Strategies in Protein Engineering to Evolve Proteins and Mimicking Evolution in the Laboratory Scale
Joydeep Chakraborty and Sagar Roy*
Department of Chemistry & Environmental Science, New Jersey Institute of Technology, USA
Submission: August 01, 2018; Published: August 22, 2018
*Corresponding author: Sagar Roy, Department of Chemistry & Environmental Science, New Jersey Institute of Technology, USA, Tel: 862-371-9145; Email: sagar@njit.edu
How to cite this article: Joydeep C, Sagar R. Strategies in Protein Engineering to Evolve Proteins and Mimicking Evolution in the Laboratory Scale. Trends Tech Sci Res. 2018; 3(1): 555601. DOI: 10.19080/TTSR.2018.03.555601
Abstract
Nature has its own way of synthesizing molecules and materials with an array of bio-catalysts known as enzymes. Enzymes comprise of a combination of proteins which in turn are the building blocks of life. With just a backbone combination of Carbon, Hydrogen, Oxygen and Nitrogen, proteins can be perceived as delightfully intelligent and adaptive. Over billions of years, nature have provided a wide range of survival questions in front of these molecules and somehow, they have evolved and survived, as proved by Darwin. This adaptive nature of proteins has been exploited and successful evolution of enzymes is possible towards a desired interest on a laboratory scale using nature’s machinery. Thus, mimicking evolution on a much shorter time-scale. In this article, we address the strategies involving the idea of evolution in a laboratory directed to fulfil our chemical needs. Also, how with time, the strategies have evolved covering up the loop-holes of previous generations. The advent of polymerase chain reactions made combinatorial libraries creation easy and then screen towards a fitness test. With time we learnt, a combination of evolution with rational designing creates shorter and smarter libraries that have a strong ‘impact factor’ upon the desired goal and reduce false-positives. However, while controlling biological systems, scientists are manipulating the factories that actually make the molecules and materials into newer forms that are not natural. Nature, out-does scientists via evolution. Through survival of the fittest, evolution collects all the beneficial mutations through multiple iterated genetic diversification and screening.
Keywords: Directed evolution; Protein engineering; Combinatorial libraries; Mutagenesis; Enzymes; Promiscuous ancestral proteins; Escherichia coli metabolism; Chemical origins; Thermoanaerobacter brockii alcohol dehydrogenase; X-ray crystallographic structures
Introduction
Enzymes are thought to be highly specific biological catalysts having a very discreet range of target molecules or substrates [1, 2]. The evolution of enzymes has been occurring under different selection pressures of nature from promiscuous ancestral proteins. A genomic analysis in Escherichia coli metabolism reveals that 37% of its enzymes have a wide variety of substrate range. However, they are found to catalyze 65% of the known metabolic reactions [3]. The evolution of enzymes in response to environmental demands is the key to the overall organism’s fitness in the nature. This is manifested in the inherent promiscuity of many enzymes. Although, we are thankful to the ‘specialist’ nature of the enzyme molecule, we also do question, how much further can the ‘generalist’ nature be pushed in-vitro or in-vivo so that an enzyme can be evolved more. In the 3-D structure analysis, the active site of an enzyme is the engine, that performs the turn-over of a substrate into the product (catalytic machinery) and more specifically amino-acid residues have been identified from the X-ray crystallographic structures that are key players in the process. Textbook definition of enzymatic activity reveals that the work of the enzyme is broadly related to the binding of the substrate into the active site, followed by a possible conformation change, then the catalytic activity happening in the core of the enzyme and finally, the product leaving the enzyme-product complex. To explain the evolution of enzymatic catalytic activities, scientists have used models of chemical origins to propose the evolutionary initiation of the diversity. One such model demonstrates via chemical models and assumes key mutations in the active site that can lead towards enhanced specificities or sometimes enhanced specificities but with compromised efficiency. Then it proposes that designing a realistic trade-off constraint is possible with mutation combinations where the evolution can be directed towards both enhanced specificity and efficiency [4].
