Occurrence of Vancomycin-Resistant Staphylococcus aureus (Vrsa) In Clinical and Community Isolates Within the University of Port Harcourt
Otobo UN*, Wala PG and Agbagwa OE
Department of Microbiology, Faculty of Science University of Port Harcourt, Nigeria
Submission: September 06, 2018; Published: September 25, 2018
*Corresponding author: Otobo UN, Department of Microbiology, Faculty of Science University of Port Harcourt, Rivers State Nigeria. P.M.B. 5323 East- West Road Choba, Nigeria; Email: obakpororo.agbagwa@uniport.edu.ng ejiroagbagwa@yahoo.com
How to cite this article: Otobo U, Wala P, Agbagwa O. Occurrence of Vancomycin-Resistant Staphylococcus aureus (Vrsa) In Clinical and Community Isolates Within the University of Port Harcourt. Adv Biotech & Micro. 2018; 11(4): 555816. DOI:10.19080/AIBM.2018.11.555816
Abstract
Staphylococcus is the major cause of infectious diseases and they have the ability to be resistant to various antibiotics. Vancomycin continues to be an important antimicrobial agent for infections caused by S. aureus that are resistant to oxacillin and other antimicrobial agents. The increased incidence of MRSA has led to more frequent use of vancomycin, which is mostly relied upon as last resorts for treating MRSA infections. As a consequence, selective pressure was established that led to the emergence of strains of Staphylococcus with decreased susceptibility to vancomycin. The aim of the study is to determine the prevalence of Vancomycin Resistant S. aureus from clinical and community samples within the University Port Harcourt.
A total of one hundred and ten samples were used for the study, fifty from clinical and sixty from the food vendors within the community. The samples were evaluated using Mannitol Salt Agar and Brain Heart Infusion Agar. Isolates were identified using colonial morphology, color indication, Gram reaction and biochemical tests. Results obtained from the study showed that out of the fifty clinical samples, thirty had significant growth out of which twenty (20) were S. aureus, from the community only 20 isolates (33.3%) were positive. Of the forty isolates, none were confirmed to be VRSA. This study shows that the occurrence of VRSA from the samples was 0%, this confirms that vancomycin is still a prevalent antibiotic against S. aureus.
Keywords: Antibiotics; Resistant; Staphylococcus aureus; Vancomycin
Abbrevations:AMP: Ampicillin; AUG: Amoxycillin/Clavulanate (Augumentin); OFL: Ofloxacin; RIF: Rifampicin; CRX: Cefuroxime; IMP: Imipenem; ERY: Erythromycin; GEN: Gentamicin
Introduction
The decrease in the effectiveness of antibiotics in treating common infections has become a world-wide burden. In recent years there has been the presence of untreated strains of carbapenem resistant enterobacteriacae [1]. The increase in resistance is facilitated by interspecies gene transmission, lack of good sanitation and hygiene in the hospital and community thereby causing an increase in the frequency of global trade and disease transmission [2]. Staphylococcus aureus is the most common cause of Staphyloccocal infection and they are responsible for different diseases such as skin infection, wound infections and toxin mediated diseases [3]. S. aureus is a major cause of infection in either hospitals or within communities across the world, this has made S. aureus develop resistance to commonly prescribed antimicrobial agents. They have the ability to acquire resistance to new antimicrobial agents. The first serious emergence of antibiotic resistant S. aureus occurred as Methicillin-Resistant Staphylococcus aureus (MRSA). The strain expressed a modified penicillin-binding protein encoded by MecA gene. MRSA strains are currently a very significant health care problem.
