Urine Biomarkers for the Detection of Clinically Significant Prostate Cancer
Pérez Lanzac A1,2*
1Professor at Alfonso X el Sabio University, Spain
2Department of Urology, Hospital Ruber Internacional, Spain
Submission:May 18, 2021;Published:September 23 2021
*Corresponding author:Alberto Pérez-Lanzac, Servicio de Urología, Hospital Ruber Internacional, Madrid Spain
How to cite this article:Pérez L. Urine Biomarkers for the Detection of Clinically Significant Prostate Cancer. JOJ Urology & Nephrology, 2021; 7(4): 555717. DOI: 10.19080/JOJUN.2021.07.555717
Abstract
Prostate cancer (PCa) biomarkers, especially urine-based ones, are useful tools able to improve the diagnosis and management of the disease. They help physicians to determine if a biopsy is required, mainly with prostate-specific antigen (PSA) levels between 4.0 and 10.0ng/mL, and therefore avoid unnecessary biopsies. Urine biomarkers can differentiate a clinically significant PCa and thus guide toward the most appropriate treatment. Diagnostic accuracy of novel urine-based test is higher than PSA level. The present manuscript aims at showing the most complete and updated information about urine-based biomarkers for the detection of clinically significant PCa, i.e. PCA3, SelectMDx, ExoDx Prostate IntelliScore, and Mi‑Prostate score. The literature basing their diagnostic efficacy is extensive in the case of PCA3, however limited for the remaining ones. Long-term, prospective, head-to-head comparisons, and real-world studies are required to complete and corroborate the available results. Besides this, the European Association of Urology has included in their guidelines the use of urine-based tests for deciding whether or not to undergo an initial or repeat biopsy. Further investigations will be the basis for the approval and widely use of these tests in routine clinical practice.
Keywords: Prostate cancer; Biomarker; Urine; PCA3; SelectMDx; IntelliScore; Mi‑Prostate score
Introduction
Prostate cancer (PCa) is the second most frequent carcinoma diagnosed among men (7.1% of all cancers), and the fifth leading cause of cancer death worldwide (3.8% of all cancer-related deaths) [1]. According to GLOBOCAN, 1,276,106 new cases and 358,989 deaths were reported in 2018. Besides the elevated incidence, survival rate has improved over the last decades because the disease is diagnosed at earlier stages [2]. PCa represents a major health concern, and is associated with important healthcare costs [3]. Advanced age, family history, and ethnicity are well-established risk factors for PCa [2]. It shows diverse clinic pathological and progression expressions. At early stage, the disease is often asymptomatic [2]. The diagnosis is based on a digital rectal examination (DRE), serum prostate-specific antigen (PSA) levels, and Trans rectal ultrasound (TRUS)-guided biopsy [4]. PSA levels >4.0ng/mL are frequently used as a threshold for recommending a prostate biopsy [5]. The positive predictive value (PPV) of PSA for detecting a PCa is 25-40% in men with a level 4-10ng/mL. Nevertheless, 65-75% of men within this PSA range have a negative result in the biopsy, and up to 15% of those with PCa present with PSA level <4.0 ng/mL [5,6].
Biomarkers for the detection of PCa
Since its introduction in 1990s, PSA testing has led to the detection of a higher number of cases; however, the lack of specificity for detecting a significant PCa has also entailed an increased number of unnecessary biopsies, over diagnosis, and overtreatment in patients with indolent disease [7]. Generally, indolent disease has not impact on the quality of life or the life expectancy, and it is managed by following watchful waiting or active surveillance [8]. Patients with indolent disease who receive a treatment can experience complications without decreasing their risk for dying due to the cancer [9,10]. Therefore, differentiating the types of PCa, i.e. very low and low risk versus intermediate and significant, by means of histological, genetic or genomic analyses has become necessary for the choice of the most appropriate management. On the other hand, one of the most relevant parameters used for determining the disease stage and grade (and, in turn, the prognosis) is the Gleason grading system [11].
Nevertheless, the quantification of the Gleason score (GS) in the biopsy is relatively subjective, and the random prostate biopsy may cause under-sampling [8,12]. For this reason, there is a need for avoiding subjectivity and interpretations when grading. Advances in molecular biology have led to the identification of biomarkers implicated in the carcinogenesis of PCa [6]. Over the last decade, a number of tests have been commercialized, including urine-based (ExoDx Prostate IntelliScore, EPI, SelectMDx, prostate cancer antigen 3, PCA3, Mi‑Prostate score, MiPS) blood-based (Four Kallikrein Score, Prostate Health Index), and tissue-based ones (Oncoytype DX, Prolaris, ConfirmMDx, Decipher) [13]. An ideal biomarker for the detection of PCa should be obtained noninvasively, specific for clinically significant PCa (csPCa), able to differentiate a significant from indolent disease, inexpensive, and easily accessible [7]. Additionally, there is a demand for improving the sensitivity and specificity of the tests [8].
