Identification of Bacteria from Substrata as Metamorphosis Inhibitors for the Coral Acropora
Ayaka Horikoshi, Chie Sasaki and Masayuki Hatta*
Department of Biology, Ochanomizu University, Japan
Submission: June 16, 2017 ; Published: August 04, 2017
*Corresponding author: Masayuki Hatta, Department of Biology, Ochanomizu University, Tokyo 112-8610, Japan, Email: hatta.masayuki@ocha.ac.jp
How to cite this article: Ayaka H, Chie S, Masayuki H.Identification of Bacteria from Substrata as Metamorphosis Inhibitors for the Coral Acropora. Fish & Ocean Opj. 2017; 4(2): 555635. DOI: 10.19080/OFOAJ.2017.04.555635
Abstract
Bacteria on substrata certainly influence metamorphosis and settlement of coral larvae. We isolated bacteria from substrata and screened them for the capability of inducing or inhibiting metamorphosis of Acropora. Out of 468 isolates, only eight isolates revealed metamorphosis- inducing activities, which were weak or non-reproducible. Three of those isolates were identified as Alteromonasspp. Based on by 16SrDNA.On the other hand, metamorphosis-inhibiting bacteria were obtained efficiently, two out of 20 isolates, indicating a majority in bacterial communities on substrata. One revealed quick but temporal inhibition and the other revealed slow but persistent inhibition onto the metamorphosis induction by a peptide hormone. Thus, at least two inhibition pathways are suggested. Both of the two strains turned out to belong to the genus Pseudoalteromonas. Responses of coral larvae to mixtures of exclusive cues on the substratum, inducers and inhibitors, seem important in the ecological strategy to select their destiny in natural situations.
Keywords: Acropora; Bacteria; Metamorphosis; Settlemen.
Introduction
The coral genus Acropora shares major components in many Indo-Pacific reefs, and its recruitment holds the key for maintenance and recovery of reef assemblages. Especially in the recovery from denuded reefs, efficient recruitment of propagules is important to reconstruct populations of various coral species and to keep genetic diversities as well. It would be critical to know the conditions suitable for settlement of acroporids and to control the environments if we aim to accelerate recruitment for reef restoration. It is empirically known that acroporids require chemical cues from substrata for the initiation of a sequence of metamorphosis and settlement, and that they have strict preference to the environmental signals. However, only a small number of micro-organisms have so far been identified as natural inducers of metamorphosis and settlement for acroporids, such as particular species of calcareous coralline algae [1] and a bacterium of the genus Pseudoalteromonas [2,3]. It is also well known that conditioned tiles, which were kept in the sea for several months, act as efficient substrata on which bacteria grow and cover the surface. Biofilms on artificial substrata were observed with implications in chemical cues for induction of coral metamorphosis and settlement [4]. Considering the efficiency of the conditioned tiles, it is expected that much more bacteria species act as inducers for coral metamorphosis and settlement.
On the other hand, it is conceivable that there are microorganisms on substrata that have inhibitory activities to coral settlement. But such inhibitory bacteria have not been focused on, even though inhibitors would also be important factors for coral recruitment in natural situations. Extensive screening is needed for finding and identifying more bacteria that induce or inhibit metamorphosis and settlement of coral larvae in order to describe bacteria compositions on substrata as controllers for coral recruitment.
Materials and Methods
Isolation and culture of bacteria
Terracotta tiles (5cm*5cm) were submerged in a tank supplied with running seawater at Akajima Marine Science Laboratory, Aka Island, Okinawa, Japan (30 ºN, 123 ºE) for 1 year to allow growth of bacteria communities on the tiles. Planula larvae of Acropora digitifera were subjected to those conditioned tiles in a bowl. Small areas where larvae settled in aggregates were selected as the source of metamorphosis-inducing bacteria and another area where larvae did not settle were selected as the source of metamorphosis-inhibiting bacteria. Tile surfaces were scraped and suspended in 0.2μm-filtered seawater (FSW). The suspension was spread on ZoBell2216E agar plates after stepwise dilutions. After culture at around 26 ºC for 1-3 days, each clonal colony was spread on a new agar plate and cultured for 1-6 days to obtain a certain quantity of bacteria cells.