Due to widespread prevalence of MRSA, the empiric therapy for MRSA was changed to vancomycin. This resulted in the emergence of vancomycin intermediate and vancomycin resistant S. aureus [4]. Lately, there have been cases of recently emerging VISA and VRSA because of reduced susceptibility to vancomycin [5]. The emergence of drug resistant organisms’ especially vancomycin-intermediate Staphylococcus aureus (VISA) and vancomycin-resistant S. aureus (VRSA) has become a major concern globally in both developing and developed countries. Studies have shown that most S. aureus isolates are usually resistant to methicillin, the glycopeptide vancomycin has been widely used as one of first proper alternative choices for the treatment of methicillin-resistant S. aureus (MRSA) [6]. People that are at high risk for S. aureus infection are the very young children, old people, immune-deficient patients and patients that have gone through surgery [6, 7]. S. aureus are responsible for most hospital-acquired infections, they infect immunocompromised humans, the infection remains localized at the point of entry by the host defenses. S. aureus have become a world- wide issue based on their drug resistant pattern and their ability to colonize [7].
In developing countries such as Nigeria, street (food) vendors have formed an integral part of the food supply chain. The main concern is the sanitary conditions of these street food vendors based on the fact Staphylococcal food poisoning can occur in which a toxic substance produced by the bacteria is ingested with the food [8]. The study was carried out to detect the presence of vancomycin -resistant S. aureus from clinical and community sources.
Materials and Methods
Sample Collection
A total of one hundred and ten (110) samples were used for the study. The clinical isolates were isolated from swabs from various sources in the hospital. The remaining sixty swabs were used to swab the hands of food vendors within the University of Port Harcourt. The characteristics of the patients were unknown. The swabs were taken to the Medical Microbiology laboratory of the University of Port Harcourt and streaked unto sterile Mannitol Salt Agar (MSA) plates. The plates were incubated at 37o C for 24 hours after which the colonial characteristics were observed.
Identification of S.aureus
Isolates that grew on the mannitol salt agar were purified by sub-culturing onto nutrient agar plates for 24 hours. The isolates were identified by carrying out standard microbiological test such Gram staining, coagulase and catalase test to identify S. aureus [9]. For microscopic observation, a pure colony was selected and subjected to Gram staining. The shape, arrangement, and Gram reactions of the isolates were observed under a light microscope (at a magnification of 100x). Confirmatory biochemical tests including catalase and triple sugar iron agar tests were performed to identify suspected S. aureus following standard microbiology protocols [10].
Antibiotic sensitivity test by Kirby-Bauer Disc Diffusion
Antibiotic sensitive test was carried out using Kirby-Bauer disc diffusion method. A sterile inoculating loop was used to pick a colony of the isolate and transferred in each of the 5ml normal saline and homogenized properly until it becomes slightly turbid. Turbidity of the suspension was cross matched with the turbidity standard (0.5 MacFarland turbidity standards). A sterile cotton swabs was dipped into the bacterial test suspension and was used to evenly inoculate the entire surface of the Muller Hinton Agar Plate (MHA). Gram positive antibiotic disc were placed on the surface and press gently using sterile forceps. The plates were incubated inverted for 24 hours at 35 oC. Antibiotic resistance and susceptibility was determined after 24 hours by measuring the zone of inhibition in millimeter. The following antibiotics were commercially impregnated in the disc; AMP: Ampicillin, AUG: Amoxycillin/Clavulanate (Augumentin), OFL: Ofloxacin. RIF: Rifampicin, CRX: Cefuroxime, IMP: Imipenem, ERY: Erythromycin and GEN: Gentamicin
Detection of Vancomycin Resistant S. aureus (VRSA)
VRSA was identified using the disc diffusion method. The isolates that were positive cocci, catalase positive and coagulase human plasma were considered S.aureus. All the confirmed S.aureus was tested for VRSA using Oxoid antimicrobial susceptibility vancomycin disc (30 μg/disc) by Kirby Bauer disc diffusion method. Muller Hinton agar plates were inoculated with the bacterial suspension which was adjusted to 0.5 McFarland standards. Sterile forceps were used to place the vancomycin disc on the agar plates. The plates were incubated at 37 oC for 24 hours. Zone diameter of bacterial growth inhibition surrounding the disc was measured and compared with CLSI standard which states that vancomycin is sensitive when zone diameter nearest to whole millimeter is ≥15 [11, 12].