Available evidence on urine biomarkers
PCA3
The Progensa PCA3 assay determines the ratio PCA3/PSA mRNA in a sample of urine collected after DRE. The PCA3 is a gene that is highly up-regulated in PCa [14]. It has been established a PCA3 cut-off value of 25, with lower values indicating a reduced risk for a positive biopsy [7]. The key validation study of PCA3 was performed by Marks et al. [15], who collected urine samples from 233 men with PSA level ≥ 2.5ng/mL and at least a previous negative biopsy. A summary of most representative results from studies using PCA3 is shown in Table 1. The biopsy identified PCa in 26.5% of patients. The area under the curve (AUC) for detecting a positive PCa was 0.68 (95% confidence interval, 95% CI, 0.60- 0.76). The PCA3 showed a differential sensitivity, specificity, PPV, and negative predictive value (NPV) depending on the cut-off value selected. By evaluating a PCA3 cut-off value of 35, the sensibility and specificity were 58% and 72%, respectively; however with a value of 50, they were 47% and 81%. Further validation data derived from 1,072 patients in the placebo arm of the REDUCE trial [16].
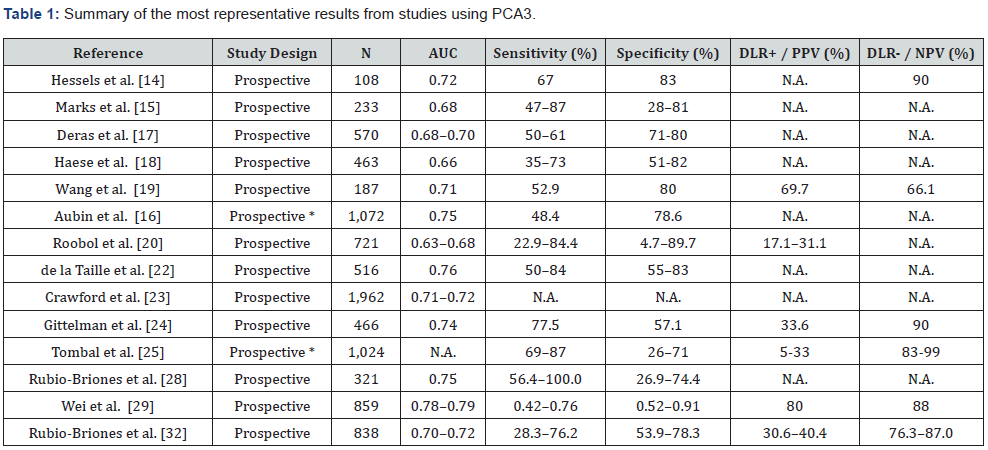
AUC: area under the curve; DLR+, positive diagnostic likelihood ratio; DLR-, negative diagnostic likelihood ratio; PPV: positive predictive value; NPV: negative predictive value; N.A.: not available. *Placebo arm from REDUCE trial.
PCA3 scores were evaluated before the biopsies at year 2 and 4. A total of 190 patients (17.7%) showed a positive biopsy over this period of time. The AUC for PCA3 was 0.69 (95% CI, 0.65- 0.74), significantly higher than serum PSA (0.612; 95% CI, 0.57- 0.66). When introducing clinical variables (prostate volume, age and family history), AUC increased to 0.75 (95% CI, 0.71-0.79). Furthermore, the PCA3 score at year 2 was able to significantly predict the biopsy outcome at year 4. Diverse further studies, systematic reviews and meta-analyses have confirmed the acceptable accuracy of PCA3 for detecting PCa [17-33]. Roobol et al. [20] in a prospective study evaluated the use of PCA3 as a firstline diagnostic test in 721 men. The number of avoided biopsies was 4.4%, 26.1%, 51.7%, and 87.5% when PCA3 score was ≥10, ≥20, ≥35, and ≥100, respectively. The AUC with PCA3 was 0.64 (95% CI, 0.58-0.69) for total patients, 0.68 (95% CI, 0.56-0.80) in men with previous negative biopsy, and 0.63 (95% CI, 0.56-0.69) in those with PSA <3.0.
