Microbial Diversity Assessment by PCR-DGGE Analysis in National Sanctuary of Ampay in Perú
Raquel A Ccoscco1, Victor H Sarmiento2 and Gretty K Villena1*
1 Laboratorio de Micología y Biotecnología, Universidad Nacional Agraria La Molina, Perú
2Universidad Nacional Micaela Bastidas de Apurímac, Perú
Submission:August 11, 2018; Published: September 20, 2018
*Corresponding author: Gretty K Villena, Laboratorio de Micología y Biotecnología, Universidad Nacional Agraria La Molina, Av. La Molina s/n, Lima 12, Perú, Tel: +51-1-6147800, Ext. 463; Email: gkvch@lamolina.edu.pe
How to cite this article: Gretty K Villena, Raquel A Ccoscco, Victor H Sarmiento. Microbial Diversity Assessment by PCR-DGGE analysis in National Sanctuary of Ampay in Perú. Adv Biotech & Micro. 2018; 11(3): 555811. DOI:10.19080/AIBM.2018.11.555811
Abstract
Microbial diversity analysis by means of PCR- DGGE fingerprinting was conducted for water and sediment samples of the National Sanctuary of Ampay in Perú. This wildlife conservation area is not disturbed by human activity and this study corresponds to the first report of microbial diversity. PCR DGGE was able to detect culturable and non-culturable bacteria and fungi. The Predominant bacterial genus in water and sediment was Pseudomonas. Fungal community was more diverse. Aspergillus species were mainly distributed in water. Sediment samples contain different fungi genera, including a non-culturable Pleosporales specie. Because the bacterial and fungal diversity found, the Sanctuary of Ampay in Peru is a very valuable natural environment for bioprospecting of new genes of enzymes and natural compounds for different biotechnological applications.
Keywords: Genes; DGGE-PCR analysis; Water samples; Enzymes; Biotechnological applications;
Introduction
Culture based methods to evaluated microbial diversity are usually unsuccessful because only 0.1-1% to 1- 10% of microbial species can growth in vitro and different microbiological and molecular strategies have been considered to improve microbial cultivation [1,2].
In the last years, culture-independent techniques had become the main tools for evaluation of taxonomic and functional structures of microbial communities. Either 16S rRNA amplicon-based metagenome sequencing or shotgun metagenome sequencing is largely used for the assessment of the microbial diversity. Recently, comparison of two next-generation sequencing techniques, amplicon and shotgun, in Brazilian river food plain systems showed that less than 50% of phyla identified via amplicon sequencing were recovered from shotgun sequencing [3]. Amplicon based metagenome sequencing are commonly performed by analyzing the prokaryotic 16S ribosomal RNA gene (16S rRNA), which is approximately 1,500 bp long and contains nine variable regions interspersed between conserved regions. Variable regions of 16S rRNA are frequently used in phylogenetic classification such as genus or species in diverse microbial populations. Improving on the amplicon sequencing techniques remain as challenge [4,5].
Between traditional and high accurate techniques for microbial diversity assessment, Denaturing Gradient Gel Electrophoresis, (DGGE) is an inexpensive fingerprinting technique and could be employed for several applications, including analysis of complex communities, monitoring of population shifts, and sequence heterogeneities. Through DGGE it is possible to obtain taxonomic information because bands can be excised, re-amplified and sequenced [6]. DGGE is still the chosen technique to assess microbial community composition in different natural or artificial ecosystems and foods [7,8]. Natural ecosystems comprise several microbial habitats including thermal water, water ponds, microbial mats and sediments. Metagenomic study of this habitats produced huge information, providing holistic images of the microbial communities. Peru has several conservancy areas distributed across marine, coast, andean and jungle ecosystems but there is not any report of microbial diversity by metagenomic approach.
National Park of Ampay Santuary is placed in Apurimac, in the south Andes (13° 34’44” S, 72° 53’ 24” W) and has forest, several water ponds and two hot springs. This place is very attractive for biodiversity studies, since is considered as conservation area. Thus, natural environments are not disturbed by human activity and biodiversity analysis has the advantage to reflect natural microbial community structure.
