosporidium babjevae. Although these yeasts are found regularly in soils, they are typically associated with the phylloplane and enter the soil profile with plant material [18]. There are different rhizospher fungi were identified from different grass family . Mostly PGPR belonging to some important genera include Serratia, Bacillus, Pseudomonas, Burkholderia, Enterobacter, Erwinia, Klebsiella, Beijerinckia, Flavobacterium and Gluconacetobacter are identified from rhizospher soil [19,20]. According to Acrophialophora fusispora, Botryotrichum piluliferum, Clonostachys rosea, Colletotrichum dematium, Fusarium oxysporum, Macrophomina phaseolina, Myrothecium roridum, Khuskia oryzae, Verticillium alboatrum fungal species were identified from the Rhizosphere and Rhizoplane Subfamily Panicoideae Grasses. According to [21]. The fungal isolates were exclusively found in the root regions of certain grass species. For example, Chloris barbata, Cynodon dactylon, Dactyloctenium aegyptium, Eleusine indica and Eragrostis unioloides harboured fungal species. Notable among them were Graphium penicilloides from Chloris barbata, Chaetomium indicum, Gilmaniella humicola, Trichoderma pseudokoningii from C. dactylon, Chaetomium bostrychodes from D. aegyptium, P. decumbense and T. viride from Eleusine indica, P. islandicum and T. koningii from E. unioloides. Rhizobacteria are isolated from teff rhizospher Pseudomonas fluorscens, Chryseomonas luteola, Pseudomonas putida, Bacillus licheniformis, Bacillus firmus, Burkholderia cepacia Stenotrophomonas maltophila,Brevibacillus brevis, Bacillus megaterium, Bacillus coagulans, Bacillus cereus, Bacillus pumilus Bacillus megaterium, Aeromonas hydrophila , Bacillus cereus, Bacillus subtilis [22]. The rise in the cost of chemical fertilizer, the lack of fertilizer industries in developing countries and the growing environmental issue and biodiversity loss using chemical fertilizer timely important concern using alternative ecofriendly bio fertilizer from rhizospher microbs investigation and evaluation for plant growth promoting character to increase yield and productivity of crop [23-28].
Conclusion
a. Understanding the Plant-microbe interactions and rhizospher microbial diversity has profound role in biotechnological application for plant productivity and ecosystem function.
b. Rhizospher soil yeast identified is mostly from genera of Cryptococcus, Trichosporon, Rhodotrula, Fellomyces, Schizoblastosporon in this study.
c. Growth and survival of a particular yeast species in soil may therefore not solely depend on the intrinsic abilities of the yeast but is the cumulative result of a number of interactions within the soil microbial community.
d. These results are promising in the field of bio-fertilizers. Application of yeasts increased the crop productivity. The mechanisms which could be involved include the bioavialbility of macro and micronutrients, production of growth hormones, and reduction of the phytopathogens’ growth.
In addition, they could improve the physical and chemical properties of soil that increase water holding capacity, prevent nutrient leaching and add more mineral nutrients to the soil. We assume that studying the production of yeasts’ secondary metaboilites and their bioactivities in the rhizospher holds exciting promise. We recommend further study dealing with the identification of the secondary metabolites of the yeasts and their bioactivities in the rhizosphere as well as their direct and indirect relationships with the plant growth and productivity./p>
References
- Soreng Robert J, Peterson Paul M, Romschenko Konstantin, Davidse Gerrit, Zuloaga Fernando O, et al. (2015) A worldwide phylogenetic classification of the Poaceae (Gramineae). Journal of Systematics and Evolution 53 (2): 117-137.
- Sarandón Ramiro (1988) Biología poblacional del gramon (Cynodon spp, Gramineae).
- Uren NC (2001) Types, amounts and possible functions of compounds released into the rhizosphere by soil-grown.
- Bowen G, Rovira A (1999) The rhizosphere and its management to improve plant growth. Advance in Agronomy 66: 1-102.
- Philippot L, Raaijmakers J M, Lemanceau P, van der Putten WH (2013) Going back to the roots: the microbial ecology of the rhizosphere. Nat Rev Microbiol 11(11): 789-799.
- Mendes L W, Kuramae EE, Navarrete AA, van Veen JA, Tsai SM (2014) Taxonomical and functional microbial community selection in soybean rhizosphere. ISME J 8(8): 1577-1587.
- Kevin Panke-Buisse, Angela C Poole, Julia K Goodrich, Ruth E Ley, Jenny Kao-Kniffin (2015) Selection on soil microbiomes reveals reproducible impacts on plant function. ISME Journal 9: 980-989.