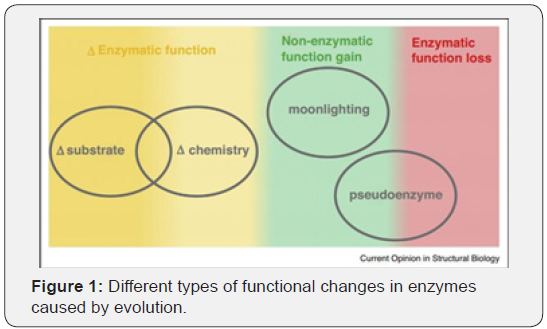
To improve the characteristics of an enzyme structural bioinformatics has been extensively used and with de-novo approaches enzymes have designed for enhanced specificity and functionality. Evolution has been a brute force that pushes the enzymes to evolve and then select the survivors based on their functionality. From Figure 1 it is evident that the evolution not only results in various changes in chemistry and substrates but are also supplemented with rarer gain or loss in the function in the form of moonlighting and pseudoenzymes respectively [5]. Due to the innate promiscuity, many enzymes do show off-target activity even when the natural selection pressure ceases to exist over evolutionary time. At that point of saturation, further catalytic/specificity changes do not improve the fitness in its highly conserved active site. The phenomenon was demonstrated by -lactam antibiotic resistance in bacterial populations over many generations. The promiscuity can also vary between orthologous enzymes which originate from the same family [5-7]. An approach of rational design relying on somewhat of a thorough knowledge of the protein structure that exists for many years is called directed evolution [8,9]. It is a popular technique that has been used to evolve a particular enzyme towards a user-defined goal. The ‘user defined’ goals narrows down the chances to produce the false-positives in comparison to natural selections. Also, this gives an opportunity to mimic natural evolution in a laboratory pilot scale operation. The limitations of this technique however is, screening may be needed for enormous random mutagenesis libraries which in-turn needs a robust high-throughput assay. A combinatorial approach has also been implied to create ‘focused’ libraries that will concentrate on regions of the enzyme active site that has been predicted by rational design to produce beneficial mutations and reduce the number of mutants to screen by a significant number. However, it must also be mentioned that our understanding of the link between the genetic sequence and function lags well behind our desire for designing new enzymes. Which makes predictions of performance-enhancing mutations, extremely challenging.
In the early 90s the most user-defined properties that were targeted for evolution included thermostability, nonnative substrates or activity in harsh organic solvents and after rounds of screening it was sometimes revealed that beneficial mutations could be nowhere near the active site of the enzyme [5,6]. Hence, it would be a hitting a deadended loop to just look for the key in the active site of the enzyme. The power of a combinatorial library is enormous. With a stable high-throughput assay it is possible to analyze millions of mutant proteins in one day. In some other cases however, the number can be less than a few hundred per day. So, development of smarter libraries is very important in this aspect. Using site-directed mutagenesis and site-directed saturation mutagenesis techniques, identification and construction of a ‘mutable landscape’ has been done in case of cytochrome P450 evolution [10-12]. This has been proved to be a very effective technique recently to tackle the very important challenge of the sequence-function relationship in case of directed evolution. The libraries thus generated on mutability landscapes can be used to engineer any fitness trait of user-defined interest [6,7,13]. We are entering into an era where the ‘user-defined’ interest has changed a lot. The demand for synthetic compounds has been growing and ‘green synthesis’ has been one of the most valued tools in the 21st century that uses enzymes as biocatalysts and minimizes the production of non-degradable bye-products and pollutants. So, directed evolution of stereo- or regio-selective enzymes that can behave as biocatalysts in asymmetric transformations is of particular interests to bio-organic chemists. Towards this goal, raised the need to develop the small and smart libraries. And hence forth different strategies including triple-code saturation mutagenesis (TCSM) at multiresidue sites of the Thermoanaerobacter brockii alcohol dehydrogenase by using distinct reduced amino acid alphabets and with requirement minimal screening have been recently reported [13-15]. So not only the physical labor has been greatly reduced, but also chances of false-postive results and codon-degeneracy can be minimized under this protocol [13].