Despite of the valuable ecological information obtained by biodiversity analysis, the deep knowledge of structure of microbial communities allows several biotechnological applications such as discovery of new genes encoding for enzymes, secondary metabolites and others.
Material and Methods
Sample collection
Samples were collected from hillside water pond and surrounding sediment of “Carbon cañana”, located in National Sanctuary of Apurímac, Perú (13° 33’26”S, 72° 51’ 43” W and; 3255m high). Environmental and water temperatures were 14 °C and 20 °C respectively.
DNA extraction
To extract the DNA from water samples, one liter of water was vacuum filtered with nylon membrane (0.2μm pore size and 47mm diameter). Then, the membrane was cut in small pieces and put them into one 2ml tube. On the other hand, the sediment sample was transferred into a 50 ml tube, centrifuged (5000 rpm, 15 min), and desiccated. Both, water and desiccated sediment samples were stored at -80°C until their DNAs were extracted.
The DNA from water sample was extracted according to Ludeña method [9], with some modifications as follows: cells were removed from filter with a vortex in 500μL of a TES buffer (10mM Tris, 1mM EDTA, pH 8, and 25% (w/v) of sucrose) for 15 sec. Then, the liquid phase was recovered into other 2ml microtube and treated with 60μl of lyzozyme (5mg/ml) and 50 μl of 0.25M EDTA at 37 °C for 15 min. RNA was eliminated by adding 175μl of H2O, 7.5μl 10% (w/v) SDS, and 10μl of 10mg/ml RNAase to the mix, homogenized by vortex 2 sec and incubated at 37 °C for 30 min). To complete the lysis process 10 μl of 10mg/ ml proteinase K was added and incubated at 37 °C for 30 min. The proteins and cell debris were precipitated by adding 90μl of TE (10 mM Tris base, 1mM EDTA, pH 8), 120μl of 5M NaCl (5M); and the mix was homogenized and incubated at 65 °C for 25 min. In addition, 8M Potassium acetate solution (75μl) was used to concentrate the sample, by incubation at 4 °C for 15 min and centrifugation at 13500 rpm for 5min. The supernatant was collected, and centrifuged at 13500 rpm for 5 min, 3 times. The DNA was precipitated using 0.6V of isopropanol, incubating at -20 °C for 30min. After centrifugation at 13500rpm for 15min, the DNA pellet was washed with 70% (v/v) ethanol, and centrifuged at 13500 rpm for 15 min. Finally, The DNA was dissolved in 25μl of free-nucleases water, and stored at 20 °C.
DNA extraction method from sediment sample was adapted from Lai, et al. [10], Zhou, et al. [11] & Murray, et al. [12] 300mg of sediment were transferred to 2ml tube, and 810μl of Extraction buffer (100mM Tris-HCl, 100mM EDTA, 100 mM sodium phosphate, 1.5M NaCl, pH 8), 90μl of 20% (w/v) SDS, and 117μl of 10%(v/v) CTAB were added. The mix was homogenized by inversion for 1 min and incubated at 68 °C for 1 hour, every 15min the sample was homogenized by inversion, and centrifuged at 13500 rpm for 5min, and the upper solution was transferred to a clean tube. This step was done for two samples by separate. The purification step was performed by adding 1 part of chloroform to each tube. Then, they were homogenized by inversion and centrifuged at 13500rpm for 10 min. The upper solution was transferred to a new 2ml tube. RNA digestion was performing by adding 15μl of 10 mg/ ml RNase to each tube and incubated at 37°C for 30min. The proteins were digested by adding 10μl of 20mg/ml proteinase K.
After that, the solution was homogenized by inversion and incubated at 37 °C for 30min. Precipitation of the CTAB-DNA complex was done by adding 250μl of 2X CTAB (1.4M NaCl, 0.1 M EDTA y 2%(w/v) CTAB) to each tube, and they were incubated at 68 °C for 15min. The CTAB-DNA complex was precipitated by adding 2.2 volumes of precipitation solution (0.5% (w/v) CTAB and 0.04M NaCl) and incubating for 1 hour in ice. Samples were centrifuged at 3000rpm for 5min, and the upper solution was discarded. 300μl of 1.2M NaCl was added to each tube to dissociate the CTAB-DNA complex. The solution of each tube was mixed in only one. Finally, The DNA was purified by adding 1 volume of chloroform and centrifuged at 13500rpm for 15min. The supernatant was recovered in a 1.5ml tube. The DNA was precipitated in 0.6 volume of isopropanol and incubated at -20 °C for 1 h. After the DNA precipitation at 13500 rpm for 15min, the pellet was washed with 70% (v/v) ethanol, solubilized in 20μl free-nuclease water, and stored at -20 °C.