Results and Discussion
Characterization of Staphylococcus aureus isolation
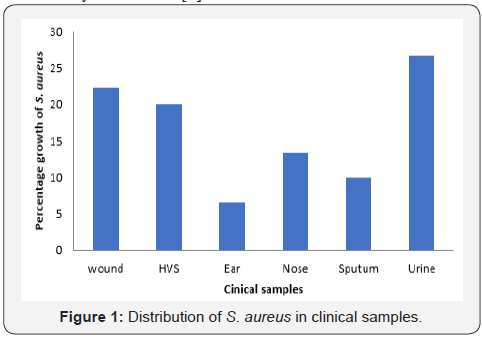
Out of the 50(110%) swabs cultured, it was observed that 30(60%) yielded growth while 20(40%) yielded no growth for the clinical samples. Percentage of the 50(100%) samples included; wound (20%), HVS (14%), Ear (12%), nose (20%), sputum (16%), and urine (18%). Isolates identified from the palms of food vendors, twenty (33.3%) were confirmed to be S. aureus. These isolates were identified by their characteristic, arrangement and biochemical test. Cultures that yielded colonies of sizes ranging from 1 mm to 3.0mm with varying shades of color ranging from pale yellow to yellow and pink. These colonies were also observed to be smooth and glistering. Based on these, 20(66.7%) isolates were confirmed characteristic S. aureus on MRA while 10(33.3%) isolates were non aureus. Percentage of the 30 isolated included; wound (22.3%), HVS (20%), ear (6.6%), nose (13.4%), sputum (10%) and urine (26.7%).
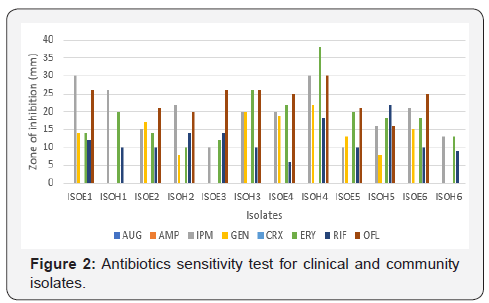
The antibiotics sensitivity result for the clinical and community isolates all the isolates were 100% resistant to ampicillin, cefuroxime and amoxicillin/clavulanate (Augmentin) but most were sensitive to ofloxacin. and showed varying resistance to rifampicin, gentamicin, imipenem and erythromycin as shown in Table 1 and Figure 2.
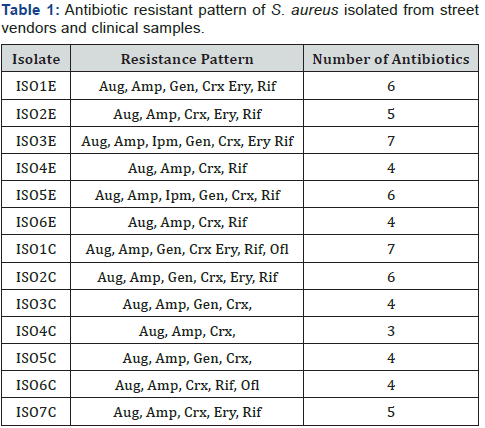
Most of the isolates were resistant to more than three antibiotics. All the isolates were resistant to Augumentin and Ampicillin, more than 50% of the isolates were also resistant to Gentamicin. ISO3E from non-clinical sources and ISO1C from clinical sources were resistant to 7 antibiotics. ISO2C and ISO5E were resistant to 6 antibiotics as shown in Table 1. Staphylococcus aureus as a ubiquitous bacterium is a leading cause of superficial infection for decades. The medical importance of S. aureus cannot be overemphasized. Antibiotics resistance to S. aureus, has been reported to be on the increase globally [13]. Since humans are colonized with Staphylococcus aureus and the use of antimicrobial agents is very common. The spread of S. aureus (including MRSA) generally is through human-to-human contact or from contaminated environment to humans so emphasis on basic hand washing techniques will therefore be effective in preventing its transmission. Over the past decade’s Staphylococcal infections have been prevalent in various communities and in healthcare institutions in both the developed and the developing countries. Numerous antibiotics have been introduced in combating Staphylococcal infections. Over time, Staphylococci developed resistance to most antibiotics by different mechanisms. The beta-lactamase enzymes hydrolyze the beta-lactam ring of Penicillin and thereby making these antibiotics ineffective to S. aureus [14].
