Misexpression of Genes Lacking Cpg Islands is a Hallmark of Aging
Jun-Yeong Lee* and Samuel Beck*
Davis Center for Regenerative Biology and Medicine, MDI Biological Laboratory, USA
Submission: March 1, 2022; Published: March 14, 2022
*Corresponding author: Jun-Yeong Lee and Samuel Beck, Davis Center for Regenerative Biology and Medicine, MDI Biological Laboratory, USA
How to cite this article: Jun-Yeong L, Samuel B. Misexpression of Genes Lacking Cpg Islands is a Hallmark of Aging. OAJ Gerontol & Geriatric Med. 2022; 6(3): 555690. DOI: 10.19080/OAJGGM.2022.06.555690
Abstract
Aging entails global disorganization of chromatin architecture characterized by disruption of the nuclear lamina and associated heterochromatin. How these structural changes contribute to age-related degenerative changes is unclear. We show that genes lacking CpG islands (CGI- genes), which form heterochromatin when transcriptionally silent, are globally misexpressed in aged nuclei with disrupted chromatin architectures. We demonstrate that CGI- gene misexpression is a common feature of mammalian aging and explains the molecular basis of various age-associated defects, ranging from loss of cellular identity and increased transcriptional noise to age-associated uncontrolled expression of secretory proteins, including signaling molecules putatively causing chronic inflammation. Our findings reveal that CGI- gene misexpression is directly associated with age-related physiological deterioration, thus provide a novel biomarker of aging.
Keywords: Aging, Chromatin architecture, Gene regulation, CpG islands, Heterochromatin, CGI- gene misexpression, aged tissues, mammalian promoter-associated elements, RNA-seq data, aged cells
Abbreviations: CGI- CpG island; DO- Diversity Outbred; LBR- Lamin B receptor; KD- Knock-down; SASP- senescence associated secretory phenotype
Introduction
Changes in the 3-D architecture of chromatin are observed in various diseases and are also a hallmark of aging. Disruption of the nuclear lamina and associated heterochromatin are commonly observed in various aging contexts, including premature aging diseases [1,2], cellular senescence [3,4], and normative aging [5]. Although these conserved structural changes have been reported for over two decades, their impacts on transcription and contribution to age-related degenerative changes remain unknown.
Our recent computational approaches demonstrated that CGIs, mammalian promoter-associated elements, provide important clues to answering this question. Specifically, we found that only genes not associated with CGIs (CGI- genes) reside within lamina-associated heterochromatin when they are transcriptionally silent. In contrast, genes associated with CGIs (CGI+ genes) remain as euchromatin even when repressed [6–8]. Based on this evidence, we hypothesized that heterochromatin decondensation during aging would specifically result in the uncontrolled expression of CGI- genes. We tested this hypothesis through large-scale transcriptome and proteome analyses of DO mice, which are a genetically diverse mouse resource that mimics the complexity of the human population with variable rates of physiological aging [9,10], and validated our results both using mice with disrupted nuclear architectures and through systematic meta-analysis of published transcriptome profiling data in our most recent study [11].
Disorganization of nuclear architecture during aging drives CGI- gene misexpression
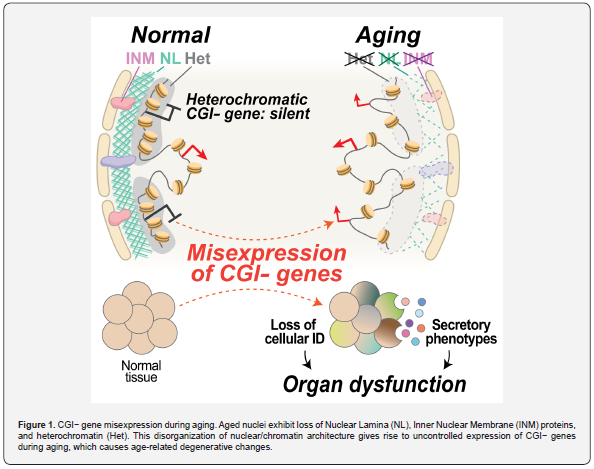
Our DO mouse transcriptome analysis indeed demonstrates that over 30% of CGI- genes are mis-activated in aged kidneys and hearts compared to young tissues, and this pattern coincides with disorganization of chromatin architecture. Our experimental validation using model mice with disrupted nuclear architectures, as well as meta-analysis of RNA-seq data generated from conditions mimicking aged nuclei further demonstrate that nuclear architecture disruption is sufficient to induce heterochromatin decondensation and CGI- gene misexpression as observed during aging Upper part of (Figure 1). Our data also show that nuclear architecture disruption and resulting CGI- gene misexpression are directly associated with physiological deterioration of aged organs.