PCR amplification of 16 and 18 S rRNA genes
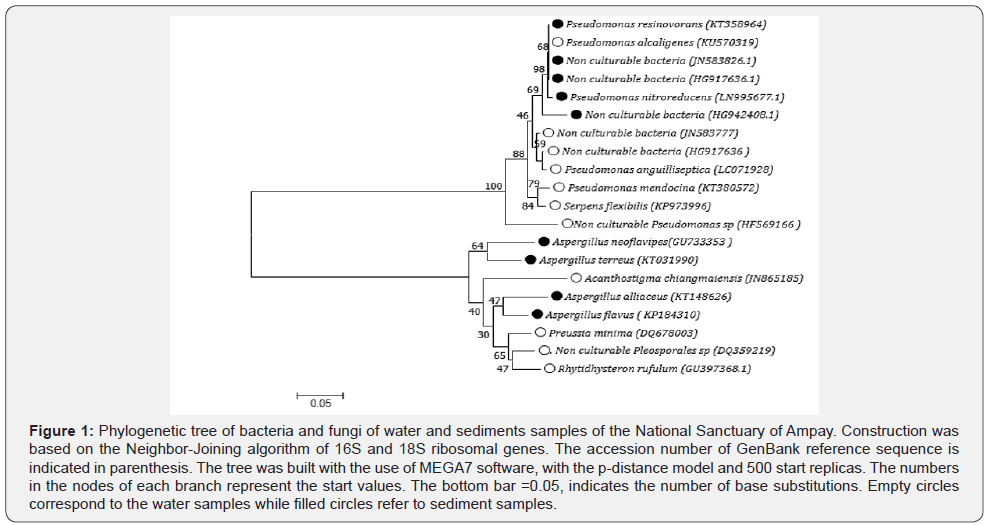
16S and 18S rRNAs were amplified by PCR with primers showed in Table 1. Temperature gradient with different temperature intervals was performed to find out the optimal temperature of annealing for Bacteria and Archaea (from 48 °C to 65 °C) and Eukarya (fungi) (from 40 °C to 60 °C).
*GCclamp (CGCCCGCCGCGCCCCGCGCCCGGCCCGCCGCCCCCGCCCC) [16].
PCR mix (final concentration) for 50 μl of volume reaction contained: PCR buffer 1 x, MgCl2 (2.5mM), dNTP’s (0.5mM for each one), primers F and R (0.2mM for each one), Go Taq Promega (1.875U/50μl), metagenomic DNA (100 ng/50μl).
PCR thermal cycles include: 1 cycle of denaturation at 94 °C, 35 cycles of 1 min denaturation at 94 °C, 1min for annealing (optimal temperature varies for each primer set) and 2min of extension at 72 °C. The final extension was performed at 72 °C for 10 min.
Denaturing Gradient Gel Electrophoresis (DGGE)
DGGE analyses was performed according to Green et al. [14] using a Cleaver scientific® equipment. The PCR products were loaded on 7% (w/v) polyacrylamide gels and run with 1X TAE (40mM Tris acetate, sodium acetate 20mM, and 1mM EDTA pH 8). The polyacrylamide gels were made with denaturing range from 36% to 65% (Bacteria and Archaea) and from 25% to 55% (Fungi). A 100% UF solution was defined as 40% (v/v) formamide and 7.0 M urea. The DGGE was conducted at 60 °C 160V for 3.5 h (for 16S rDNA amplicons), and at 56 °C, 50V for 16h (for 18S rDNA amplicons). After the electrophoresis, the gel was stained with Gel Red™ for 30min. The stained gel was immediately photographed (ChemiDoc XRS-BioRad).