Metamorphosis assays
Planula larvae of Acropora tenuis and A. digitifera were prepared according to Iwao et al. [5]. Planulae were reared by daily transfer to fresh FSW, and also metamorphosis assays were done, in the laboratory kept at around 26 °C. Planula larvae of day 5-13 post fertilization were used for the assays. Metamorphosis was defined as appearance of 6 septations of the tissue, even if faintly, in rounded individuals.
The metamorphosis-inducing assay was performed by administration of each strain of bacterial cells in suspension into 1ml of FSW containing 5 planulae in a well of 24-well cell culture multiplates. Approximately 20μil volume of bacterial cells was used for each assay, and the cell numbers were confirmed as 1-4x108cfu by dilution and culture on agar plates. The number of metamorphosed animals was counted 1 day after the initiation of treatment. Larvae of A. tenuis were used except for some initial assays using A. digitifera.
The metamorphosis-inhibition assay was achieved by combining bacteria and a metamorphosis-inducing peptide Hym-248 [5]. Approximately 20μiL volume of bacterial cells were administrated in suspension to 10 planulae in 1mL of FSW in a well of 24-well cell culture multiplate simultaneously with Hym-248 at a final concentration 2x10-6M, and the number of metamorphosed animals was counted along time up to 12hrs and compared with positive controls, which were treated with only Hym-248. Cell densities in assays were approximately 3x106cfu and 1x107cfu respectively for the two examined strains named MIP01 and MIW01.
Results and Discussion
Screening of metamorphosis-inducing bacteria
In total 468 independent strains of bacteria were screened for the metamorphosis-inducing activity, and 7 strains were found positive in the first screening (Table 1). The rest showed no activities of metamorphosis-induction and also no toxicity to planula larvae. Although the functions remain unknown, 89 strains caused discontinuation of movements and elongation of the body on planulae. These 89 strains were tested again, and one additional strain (MA505) was found to show metamorphosis- inducing activity (Table 1). The frequency of positive strains, 8 per 468, corresponded to 1.7%. Negri et al. [2] obtained a similar frequency, 1.3%; one metamorphosis-inducing strain from 80 strains screened. Note that these values reveal frequencies among bacteria culturable by conventional isolation culture methods.
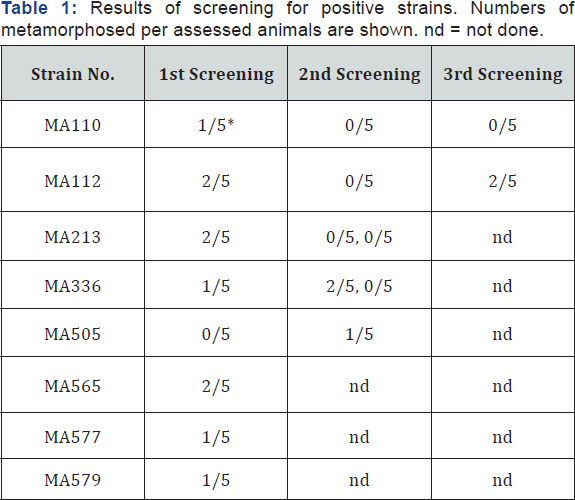
*Results using larvae of A. digitifera.
Repeated tests were done for 5 out of 8 positive strains, but the rest could not be assessed due to over-aging of the planulae. Only 1-2 per 5 larvae metamorphosed in the positive cases of the 8 strains (Table 1) revealing weak activities compared with the results by Negri et al. [2], which showed 50-80% efficiency of metamorphosis induction. Reproducibility was also poor in our tests (Table 1). Some improvements may remain in experimental designs; such as bacteria densities in the assay, use of bacteria grown on substrata instead of suspension, and tests using other coral species. However, positive activities seem specific, even though the activities were low, because a large number of strains showed no effects.
The 8 positive strains were maintained by 2-weeks passages for 1 year and assessed again. None of them retained the activity (data not shown). This loss of activity might be due to alteration of physiological conditions during the isolated culture of bacteria.