Misexpressed CGI- genes is responsible for physiological deterioration during aging
Notably, CGI- gene misexpression explains the molecular basis of various degenerative changes previously reported throughout the aging process Lower part of (Figure 1). For example, nuclear architecture disruption leads to the expression of tissue-specific CGI- genes in tissues where they are not normally expressed. Our single-cell RNA-seq analyses reveal that CGI- gene misexpression is responsible for age-associated transcriptional noise (cell-to-cell transcriptional variability). These data indicate that disrupted regulation of CGI- genes during aging results in loss of functional identity of aged cells. We also show that uncontrolled secretory phenotypes commonly observed during aging, such as SASP [12,13], are largely attributable to CGI- gene misexpression. Our findings indicate that nuclear architecture disruption and resulting CGI- gene misexpression drive disruption of intercellular communication and fuel chronic inflammation in aged tissues.
CGI- gene misexpression is a common feature of aging and age-associated diseases
Our large-scale meta-analysis of transcriptome profiling data further demonstrates that CGI- gene misexpression is a common feature of mammalian aging and age-associated diseases. Various aged tissue types, such as brain, liver, and muscle, significantly misexpress CGI- genes compared to young tissues. Interestingly, CGI- gene misexpression can be suppressed by previously validated anti-aging interventions, such as caloric restriction, rapamycin/ acarbose treatment, and heterochronic parabiosis. Lastly, a broad range of age-associated diseases, such as myocardial, neurodegenerative, hepatic, and macular diseases, also exhibit dramatic CGI- gene misexpression compared to healthy controls. These results demonstrate that CGI- gene misexpression provides a novel biomarker for physiological deterioration associated with aging.
Conclusion
Our study establishes that loss of transcriptional homeostasis of CGI- genes is a shared trait of mammalian aging, providing a fundamental basis for understanding aging and its associated degenerative changes. Our data also suggest that age-associated CGI- gene misexpression is a novel biomarker of physiological aging which offers an effective therapeutic target for delaying or ameliorating degenerative changes associated with aging/the aging process.
Acknowledgements
This study was supported by the Morris Scientific Discovery Funds and the National Institutes of Health under grant number R01AG068179.
Conflict of Interest
The authors declare that they have no competing interests.
References
- Zhang W, Li J, Suzuki K, Qu J, Wang P (2015) A Werner syndrome stem cell model unveils heterochromatin alterations as a driver of human aging. Science 348: 1160–1163.
- Beyret E, Liao HK, Yamamoto M, Hernandez-Benitez R, Fu Y (2019) Single-dose CRISPR–Cas9 therapy extends lifespan of mice with Hutchinson–Gilford progeria syndrome. Nat. Med 25: 419-422.
- Shah PP, Donahue G, Otte GL, Capell BC, Nelson DM (2013) Lamin B1 depletion in senescent cells triggers large-scale changes in gene expression and the chromatin landscape. Genes Dev 27: 1787-1799.
- Chandra T, Ewels PA, Schoenfelder S, Furlan-Magaril M, Wingett SW (2015) Global reorganization of the nuclear landscape in senescent cells. Cell Rep 10(4): 471-483.
- Scaffidi P, Misteli T (2006) Lamin A-dependent nuclear defects in human aging. Science 312: 1059-1063.
- Beck S, Lee BK, Rhee C, Song J, Woo AJ (2014) CpG island-mediated global gene regulatory modes in mouse embryonic stem cells. Nat Commun. 5: 1–12.
- Beck S, Rhee C, Song J, Lee B-K, LeBlanc L (2018) Implications of CpG islands on chromosomal architectures and modes of global gene regulation. Nucleic Acids Res 46(9): 4382-4391.
- Lee J-Y, Song J, Leblanc L, Davis I, Kim J (2021) Conserved dual-mode gene regulation programs in higher eukaryotes. Nucleic Acids Res 49: 2583-2597.
- Gyuricza IG, Chick JM, Keele GR, Deighan AG, Munger SC (2020) Genome-wide transcript and protein analysis reveals distinct features of aging in the mouse heart. bioRxiv 2020: 272260.
- Takemon Y, Chick JM, Gerdes Gyuricza I, Skelly DA, Devuyst O (2021) Proteomic and transcriptomic profiling reveal different aspects of aging in the kidney. Elife 10: 62585.
- Lee J-Y, Davis I, Youth EHH, Kim J, Churchill G (2021) Misexpression of genes lacking CpG islands drives degenerative changes during aging. Sci Adv 7: 9111.
- Childs BG, Gluscevic M, Baker DJ, Laberge RM, Marquess D (2017) Senescent cells: An emerging target for diseases of ageing. Nat Rev Drug Discov 16(10): 718-735.
- Coppé JP, Patil CK, Rodier F, Sun Y, Muñoz DP (2008) Senescence-associated secretory phenotypes reveal cell-nonautonomous functions of oncogenic RAS and the p53 tumor suppressor. PLoS Biol. 6: e0060301.






