DGGE PCR
DNA bands from the DGGE gel were cut and put in 600 μl tubes, then, submerged with 30 μl of ultrapure free-nuclease water. The DNA was allowed to passively diffuse into water at 37 °C for 30 min, and then at 4 °C overnight [15]. Aqueous phase for each band was recovered in 1.5 ml tube and stored at 4 °C 5μl of aqueous phase was used as template for PCR reaction with conventional rRNA gene primers without clamp (Table 1), following the same conditions of metagenomic DNA PCR.
Sequence analysis
DGGE-PCR amplicons were sequenced in MACROGEN (South Korea). The sequences obtained were analyzed with BLAST program of the National Centre for Biotechnology Information (NCBI) Phylogenetic tree was constructed by Neighbor-joining method from MEGA7 software.
Results and Discussion
DGGE analysis, a molecular culture independent method for assessment of microbial diversity was performed with hill inside water and sediment samples from National Sanctuary of Ampay in Perú.
This is the first report of microbial diversity in this wildlife conservation area, although previous reports on faunal inventory of Satyrinae butterflies and diversity of arboreal biomass were published [16,17].
To evaluate the microbial diversity of water and sediment samples, some protocols for metagenomic DNA extraction were standardized to obtain both, better yield and quality of DNA. DNA recovery was as tenfold (10 x) higher from sediment samples compared with water samples. Yields for DNA extraction were 2μg per liter of water and 6μg per gram of sediment (dry weight). As expected, DNA quality from water samples was better; since sediment and soils commonly have humic acids which co precipitate with nucleic acids [18,19]. Also, the average size of the metagenomic DNA obtained from both types of samples was up to 18 Kbp, which was comparable with other optimized methods [20].
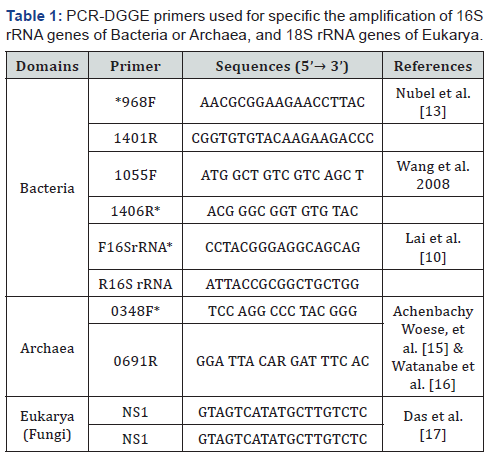
For PCR-DGGE, reference primers were used for amplification of a fragment of the 16S rRNA genes, 23S rRNA and 18S rRNA belonging to bacteria, archaea and eukarya (fungi). For each pair of primers, a gradient temperature PCR was done in order to obtain the optimal annealing temperatures (48 °C for GC- 968F/1401R and 1055F/ 1406R-GC, 55 °C for F16SrRNA-GC/ R16S rRNA and 55 °C, NS1/ GC-Fung). No amplification was obtained for the 16S rRNA target sequences of archaea, since these sequences belong mainly to methanogenic archaea [16]. Although r16SRNA target sequences for Archaea were limited to few phyla, more sequences have been validated in the past years [21-24]. Bacterial and fungal diversity found in water and sediment samples of Ampay are showed in Figure 1. The most prevalent bacterial genus in water and soil was Pseudomonas, which is widely distributed in the environment [25]. Species of Pseudomonas comprised culturable and non-culturable strains. Non culturable bacteria refers to microorganisms with low viability because any stress factor in their environment limits their growth on the routine bacteriological media but are still able to show metabolic activity [26].
Common species found in Ampay include P. resinovorans and P. nitroreducens in sediment and P. alcaligenes in water respectively. P. resinovorans could be a rare human pathogen but also has biotechnological importance as lipase producer as well as P. resinovorans for the vanillin production and carbazol degradation [27-30]. Another detected bacteria in sediment, Pseudomonas nitroreducens, has a great metabolic capability for the catabolism of aromatic compounds [31].