The sensitivity and specificity ranged between 22.9-84.4% and 4.7-89.7%, respectively. The NPV varied from 17.1% to 31.1%. Hu et al. [26] in a systematic review and meta-analysis involving 16 research studies and 6,693 individuals, explored the diagnostic value of PCA3 with a cut-off value of 35μg/L. Overall results were: sensitivity (0.57; 95 % CI, 0.55-0.59), specificity (0.71; 95 % CI, 0.70-0.73), positive likelihood ratio (2.12; 95 % CI, 1.89-2.38), negative likelihood ratio (0.55; 95% CI, 0.50-0.61), diagnostic odds ratio (3.93; 95 % CI, 3.28-4.72), and AUC of 0.71. Similarly, Cui Y et al. in a systematic review and meta-analysis with 46 clinical trials and 12,295 subjects, showed a pooled sensitivity of 0.65 (95% CI, 0.63-0.66), specificity of 0.73 (95% CI, 0.72-0.74), positive likelihood ratio of 2.23 (95% CI, 1.91-2.62), negative likelihood ratio of 0.48 (95% CI, 0.44-0.52), diagnostic odds ratio of 5.31 (95% CI, 4.19-6.73, and AUC of 0.75 (95% CI, 0.74-0.77).
SelectMDx
SelectMDx is a molecular test that detect HOXC6 and DLX1 mRNA levels in urine obtained after DRE [7]. The most important validation study was carried out by Van Neste et al. [9] they collected urine samples of patients after DRE and previously to the prostate biopsy (initial or repeated). The summary of most representative results from studies using SelectMDx is shown in Table 2. The biopsy was underwent based on a PSA level ≥3 ng/ mL, abnormal DRE, or family history of PCa. A first cohort of 519 patients participated in the exploratory analysis of RNA levels (HOXC4, HOXC6, TDRD1, DLX1, KLK3, and PCA3 gene) [9]. The combination of HOXC6 and DLX1 mRNA levels showed the highest AUC (0.76; 95% CI, 0.71-0.81) for detecting high-grade PCa (GS ≥7). Moreover, the PPV and NPV were 27% and 94%, respectively. A second cohort of 386 patients participated in the validation study, where the HOXC6/DLX1 gene signature was tested together with traditional clinical variables (PSA density, previous biopsies, PSA level, age, family history, DRE) [9]. A highest AUC, 0.87 (95% CI, 0.81-0.93), was found when variables were introduced in the model, a 98% of NPV and 53% reduction in unnecessary biopsies. Subsequently, Haese et al. [34], in a multicenter and multinational study (The Netherlands, France and Germany), evaluated SelectMDx for detecting csPCa prior to an initial biopsy in a training (1,039 men) and a validation (916) cohort.
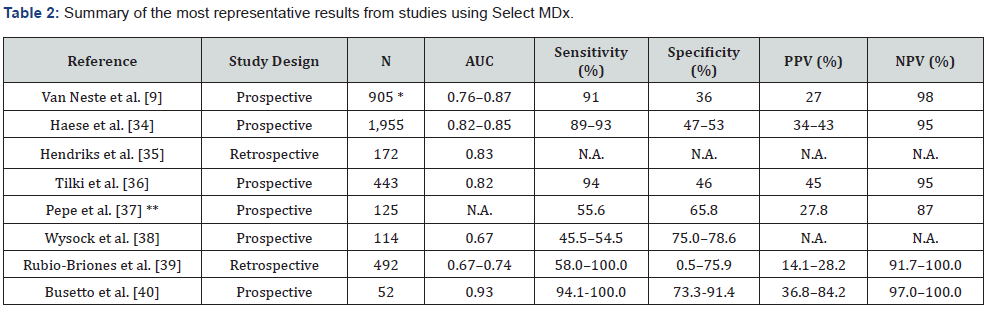
AUC: area under the curve; PPV: positive predictive value; NPV: negative predictive value; N.A.: not available
* Total patients (training and validation cohorts).
** Use of Select MDx in individuals in active surveillance (off-label).
In the latter, the AUC for HOXC6/DLX1 mRNA was 0.85 (95% CI, 0.83-0.88), 93% of sensitivity, 47% of specificity, 43 of PPV, and 95% of NPV. Individuals with PSA ≤10 ng/mL (n=715) showed an AUC of 0.82 (95% CI, 0.79-0.86), 89% of sensitivity, 53% of specificity, 34% of PPV, and 95% of NPV. Further studies have complemented the information of provided in the validation studies [35-40]. Tilki et al. [36], in a prospective study with 443 men undergoing an initial biopsy, revealed an AUC with SelectMDx of 0.82 (95% CI, 0.78-0.86) for the detection of GS ≥7 PCa. Sensitivity, specificity, PPV, and NPV were 94% (95% CI, 89- 98%), 46% (95% CI, 40-52%), 45% (95% CI, 42-47%) and 95% (95% CI, 90-97%), respectively.