A lot of concern is placed on S. aureus because it can adapt to different environmental conditions and its intrinsic virulence that enables it to cause various types of infection that can threaten human life [14]. They can cause infection in hospitals and communities, they show variability in susceptibility towards antibiotics, this has not reduced even with the existence of different antibiotics [15]. In our study the susceptibility of S. aureus isolates to vancomycin was 100%, none was resistant indicating that the prevalence rate of VRSA is 0%. The findings of this study could be attributed to the abuse of this antibiotics by patients and street vendors is therefore limited and could be the reason why vancomycin was active against the S. aureus isolates. The presence of multi-drug resistant strains of S. aureus observed among the isolates can be due to antibiotic mixture that could have arisen from self-medication in suspected bacterial infections [16].
Staphylococcus is one of the most common clinical isolates that are responsible for most suppurated infections, they can also acquire resistance by more than one resistance mechanism from one location to the other. This is also applicable to the resistance pattern which depends on the local antibiotic policy applicable in that particular location. Other factors that could influence the prevalence of S. aureus include the infection control activities, the time of the study, the number of cases and the biological characteristics of the strains involved in the study [17]. The results obtained from this study showed that staphylococcus aureus is a common pathogen of humans’ observation is similar to those observed by [18] and is also found in the environment. A total of 1789 S. aureus was isolated from a hospital in Tehran. From all the isolates four were confirmed to be VRSA and two were VISA, this shows that continued surveillance of VRSA in other to prevent the transmission of these isolates [19].
A study carried out by [20] in a Tertiary hospital in Nigeria showed that the susceptibility of S. aureus strains to vancomycin was 76.6% (272) out of 355 S. aureus isolates that were tested. Our study is also in agreement with the findings of [21] where out of 335 S. aureus clinical isolates showed that 272(76%) were susceptible. When they were subjected to statistical analysis it showed that the significant rate was non- significant. The result when subjected to statistical analysis the result was not significant (P˂0.05) which is in conformity with our findings in the present study. Another study carried out in Karachi, Pakistan where a total of 850 clinical isolated was tested for VRSA, none of the isolates were found to be resistant to vancomycin which is also in conformity with the present study. S. aureus developed resistance against vancomycin due to the presence of Van A gene that codes for an altered target such that the binding of vancomycin to the target is significantly reduced and thus cannot carry out its normal function of inhibiting bacterial cell wall synthesis. Research has shown that the source of the Van A gene isolated in VRSA suspects to have come from co-infection with vancomycin resistant enterococcus (VRE) [22]. In recent years the number of cases of VISA and VRSA reported has remained relatively small indicating that that the epidemiology and the risk factors associated with infection with these organisms is not totally known.
Conclusion
Good hygiene an sanitary habit should be emphasized on the reduce carriage of S. aureus and vancomycin resistant Staphylococcus aureus, since their presence could act as a source of food poisoning and also predispose an individual to other endogenous infections. Early detection of isolates resistant to newer antibiotics is important as this will allow appropriate treatment of affected patients. Further studies are needed to evaluate the relevance of VRSA in patients with clinical failure to vancomycin and in order to conduct such studies, a means of accurately identifying these strains is essential and attempts should be made to keep the prevalence of such strains low
References
- Sievert DM, Boulton ML, Stolman G, Johnson D, Stobierski MG, et al. (2013) S. aureus resistant to vancomycin. MMWR Morb Mortal Wkly REP 51: 565-567.
- Abraham EP, Chain E (2008) An enzyme from bacteria able to destroy penicillin. RevInfect Dis. 10: 677-678.