Other species in water samples include Serpens flexibilis, recently renamed as Pseudomonas flexibilis [32] and P. mendocina an inhabitant of soil and water environments and previously reported also as an opportunistic human pathogen [33]. Interestedly, P.mendocina strains were recently reported as cellulase and medium-chain-length polyhydroxyalkanoate producers [34,35]. Finally, Pseudomonas anguilliseptica has been reported as a potential fish pathogen [36]. Because nonhuman activity has taken place in hill inside water in Ampay, strain identification should be confirmed
With respect with the fungal diversity, the prevalent genus in sediment samples was Aspergillus. Even fungi are predominantly saprophytic, some of them are also considered as opportunistic species. Aspergillus flavus is a crop contaminant and aflatoxin producer [37], Aspergillus flavipes was recently studied for cytochalasans production and Aspergillus terreus which is the main producer of lovastatin, a cholesterol-lowering commercial drug [38,39].
Water samples showed more diverse genera of fungi. Those include: The new aquatic specie Acanthostigma chiangmaiensis [38], the endophytic specie Preussia minima which produces amylases and the Ascomycete, pan tropical specie, Rhytidhysteron rufulum [40,41]. Also, non-culturable species were detected, belonging to the Pleosporales, which comprises epiphytes,
Conclusion
According to the overall species and metabolic diversity found in water and sediment by PCR-DGGE analysis, the Sanctuary of Ampay, constitutes an environment with high potential for bioprospecting of new genes of enzymes and natural compounds with biotechnological applications. The use of culture independent methods, as DGGE, to recover DNA directly from the environment, avoids the manipulation of opportunistic or pathogenic strains, allowing a safe use of genes related to metabolic diversity in natural environments.
Acknowledgement
This work was supported by Programa Nacional de Innovación para la Competitividad y Productividad- Innóvate Perú, through the grant Nº 143-FINCyT-IB-2013.
This article is dedicated in memory of Dr. Marcel Gutiérrez- Correa, great mentor.
References
- Amann RL, Ludwig W, Schleifer Karl (1995) Phylogenetic identification and in situ detection of individual microbial cell without cultivation. Microbiol Rev 59(1): 43-69.
- Prakash O, Shouche Y, Jangid K, Kostka JE (2013) Microbial cultivation and the role of microbial resource centers in the omics era. Appl Microbiol Biotechnol 97(1): 51-62.
- Tessler M. et al. (2017) Large-scale differences in microbial biodiversity discovery between 16S amplicon and shotgun sequencing. Scientific Reports 7: 6589.
- Barb JJ, Oler AJ, Kim HS, Chalmers N, Wallen GR, et al. (2016) Development of an Analysis Pipeline Characterizing Multiple Hypervariable Regions of 16S rRNA Using Mock Samples. PLoS One 11(2): e0148047.
- de Muinck EJ, Trosvik P, Gilfillan GD, Hov JR, Sundaram AYM (2017) A novel ultra-high-throughput 16S rRNA gene amplicon sequencing library preparation method for the Illumina HiSeq platform. Microbiome 5(1): 68.
- Duarte S, Cássio F, Pascoal C (2012) Denaturing gradient gel electrophoresis (DGGE) in microbial ecology - insights from freshwaters. In: Magdeldin S, (Eds,) Gel electrophoresis - principles and basics. Rijeka: In Tech, p. 173-196.
- Tapia Estella, Andrés Donoso-Bravo, Léa Cabrol, Maria Madalena Alves, Alcina Pereira, et al (2014). A methodology for a functional interpretation of DGGE with the help of mathematical modelling. Application in bio-hydrogen production. Water Science and Technology 69(3): 511-517.
- Benítez-Cabello A, Bautista-Gallego J, Garrido-Fernández A, Rantsiou K, Cocolin L, et al. (2016) RT-PCR–DGGE Analysis to Elucidate the Dominant Bacterial Species of Industrial Spanish-Style Green Table Olive Fermentations. Front Microbiol 7: 1291.
- Ludeña Hinojosa Y (2007) Cianobacterias en la Bahia de Mayagüez: Abundancia, distribución y su relación con las propiedades bio-ópticas. Tesis de maestría Universidad de Puerto Rico Recinto Universitario de Mayagüez.