By following SelectMDx, 34% of biopsies could be avoided. Hendriks et al. [35] evaluated the association between SelectMDx and multiparametric magnetic resonance imaging (mpMRI) outcomes in 172 patients from the study of Van Neste et al. [9] who had undergone this imaging technique. The AUC of SelectMDx with mpMRI was 0.83 (95% CI, 0.77-0.89), in comparison with 0.66 (95% CI, 0.58-0.74) of PSA and 0.65 (95% CI, 0.57-0.74) with PCA3. Pepe et al. [37] evaluated the diagnostic accuracy of SelectMDx for detecting csPCa in 125 patients in active surveillance. Post-DRE urine was obtained from 45 patients, and of them, 9 (20%) were re-classified as csPCa. Results with SelectMDx were: 55.6% of sensitivity, 65.8% of specificity, 27.8% of PPV, 87% of NPV, and 70.3% of diagnostic accuracy. Finally, Wysock et al. [38] compared a blood-based test, Four Kallikrein score, with SelectMDx in 114 men with elevated PSA. The AUC for SelectMDx was 0.67 (95% CI, 0.52-0.83) for detecting a csPCa. With a cut-off value of 7.5, sensitivity and specificity were 54.5% and 75.0%, respectively; however, with a value of 12 were 45.5% and 78.6%.
ExoDx Prostate IntelliScore
The EPI is an assay aimed at determining the expression of a 3-gene signature (Exosomal PCA3, the oncogenic gene fusion rearrangement TMPRSS2: ERG, and the SAM-pointed domaincontaining Ets transcription factor, SPDEF) in a first-catch urine sample [41]. In contrast to PCA3 and SelectMDx, EPI does not require a previous DRE. It provides a risk score, between 0 and 100. A score ≥15.6 is associated with an increased risk for grade group (GG) ≥2 PCa [41]. An early evaluation of EPI in 195 men with suspicious DRE and/or PSA levels revealed a good prediction of both any cancer and high-grade PCa [42]. The summary of the most representative results from studies using EPI is shown in Table 3. The combination of EPI with standard of care (SoC; PSA, age, race, and family history) showed a greater differentiation between high-grade (GS ≥7) and GS ≤6 biopsy, with an AUC of 0.80 (95% CI, 0.73-0.88) versus 0.67 (95% CI, 0.58-0.78) with SoC alone. The main validation study was conducted by McKiernan et al. [43], when compared EPI plus SoC versus SoC alone to discriminate GS ≥7 from GS 6 and benign PCa on initial biopsy. In an initial training cohort (255 men), the ability for detecting a high-grade PCa with EPI plus SoC was significantly higher than SoC alone in terms of AUC (0.77; 95% CI, 0.71-0.83 versus 0.66; 95% CI, 0.58-0.72).
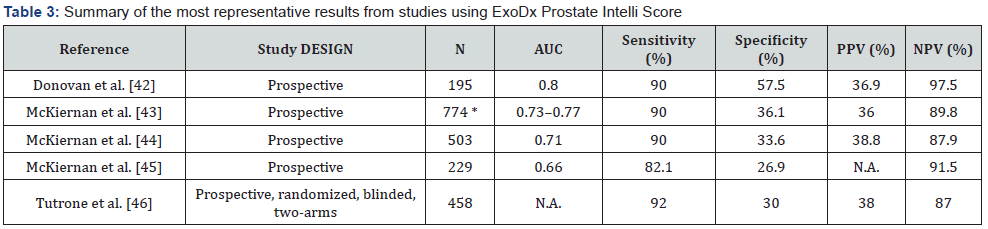
AUC: area under the curve; PPV: positive predictive value; NPV: negative predictive value; N.A.: not available
* Total patients (training and independent cohorts).