Discussion
To push the promiscuous nature inherent in enzymes has been what scientists been brain-storming over the decades. The further challenges are to create enzymes for reactions that will let them solve problems that neither synthetic organic chemistry nor biological chemistry can address. Being macromolecular catalysts, it would be easier for enzymes to stabilize transition-states which otherwise would be inaccessible to chemical small molecule catalysts as they would likely compete with other lower-energy reaction pathways. Such was reported from cytochrome P450 from Labrenzia arregata [16]. Strategies and efforts are currently being invested to design enzymes with artificial cofactors as well. However, designed enzymes are yet to have the sophistication of nature’s products and struggles remain with other co-factors and prosthetic groups that are currently being addressed.
Conclusion
Over time, nature has been the best template to mankind. In our case, it has been the best chemistry teacher, solving the very challenges of existence of life over billions of years through a range of astonishing conditions. This has been possible only at the molecular level because of the enzymes which act as the fundamental bio-molecules. These systems are good models for a sustainable chemical industry or energy industry that uses renewable natural resources and recycles almost all of it without causing significant damage to the environment. It is not far where we stand today that DNA programmable organisms can produce many of our desired chemical needs.
References
- Van Beilen JB, Z Li (2002) Enzyme technology: an overview. Curr Opin Biotechnol 13(4): 338-344.
- Cooper GM (2000) The central role of enzymes as biological catalysts. Sinauer Associates.
- Hojung Nam, Nathan E Lewis, Joshua A Lerman, Dae-Hee Lee, Roger L Chang, et al. (2012) Network context and selection in the evolution to enzyme specificity. Science 337(6098): 1101-1104.
- Könnyű B, T Czárán (2011) The evolution of enzyme specificity in the metabolic replicator model of prebiotic evolution. PLoS One 6(6): e20931.
- Tyzack JD, Furnham N, Sillitoe I, Orengo CM, Thornton JM, et al. (2017) Understanding enzyme function evolution from a computational perspective. Curr Opin Struct Biol 47: 131-139.
- Arnold FH (2018) Directed evolution: bringing new chemistry to life. Angew Chem Int Ed Engl 57(16): 4143-4148.
- Acevedo-Rocha CG, M Ferla, MT Reetz (2018) Directed evolution of proteins based on mutational scanning, Methods Mol Biol: 87-128.
- Clark DP, NJ Pazdernik (2015) Biotechnology: Newnes.
- Steiner K, H Schwab (2012) Recent advances in rational approaches for enzyme engineering. Comput Struct Biotechnol J 2(3): p. e201209010.
- Li A, CG Acevedo-Rocha, MT Reetz (2018) Boosting the efficiency of site-saturation mutagenesis for a difficult-to-randomize gene by a two-step PCR strategy. Appl Microbiol Biotechnol.102(14): 6095-6103.
- Carlos G Acevedo-Rocha, Charles G Gamble, Richard Lonsdale, Aitao Li, Nathalie Nett, et al. (2018) P450-Catalyzed Regio-and Diastereoselective Steroid Hydroxylation: Efficient Directed Evolution Enabled by Mutability Landscaping. ACS Catalysis 8(4): 3395-3410.
- Badenhorst, CP, UT Bornscheuer (2018) Getting Momentum: From Biocatalysis to Advanced Synthetic Biology. Trends in biochemical sciences 43(3): 180-198.
- Qu G, Lonsdale R, Yao P, Li G, Liu B, et al. (2018) Methodology Development in Directed Evolution: Exploring Options when Applying Triple‐Code Saturation Mutagenesis. Chembiochem 19(3): 239-246.
- Bougioukou DJ, Sabrina Kille, Andreas Taglieber, Manfred T Reetz (2009) Directed evolution of an enantioselective enoate‐reductase: testing the utility of iterative saturation mutagenesis. Advanced synthesis & catalysis 351(18): 3287-3305.
- Sun Z, Lonsdale R, Li G, Reetz MT (2016) Comparing Different Strategies in Directed Evolution of Enzyme Stereoselectivity:Singleversus Double‐Code Saturation Mutagenesis. Chembiochem 17(19): 1865-1872.
- Hammer SC, Kubik G, Watkins E, Huang S, Minges H, et al. (2017) Anti-Markovnikov alkene oxidation by metal-oxo-mediated enzyme catalysis. Science 358(6360): 215-218.






