- Lai X, Zeng X, Fang S, Huang Y, Cao L, et al. (2006) Denaturing gradient gel electrophoresis (DGGE) analysis of bacterial community composition in deep-sea sediments of the South China Sea. World Journal of Microbiology and Biotechnology 22(12): 1337-1345.
- Zhou J, Bruns MANN, Tiedje JM (1996) DNA recovery from soils of diverse composition. Appl Environ Microbiol 62(2): 316-322.
- Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8(19): 4321-4326.
- Nübel U, Engelen B, Felske A, Snaidr J, Wieshuber A, et al. (1996) Sequence heterogeneities of genes encoding 16S rRNAs in Paenibacillus polymyxa detected by temperature gradient gel electrophoresis. J Bacteriol 178(19): 5636-5643.
- Green SJ, Leigh MB, Neufeld JD (2009) Denaturing gradient gel electrophoresis (DGGE) for microbial community analysis. In: Timmis KN, (Eds,) Handbook of Hydrocarbon and Lipid Microbiology, Springer (Heidelberg, Germany) pp 4137-4158.
- Achenbach L, Woese C (1995) 16S and 23S rRNA-like primers. In: Sowers KR, Schreier HJ, (Eds.) Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, USA, pp. 521-523.
- Watanabe T, Asakawa S, Nakamura A, Nagaoka K, Kimura M (2004) DGGE method for analyzing 16S rDNA of methanogenicarchaeal community in paddy field soil. FEMS Microbiol Lett 232(2): 153-163.
- Das M, Royer TV, Leff LG (2006) Diversity of Fungi, Bacteria, and Actinomycetes on leaves decomposing in a stream. Appl Environ Microbiol 73(3): 756-767.
- Pyrcz TW, AL Viloria, G Lamas, P Boyer (2011) La fauna de mariposas de la subfamilia Satyrinae del macizo del Ampay (Perú): diversidad, endemismo y conservación (Lepidoptera: Nymphalidae). SHILAP Revta. Lepid 39(154): 205-232.
- Huamantupa Isau, Luza Victorio Miguel, Alfaro-Curitumay Lucero, Ururi Michel, Huaman-Arque, et al. (2017) Diversidad y Biomasa Arbórea en los Bosques Andinos del Santuario Nacional del Ampay, Apurímac-Perú. Rev Q’EUÑA 8: 7-26.
- Felczykowska A, Krajewska A, Zielińska S, Łos JM (2015) Sampling, metadata and DNA extraction. Important steps in metagenomic studies. Acta Biochimica Pol 62(1): 151-160.
- Lombard Nathalie, Emmanuel Prestat, Jan Dirk van Elsas, Pascal Simonet (2011) Soil-specific limitations for access and analysis of soil microbial communities by metagenomics. FEMS Microbiology Ecol 78(1): 31-49.
- Bag S, Bipasa Saha, Bhabatosh Das (2016) An Improved Method for High Quality Metagenomics DNA Extraction from Human and Environmental Samples. Scientific Reports 6: 26775.
- Gantner, Stephan Gantner, Anders F. Andersson, Laura Alonso-Sáez, Stefan Bertilsson (2011) Novel primers for 16S rRNA-based archaeal community analyses in environmental samples. J Microbiol Methods 84(1): 12-18.
- Hong JK, Kim HJ, Cho JC (2014) Novel PCR Primers for the Archaeal Phylum Thaumarchaeota Designed Based on the Comparative Analysis of 16S rRNA Gene Sequences. PLoS ONE 9(5): e96197.
- Fischer Martin A, Güllert Simon, Neulinger Sven C, Streit Wolfgang R, Schmitz Ruth A (2016) Evaluation of 16S rRNA Gene Primer Pairs for Monitoring Microbial Community Structures Showed High Reproducibility within and Low Comparability between Datasets Generated with Multiple Archaeal and Bacterial Primer Pairs. Front Microbiol 7: 1297.
- Özen Asli I, Ussery David W (2012) Defining the Pseudomonas Genus: Where Do We Draw the Line with Azotobacter? Microb Ecol 63(2): 239-248.
- Oliver JD (2005) The Viable but Nonculturable State in Bacteria. J Microbiol 93-100.