In a subsequent independent validation (519 patients), the AUC was also significantly superior with EPI plus SoC (0.73; 95% CI, 0.68-0.77 versus 0.63; 95% CI, 0.58-0.68) [42]. A total of 27% of biopsies could be avoided when used a cut-off value of ≥15.6. A further validation study with 503 patients with PSA level 2-10ng/ mL corroborated the greater accuracy of EPI plus SoC than SoC alone (AUC of 0.71; 95% CI, 0.66-0.76 versus 0.62; 95% CI, 0.57- 0.67) [44]. With a cut-off value of 15.6, up to 26% of biopsies could be avoided, missing 7% of GG ≥2 PCa, and achieving a NPV of 89%. When evaluated to 226 patients with a previous negative biopsy and underwent repeat biopsies, EPI alone showed an NPV of 92% and could avoid 26% of unnecessary biopsies [45]. The AUC with EPI alone (0.66; 95% CI, 0.55-0.78) was higher than the European Randomised Study of Screening for Prostate Cancer risk calculator (0.47; 95% CI, 0.36-0.58) and PSA (0.54; 95% CI, 0.43-0.66). In a recent prospective, randomized, blinded, twoarmed clinical study, Tutrone et al. [46] have evaluated the impact of EPI on the shared biopsy decision process between patients and urologists. All patients had an EPI, however in the EPI arm (n=458) patients received EPI results for the biopsy decisionmaking versus the control arm (n=484) in which patients did not receive the EPI results, and urologists proceeded with SoC. As a result, EPI arm was able to detect 30% more cases of high-grade PCa than the control arm, missing approximately 49% fewer cases than SoC.
Mi‑Prostate score
The MiPS is a test that evaluates the combination of PCA3 and TMPRSS2: ERG together with PSA or the Prostate Cancer Prevention Trial risk calculator in a post-DRE urine sample. In an early study, Salami et al. [47] revealed that the combination of PSA, TMPRSS2: ERG, and PCA3 provided an AUC of 0.88 (95% CI, 0.75-0.98) for predicting PCa on a subsequent biopsy. The summary of the most representative results from studies using MiPS is shown in Table 4. Cornu et al. [48] also demonstrated that PSA density, TMPRSS2: ERG, and PCA3 was independently associated with prostatic biopsy outcome (AUC of 0.73). Tomlins et al. [49] evaluated and validated risk models, including MiPS, for the detection of PCa and PCa (GS>6) on biopsy in a cohort of 1,225 men. The AUC for detecting high-grade PCa was 0.77 when using PSA, TMPRSS2: ERG, and PCA3; and 0.78 when using the Prostate Cancer Prevention Trial high-grade risk calculator (instead of PSA). Between 35% and 47% of biopsies could be avoided by using MiPS.
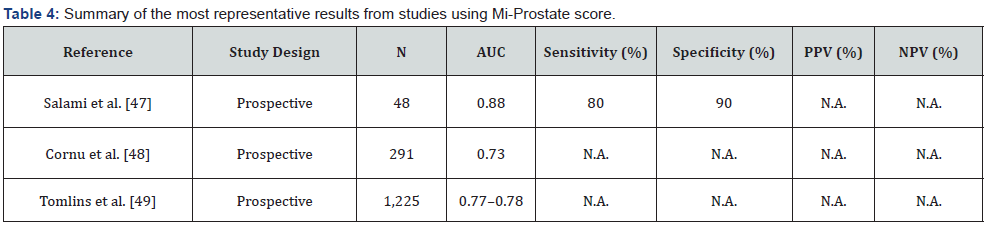
AUC: area under the curve; PPV: positive predictive value; NPV: negative predictive value; N.A.: not available
* Total patients (training and independent cohorts).
Clinical implications of urine biomarkers
Biomarkers have demonstrated to improve the diagnosis and discriminate a PCa with indolent course from those with unfavorable prognosis [7,50]. Post-DRE urine samples are a rich source of biomarkers for PCa [51]. The urine can be easily collected in a urology clinic, and it does not constitute an extra burden in the routine clinical practice. Therefore, post-DRE urine represents a minimally invasive option that provide enough material for an accurate diagnosis [51]. Moreover, novel urinebased tests have the potential to help physicians to determine the need for a biopsy, especially in men with PSA levels 4.0-10.0 ng/mL [13]. Urine biomarkers have demonstrated the ability to avoid unnecessary biopsies in clinical practice; up to 26% with PCA3, 53% with SelectMDx, 27% with EPI, and 47% with MiPS. Although there are no comparison studies between biomarkers so far, the diagnostic accuracy of SelectMDx seems to be higher than PCA3, in terms of sensitivity, specificity, PPV, NPV, and AUC. By contrast, PCA3 is also the only urine biomarkers that has also been associated with prognosis, by predicting the upgrading to GS ≥7 during active surveillance, or the tumor volume, extracapsular extension and positive surgical margins in prostatectomy specimens [52]. At the same time, the use in clinical practice of mpMRI has increased among men with a negative biopsy or even biopsy naïve [50]. The median NPV for a csPCa is about 88.1 [53]. Besides this, mpMRI is time-consuming, resource-intensive, and needs sufficient access to an MRI scanner [6]. Urine-based biomarkers can be used in clinical practice for the selection of candidates requiring a further evaluation, especially those with no mpMRI-visible lesions. However, the published literature points to a complementary effect of biomarkers and mpMRI6, achieving a NPV of up to 91% [32,50].