- Azeez-Akande O, Utosala SJ, Epoke J (2008) Distribution and antibiotic susceptibility pattern of Methicillin Resistant Staphylococcus aureus isolates in University Teaching Hospital in Nigeria 11(4): 142-147.
- Saderi H, Owlia P, Shahrbanooie R (2005) Vancomycin resistance among clinical isolates of Staphylococcus aureus. Archive Iranian Med, 8(2): 100–103.
- Chamber HF (2001) The changing of Staphylococcus aureus? Emerg. Infect. Dis 7(2): 178-182.
- Katzung B G (2009) Basic and clinical pharmacology. New York, NY: Hill.
- Katzung B G (2007) Basic and clinical pharmacology (10th edn.) New York, NY: Hill.
- Tuo L, Cui L, Ma X, Sato K (2005) Cell wall thickening: A common feature of vancomycin resistant Staphylococcus aureus. J Clin Microbiol 41: 5-14.
- Cheesbrough M (2000) Medical laboratory manual for tropical countries. Microbiology. Cambridge University Press 2: 62-70.
- Cappuccino JG, Sherman N (1996) Microbiology A laboratory manual. (5th edn). Menlo Park, (CA): The Benjamin/Cummings Publishing Co, Inc.
- Kaiser ML, Thompson DJ, Malinoski D, Lane C, Cinat ME (2011) Epidemiology and risk factors for hospital-acquired methicillinresistant Staphylococcus aureus among burn patients. J Burn Care Res 32(3): 429–434.
- Schweizer M, Ward M, Cobb S, McDanel J, Leder L, Wibbenmeyer L (2012) The epidemiology of methicillin-resistant Staphylococcus aureus on a burn trauma unit. Infect Control Hosp Epidemiol 33(11): 1118–1125.
- Shittu AA, Johnson PB (2006) Triends in the incidence of methicillin resitant S. aureus infection in children’s hospital in the United States. Clin Infect Dis 49(1): 65.
- Obasi EA, Kobayashi SD, Musser JM, DeLeo FR (2007) Genomic analysis of the emergence of VRSA, MBio 3(4): 170-12.
- Newman NR, Polk R, Fox C, Mahoney A (2006) Measurement of adult antibacterial drug use in 130 US hospitals: comparison of defined daily dose and days of therapy. Clin Infec Dis 44 (5): 664-670.
- Mallick SK, Basak S (2010) MRSA too many hurdles to overcome: a study from Central India. Tropical Doctor 40(2): 108-110.
- Uemura E, Kakinohana S, Higa N, Toma C, Nakasone N (2004) Comparative characterization of Staphylococcus aureus from throat and nose of healthy volunteers. Jpn. J. Infect. Dis 57: 21 -24.
- Shekarabi M, Hajlkhari B, Salimi Chirani A, Fazeli M, Goudaezi M (2017) Vancomycin-resistant Staphylococcus aureus strains isolated from clinical samples: A three-year study in Tehran, Iran. PLOSONE 12(8): e0183607.
- Wanyne PA (2007) Methods of antimicrobial susceptibility tests for bacteria that grow aerobically. (5th edn). Approved standard, M7-A5. National Committee for Clinical Laboratory Standards.
- Aflo M, Ugah U, Okoro N (2013) Epidemiology of vancomycin resistant Staphylococcus aureus among clinical isolates in a tertiary hospital in Abakaliki, Nigeria. American journal of epidemiology and infectious diseases 1(3): 24 -26.
- Whitner CJ, Park SY, Browne FA, Parent LJ, Julian K, et al. (2004) Vancomycin-resistant Staphylococcus aureus in the absence of vancomycin exposure. Clinical Infectious Diseases 38(8): 1049 -1055.
- Laxminarayan RE, Derlot J, Duval J, Courvalin P (2012) Plasmidmediated resistance to vancomycin and teicoplanin in Enterococcus faecium. N Engl J Med 19(3): 157-161.






