- Suzuki M, Suzuki S, Matsui M, Hiraki Y, Kawano F, et al. (2013) Genome sequence of a strain of the human pathogenic bacterium Pseudomonas alcaligenes that caused bloodstream infection. Genome Announc 1(5): e00919-13.
- Krzeslak Joanna, Gerritse Gijs, van Merkerk Ronald, Cool Robbert H, Wim J Quax (2008) Lipase Expression in Pseudomonas alcaligenes Is Under the Control of a Two-Component Regulatory System. Appl Environ Microbiol 74(5): 1402-1411.
- Ashengroph Morahem, Iraj Nahvi, Hamid Zarkesh-Esfahani, Fariborz Momenbeik (2011) Pseudomonas resinovorans SPR1, a newly isolated strain with potential of transforming eugenol to vanillin and vanillic acid. N Biotechnol 28(6):656-664.
- Shintani M, Hosoyama A, Ohji S, Tsuchikane K, Takarada H, et al. (2013) Complete genome sequence of the carbazole degrader Pseudomonas resinovorans strain CA10 (NBRC 106553). Genome Announc 1(4): e 00488-13.
- Iyer Rupa, Ikena Brian, Damaniab Ashish (2017) Genome of Pseudomonas nitroreducens DF05 from dioxin contaminated sediment downstream of the San Jacinto River waste pits reveals a broad array of aromatic degradation gene determinants. Genom Data 14: 40-43.
- SK Shin, CY Hwang, YJ Cho, H Yi (2015) Reclassification of Serpens flexibilis Hespell 1977 as Pseudomonas flexibilis comb. nov., with Pseudomonas tuomuerensis Xin et al. 2009 as a later heterotypic synonym. Syst Appl Microbiol 38(8): 563-566.
- Aragone MR, DM Maurizi, LO Clara, JL Navarro Estrada, A Ascione (1992) Pseudomonas mendocina, an environmental bacterium isolated from a patient with human infective endocarditis. J Clin Microbiol 30(6): 1583-1584.
- Guo W, Wang Y, Song C, Yang C, Li Q, et al. (2011) Complete genome of Pseudomonas m endocina NK-01, which synthesizes mediumchain- length polyhydroxyalkanoates and alginate oligosaccharides. J Bacteriol 193(13): 3413-3414.
- Zhang J, Hou H, Chen G, Wang S, Zhang J (2016) The isolation and functional identification on producing cellulase of Pseudomonas mendocina. Bioengineered 7(5): 382-391.
- Fadel A, Mabrok M, Aly S (2018) Epizootics of Pseudomonas anguilliseptica among cultured seabream (Sparus aurata) populations: Control and treatment strategies. Microb Pathog 121: 1-8.
- Amaike Saori, Nancy P Keller (2011) Aspergillus flavus. Annual Review of Phytopathology 49: 107-133.
- Wei G, Tan D, Chen C, Tong Q, Li XN, et al. (2017) Flavichalasines A-M, cytochalasan alkaloids from Aspergillus flavipes. Sci Rep 7: 42434
- Guo CJ, Wang CC (2014) Recent advances in genome mining of secondary metabolites in Aspergillus terreus. Front Microbiol 5: 717.
- Luo Z-L, et al. (2017) Molecular phylogeny and morphological characterization of asexual fungi (Tubeufiaceae) from freshwater habitats in Yunnan, China. Cryptogamie, Mycologie 38(1): 27-53.
- Murillo C, Albertazzi FJ, Carranza J, Lumbsch HT, Tamayo G (2009) Molecular data indicate that Rhytidhysteron rufulum (Ascomycetes, Patellariales) in Costa Rica consists of four distinct lineages corroborated by morphological and chemical characters. Mycol Res 113(Pt 4): 405-416.
- Zaferanloo B, Bhattacharjee S, Ghorbani MM, Mahon PJ, Palombo EA (2014) Amylase production by Preussia minima, a fungus of endophytic origin: optimization of fermentation conditions and analysis of fungal secretome by LC-MS. BMC Microbiol 14: 55.
- Zhang Ying, Pedro W Crous, Conrad L Schoch, Kevin D Hyde (2011) Pleosporales. Fungal Diversity 53(1): 1-221.






