Caveats and Limitations with their Clinical Use
Although urine biomarkers provide worthwhile information about the disease, they should be considered as a complementary tool for the decision-making process, and not be used as a firstline for the diagnosis of PCa [54]. Diverse international Societies (such as the European Association of Urology, EAU, National Comprehensive Cancer Network, or the American Urological Association), have highlighted the lack of clear evidence about their benefit as primary screening test [55-57]. Yet, to avoid unnecessary biopsies, the EAU recommends further riskassessment (risk-calculator, imaging; strong recommendation), or an additional serum or urine-based test (weak recommendation) in asymptomatic men with normal DRE and PSA level 2-10ng/ mL [57]. ExoDx Prostate IntelliScore and MiPS assay are currently considered as investigational; thus, PCA3 and SelectMDx are the only urine-based tests that could be widely implemented in clinical practice for this setting.
It is necessary to indicate that most of patients (84%) from the main validation study of SelectMDx were prostate biopsy naïve [9]. PCA3 is the unique urine test that has been approved by the U.S.
Food and Drug Administration for patients who are candidates for repeat biopsy [58]. The EAU also suggest the complementary use of urine-based tests (PCA3 and SelectMDx) in the decisionmaking for men with an elevated risk of PCa with a prior negative biopsy and candidate for repeat biopsy [57]. The development and validation process of candidate urine biomarkers is long and requires large prospective studies, with a significant number of participants, and an adequate follow-up time. To date, the literature basing their diagnostic efficacy is extensive in the case of PCA3; however it is limited for SelectMDx, EPI, and MiPS. There are also no studies evaluating the implementation of each urine biomarker in routine clinical practice. Additional head-to-head comparison studies between biomarkers are also necessary. On the other hand, the price (PCA3 $300-500, SelectMDx $300, EPI $600) and availability of the biomarkers are factors that may limit their progress and implantation to clinical practice [59].
Future Perspectives
The increasing use of urine-based biomarkers for PCa will improve patient stratification and diagnosis, and thus allow more personalized therapies and follow-up schemes [60]. Further long-term, prospective, and real-world studies will complete and validate the available information about urine-based biomarkers, required to be implemented in routine clinical practice. Moreover, given the number of potential biomarkers, it is likely that the combination of biomarkers may perform better than the simple pairing [61]. In addition, urine biomarkers together with mpMRI has already demonstrated to provide a more accurate risk assessment of the disease [13].
Conclusion
A new era has emerged with the development of biomarkers having higher specificity than PSA for the detection of csPCa. The PCa biomarkers, especially urine-based ones, are useful tools able to improve the diagnosis and management of the disease. Urine biomarkers can help physicians to determine if a biopsy is required, mainly with PSA levels, especially in the grey zone (between 4.0 and 10.0ng/mL), and therefore avoid unnecessary biopsies. Moreover, they are valuable tools to differentiate csPCa from indolent disease, provide information about prognosis, and thus guide toward the most appropriate treatment. Further investigations will be the basis for the approval and widely use of these tests in routine clinical practice.
Declaration
i. Ethics approval and consent to participate: Not applicable
ii. Consent to publish: Not applicable
iii. Availability of data and materials: Data sharing is not applicable to this article as no datasets were generated or analysed during the current study
iv. Competing interests: Author declare no competing interest.
v. Funding: This manuscript was solely funded by Ferrer International S.A. Medical writing support was provided by Meisys (Madrid, Spain), and was funded by Ferrer International S.A.
vi. Authors’ Contributions: APL take responsibility for the integrity of the work as a whole: conception and designing, interpretation of the data, substantial revision and approving of the submitted version.
Acknowledgements: The author would like to express gratitude to Pablo Vivanco (PhD, Meisys) for helping in the elaboration of the manuscript.
References
- Bray F, Jacques Ferlay, Isabelle Soerjomataram, Rebecca L Siegel, Lindsey A Torre, et al. (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68(6): 394-424.
- Rawla P (2019) Epidemiology of prostate cancer. World J Oncol 10(2): 63-89.
- Roehrborn C G, Black L K (2011) The economic burden of prostate cancer. BJU int 108(6): 806-813.
- Shore N (2014) Management of early-stage prostate cancer. Am J Manag Care 20(12): 260-272.
- Hendriks R J, van Oort I M, Schalken J A (2017) Blood-based and urinary prostate cancer biomarkers: a review and comparison of novel biomarkers for detection and treatment decisions. Prostate Cancer Prostatic Dis 20(1): 12-19.