Metamorphosis-inhibiting bacteria

As a pre-screening, 20 strains were checked to see whether they made planulae stop swimming and elongate their body. Out of 5 strains showing those activities, 2 strains were chosen for further testing as the inhibition assay is laborious. One strain, named MIP01, formed purple color colonies and another, named MIW01, formed white colonies. Simultaneous application of MIP01 and the peptide Hym-248 resulted in complete inhibition of metamorphosis during the observation period, whereas MIW01 reduced metamorphosis efficiency to about 43-73% (Table 2). These results were reproduced after 2-weeks passages for 1 year (data not shown). Contrary to the inducer strains, the both inhibitor strains retained their functions stably in the culture conditions. The other 3 strains showed no effects (data not shown).
When larvae were treated with MIW01 alone for 12hrs and afterwards Hym-248 was added, strong inhibition was observed (Table 2). In contrast, MIP01 pre-treatment resulted in no inhibition onto the metamorphosis induction by Hym- 248, suggesting desensitization of planulae to MIP01 stimuli. These results suggest that the inhibitory action by MIP01 is quick and temporal and that by MIW01 is slow and persistent. When the bacteria were boiled for 5 minutes, those inhibiting activities were reduced in MIP01 and enhanced in MIW01 (data not shown). Thus, at least two pathways of metamorphosis inhibition in planlae responding to distinct bacterial molecules were found in this study.
The frequency of getting inhibitor bacteria was much higher than that of obtaining inducer bacteria. If these differences in frequency reflect abundance of those bacteria on the substrata, coral larvae would find their places for settlement after passing through lots of negative patches. Only metamorphosis induction has been focused in corals based on interests in propagation of corals from larvae, however, metamorphosis inhibition should be considered to understand mechanisms of recruitment in corals.
Molecular identification of the bacteria
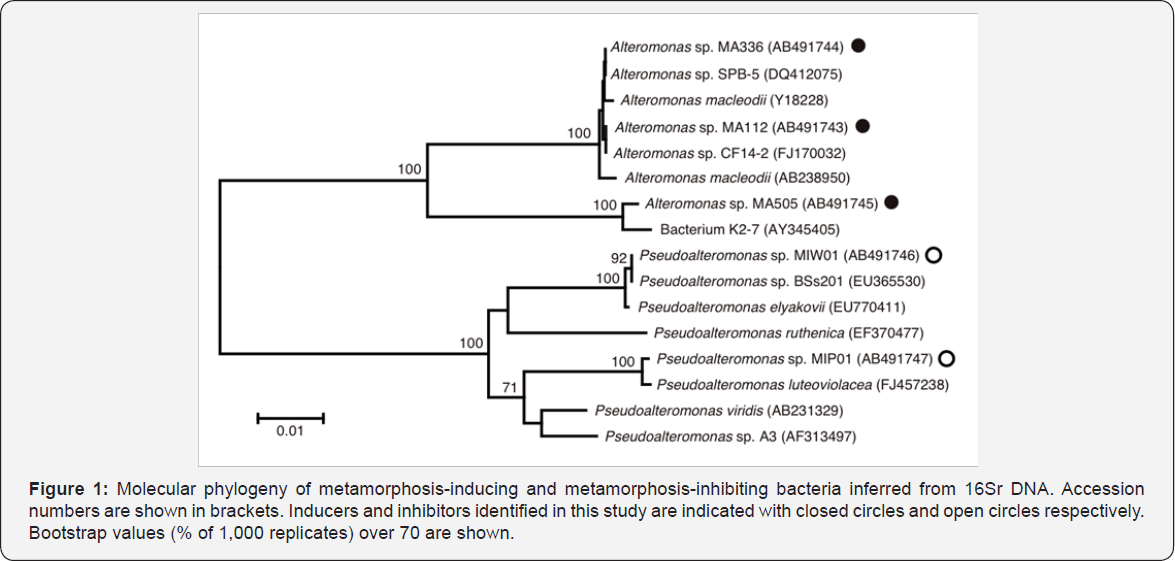
Using conventional methods, 16SrDNA sequences were determined for the isolated strains. Among the metamorphosis- inducing bacteria, 3 strains (strain MA112, MA336 and MA505) were chosen for molecular identification, and the results of BLAST search revealed that the 3 strains are all included in the genus Alteromonas. The 2 metamorphosis-inhibiting strains (strain MIP01 and MIW01) were both included in the genus Pseudoalteromonas. Sequence data revealing high similarity to the queries were retrieved from the database, and a phylogenetic tree was constructed by the Neighbor-Joining method (Figure 1). Metamorphosis-inducing strains MA112 and MA336 are closely related to each other and MA505 falls into a separate branch within the Alteromonas clade. Metamorphosis-inhibiting strains MIP01 and MIW01 are inferred as two independent species of Pseudoalteromonas. This genus contains also a metamorphosis- inducing strain isolated by Negri et al. [2] (Figure 1, Pseudoalteromonas sp. A3). Simply, the two features are pointed out for the phylogenies; inducer bacteria are found in different genera (Alteromonas and Pseudoalteromonas), and a single genus (Pseudoalteromonas) includes both inducer and inhibitor species. The former would be a result of either production of the same compound(s) to stimulate coral metamorphosis or a broad preference of acroporids' planulae for various chemical compounds as environmental cues for metamorphosis initiation. The latter would be due to diversity within a genus with implication in influencing coral metamorphosis, and diversity of compound production as well [6-8].

Conclusion
We screened nearly 500 isolates of bacteria to find inducers of metamorphosis of acroporid larvae. However, only several strains revealed weak or non-reproducible activities that were finally lost after passages. There would be a limitation of isolating culture as more than 99% of bacteria in environments are thought unculturable by conventional culture methods due to lack of proper culture conditions. On the other hand, metamorphosis-inhibiting bacteria were efficiently obtained from small numbers of isolates. This is the first report of isolation of bacteria that are inhibitory to metamorphosis of the coral Acropora. The 2 inhibitor strains revealed different modes of action, hence larvae should have at least 2 signal transduction pathways in suspending metamorphosis. It is conceivable that those inducing and inhibiting bacteria grow together in a small area of substrata in nature. Responses of planula larvae to mixtures of inducers and inhibitors on the substratum may be important as the ecological strategy to choose their destiny in natural situations.
Acknowledgment
We thank Akajima Marine Science Laboratory which enabled us to perform major part of this work. This work was supported by SUMITOMO Foundation, Yamada Science Foundation, and the Grants-in-Aid for Scientific Research by MEXT Japan.
References
- Morse ANC, Iwao K, Baba M, Shimoike K, Hayashibara T, Omori M (1996) An ancient chemosensory mechanism brings new life to coral reefs. Biol Bull 191(2): 149-154.
- Negri AP, Webster NS, Hill RT, Heyward AJ (2001) Metamorphosis of broadcast spawning corals in response to bacteria isolated from crustose algae. Mar Ecol Prog Ser 223: 121-131.
- Tebben J, Tapiolas DM, Motti CA, Abrego D, Negri AP, et al. (2011) Induction of larval metamorphosis of the coral Acropora millepora by tetrabromopyrrole isolated from a Pseudoalteromonas bacterium. PLoS one 6(4): e19082.
- Webster NS, Smith LD, Heyward AJ, Watts JEM, Webb RI, et al. (2004) Metamorphosis of a scleractinian coral in response to microbial biofilms. Appl Env Microbiol 70(2): 1213-1221.
- Iwao K, Fujisawa T, Hatta M (2002) A cnidarian neuropeptide of the GLWamide family induces metamorphosis of reef-building corals of the genus Acropora. Coral Reefs 21(2): 127-129.
- Fukami N, Omori M, Hatta M (2000) Phylogenetic relationships in the coral family Acroporidae, reassessed by inference from mitochondrial genes. Zool Sci 17(5): 689-696.
- Reysenbach AL, Giver LJ, Wickham GS, Pace NR (1992) Differential amplification of rRNA genes by polymerase chain reaction. Appl Environ Microbiol 58(10): 3417-3418.
- Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173(2): 697-703.






