- Kent AD, Triplett EW (2002) Microbial communities and their interactions in soil and rhizosphere ecosystems. Annu Rev Microbiol 56: 211-236.
- Bais HP, Weir TL, Perry LG, Gilroy S, Vivanco JM (2006) The role of root exudates in rhizosphere interactions with plants and other organisms. Ann Rev Plant Biol 57: 233-266.
- Biolog (1993) YT Microplate instruction for use. Biolog Inc.
- Hartmann A, Schmid M, van Tuinen D, Berg G (2009) Plant-driven selection of microbes. Plant and Soil 321(1-2): 235-257.
- Sajid Mahmood Nadeem, Muhammad Naveed, Zahir A Zahir, Hafi z Naeem Asghar (2013) Plant–Microbe Interactions for Sustainable Agriculture: Plant Microbe Symbiosis: Fundamentals and Advances 51-102
- Rosellό-Mora, Amann R (2001) The species concept prokaryotes. FEMS Microbiol Rev 25(1): 39-67.
- Bab’eva IP, Chernov IYu (1995) Geographical aspects of yeast ecology. Physiology and General Biology Reviews 9: 1-54.
- Lachance MA, Starmer WT (1998) Ecology and yeasts. In: Kurtzman CP, et al. (Eds). The Yeasts, a Taxonomic Study, (4th edn). Elsevier, Amsterdam, pp. 21-30.
- Vishniac HS (2006) A multivariate analysis of soil yeasts isolated from a latitudinal gradient. Microb Ecol 52(1): 90-103.
- Botha A (2011) The importance and ecology of yeasts in soil. Soil Biology & Biochemistry 43(1): 1-8.
- Fonseca A, Inacio J (2006) Phylloplane yeasts. In: Peter G, Rosa C (Eds), Biodiversity and Ecophysiology of Yeasts. SpringereVerlag, Berlin, pp. 263-301.
- Podile AR, Kishore GK (2006) Plant growth-promoting rhizobacteria. In: Gnanamanickam SS, (Eds.) Plant-Associated Bacteria. Springer; Netherlands. pp. 195-230.
- Dardanelli M S, Manyani H, Gonzalez-Barroso S, Rodriguez-Carvajal M A, Gil-Serrano A M, et al. (2010) Effect of the presence of the plant growth promoting rhizobacterium (PGPR) Chryseobacterium balustinum Aur9 and salt stress in the pattern of flavonoids exuded by soybean roots. Plant and Soil 328(1-2): 483-493
- Shivanna MB, Vasanthakumari MM (2011) Temporal and spatial variability of rhizosphere and rhizoplane fungal communities in grasses of the subfamily Chloridoideae in the Lakkavalli region of the Western Ghats in India. Mycosphere 2(3): 255-271.
- Delelegn Woyessa, Fassil Assefa (2011) Diversity and Plant Growth Promoting Properties of Rhizobacteria Isolated from tef (Eragrostis tef) Ethiop J Educ & Sc 6(2).
- Ahemad M, Kibret M (2014) Mechanisms and applications of plant growth promoting rhizobacteria: current perspective. Journal of King Saud University- Science 26(1): 1-20.
- Barnhart, John Hendley (1895) Family Nomenclature-Bulletin of the Torrey Botanical Club. Lancaster, PA, USA 22(1): 1-24.
- Luo SL, Xu TY, Chen L, Chen JL, Rao C, et al. (2012) Endophyte-assisted promotion of biomass production and metal-uptake of energy crop sweet sorghum by plant-growth promoting endophyte Bacillus sp. SLS18. Appl Microbiol Biotechnol 93(4): 1745-1753.
- Vasanthakumari MM, Shivanna MB (2011) Fungal Assemblages in the Rhizosphere and Rhizoplane of Grasses of the Subfamily Panicoideae in the Lakkavalli Region of Karnataka, India. Microbes Environ 26(3): 228-236.
- Soreng RJ, Peterson PM, Romaschenko K, Davidse G, Teisher JK, et al. (2017) A worldwide phylogenetic classification of the Poaceae (Gramineae) II. An update and a comparison of two 2015 classifications. Journal of Systematics and Evolution 55(4): 259-290.
- Weyens N van der Lelie D, Taghavi S, Newman L, Vangronsveld J (2009) Exploiting plant–microbe partnerships to improve biomass production and remediation. Trends Biotechnol 10: 591-598.






