- Brönimann S, Pradere B, Karakiewicz P, Abufaraj M, Briganti A, et al. (2020) An overview of current and emerging diagnostic, staging and prognostic markers for prostate cancer. Expert Rev Mol Diagn 20(8): 841-850.
- Narayan V M (2020) A critical appraisal of biomarkers in prostate cancer. World J Urol 38(3): 547-554.
- Matulay J T, Wenske S (2018) Genetic signatures on prostate biopsy: clinical implications. Trans Can 7(6): 640-650.
- Van Neste L, Rianne J Hendriks, Siebren Dijkstra, Geert Trooskens, Erik B Cornel, et al. (2016) Detection of high-grade prostate cancer using a urinary molecular biomarker-based risk score. European Urology 70(5): 740-748.
- Welch H G, Albertsen P C (2009) Prostate cancer diagnosis and treatment after the introduction of prostate-specific antigen screening: 1986-2005. J Natl Cancer Inst 101(19): 1325-1329.
- Epstein J I, Feng Z, Trock B J, Pierorazio P M (2012) Upgrading and downgrading of prostate cancer from biopsy to radical prostatectomy: incidence and predictive factors using the modified Gleason grading system and factoring in tertiary grades. Eur Urol 61(5): 1019-1024.
- Goodman M, Kevin C Ward, Adeboye O Osunkoya, Milton W Datta, Daniel Luthringer, et al. (2012) Frequency and determinants of disagreement and error in gleason scores: a population-based study of prostate cancer. Prostate 72(13): 1389-1398.
- Kohaar I, Petrovics G, Srivastava S (2019) A rich array of prostate cancer molecular biomarkers: Opportunities and challenges. Int J Mol Sci 20(8): 1813.
- Hessels D, Jacqueline M T Klein Gunnewiek, Inge van Oort, Herbert F M Karthaus, Geert J L van Leenders, et al. (2003) DD3(PCA3)-based molecular urine analysis for the diagnosis of prostate cancer. Eur Urol 44 (1): 8-16.
- Marks L S, Yves Fradet, Ina Lim Deras, Amy Blase, Jeannette Mathis, Sheila M J Aubin, et al. (2007) PCA3 molecular urine assay for prostate cancer in men undergoing repeat biopsy. Urology 69(3): 532-535.
- Deras I L, Sheila M J Aubin, Amy Blase, John R Day, Seongjoon Koo, et al. (2008) PCA3: a molecular urine assay for predicting prostate biopsy outcome. The Journal of Urology 179(4): 1587-1592.
- Haese A, Alexandre de la Taille, Hendrik van Poppel, Michael Marberger, Arnulf Stenzl, et al. (2008) Clinical utility of the PCA3 urine assay in European men scheduled for repeat biopsy. Eur Urol 54(5): 1081-1088.
- Wang R, Chinnaiyan A M, Dunn R L, Wojno K J, Wei J T (2009) Rational approach to implementation of prostate cancer antigen 3 into clinical care. Cancer 115(17): 3879-3886.
- Aubin S M, Jennifer Reid, Mark J Sarno, Amy Blase, Jacqueline Aussie, Harry Rittenhouse, et al. (2010) PCA3 molecular urine test for predicting repeat prostate biopsy outcome in populations at risk: validation in the placebo arm of the dutasteride REDUCE trial. J Urol 184(5): 1947-1952.
- Roobol M J, Fritz H Schröder, Pim van Leeuwen, Tineke Wolters, Roderick C N van den Bergh, et al. (2010) Performance of the prostate cancer antigen 3 (PCA3) gene and prostate-specific antigen in prescreened men: exploring the value of PCA3 for a first-line diagnostic test. European Urology 58(4): 475-481.
- de la Taille, Jacques Irani, Markus Graefen, Felix Chun, Theo de Reijke, et al. (2011) Clinical evaluation of the PCA3 assay in guiding initial biopsy decisions. J Urol 185(6): 2119-2125.
- Crawford E D, Kyle O Rove, Edouard J Trabulsi, Junqi Qian, Krystyna P Drewnowska, et al. (2012) Diagnostic performance of PCA3 to detect prostate cancer in men with increased prostate specific antigen: a prospective study of 1,962 cases. J Urol 188(5): 1726-1731.
- Gittelman M C, Bernard Hertzman, James Bailen, Thomas Williams, Isaac Koziol, et al. (2013) PCA3 molecular urine test as a predictor of repeat prostate biopsy outcome in men with previous negative biopsies: a prospective multicenter clinical study. J Urol 190(1): 64-69.
- Tombal B, Gerald L Andriole, Alexandre de la Taille, Paolo Gontero, Alexander Haese, et al. (2013) Clinical judgment versus biomarker prostate cancer gene 3: which is best when determining the need for repeat prostate biopsy? Urology 81(5): 998-1004.
- Rubio Briones J, J Casanova, R Dumont, L Rubio, A Fernandez Serra, et al. (2014) Optimizing prostate cancer screening; prospective randomized controlled study of the role of PSA and PCA3 testing in a sequential manner in an opportunistic screening program. Actas Urol Esp 38(4): 217-223.
- Wei J T, Ziding Feng, Alan W Partin, Elissa Brown, Ian Thompson, et al. (2014) Can urinary PCA3 supplement PSA in the early detection of prostate cancer? J Clin Oncol 32(36): 4066-4072.
- Ruiz Aragón J, Márquez Peláez S (2010) Assessment of the PCA3 test for prostate cancer diagnosis: a systematic review and meta-analysis. Actas Urol Esp 34(4): 346-355.
- Hu B, Yang H, Yang H (2014) Diagnostic value of urine prostate cancer antigen 3 test using a cutoff value of 35 μg/L in patients with prostate cancer. Tumour biol 35(9): 8573-8580.
- Luo Y, Gou X, Huang P, Mou C (2014) The PCA3 test for guiding repeat biopsy of prostate cancer and its cut-off score: a systematic review and meta-analysis. Asian J Androl 16(3): 487-492.
- Cui Y, Wenzhou Cao, Quan Li, Hua Shen, Chao Liu, et al. (2016) Evaluation of prostate cancer antigen 3 for detecting prostate cancer: a systematic review and meta-analysis. Sci Rep 6: 25776.
- Xue W J, Ying X L, Jiang J H, Xu Y H (2014) Prostate cancer antigen 3 as a biomarker in the urine for prostate cancer diagnosis: a meta-analysis. J Cancer Res Ther 10: 218-221.
- Rubio Briones J, J Casanova, F Martínez, J L Domínguez Escrig, A Fernández Serra, et al. (2017) PCA3 as a second-line biomarker in a prospective controlled randomized opportunistic prostate cancer screening programme. PCA3 como biomarcador de segunda línea en un programa de screening oportunista prospectivo, aleatorizado y controlado. Actas Urol Esp 41(5): 300-308.
- Jiang Z, Zhao Y, Tian Y (2019) Comparison of diagnostic efficacy by two urine PCA3 scores in prostate cancer patients undergoing repeat biopsies. Minerva Urol Nefrol 71(4): 373-380.
- Haese A, Geert Trooskens, Sandra Steyaert, Daphne Hessels, Michael Brawer, et al. (2019) Multicenter optimization and validation of a 2-Gene mRNA urine test for detection of clinically significant prostate cancer before initial prostate biopsy. J Urol 202(2): 256-263.
- Hendriks R J, Marloes M G van der Leest, Siebren Dijkstra, Jelle O Barentsz, Wim Van Criekinge, et al. (2017) A urinary biomarker-based risk score correlates with multiparametric MRI for prostate cancer detection. Prostate 77(14): 1401-1407.
- Tilki D, Daphne Hessels, Geert Trooskens, Susan Mulders, Michael Brawer, et al. (2019) Validation of a two-gene mRNA urine test for detection of high-grade prostate cancer in German men. J Clin Oncol 37(7).
- Pepe P, Dibenedetto G, Pepe L, Pennisi M (2020) Multiparametric MRI versus SelectMDx accuracy in the diagnosis of clinically significant PCa in men enrolled in active surveillance. In Vivo 34(1): 393-396.
- Wysock J S, Becher E, Persily J, Loeb S, Lepor H (2020) Concordance and performance of 4Kscore and SelectMDx for informing decision to perform prostate biopsy and detection of prostate cancer. Urology 141: 119-124.
- Rubio Briones J, Angel Borque Fernando, Luis M Esteban, Juan M Mascarós, Miguel Ramírez Backhaus, et al. (2020) Validation of a 2-gene mRNA urine test for the detection of ≥GG2 prostate cancer in an opportunistic screening population. Prostate 80(6): 500-507.
- Busetto G M, Francesco Del Giudice, Martina Maggi, Ferdinando De Marco, Angelo Porreca, et al. (2020) Prospective assessment of two-gene urinary test with multiparametric magnetic resonance imaging of the prostate for men undergoing primary prostate biopsy. World J Urol 39(6): 1869-1877.
- Becerra M F, Atluri V S, Bhattu A S, Punnen S (2020) Serum and urine biomarkers for detecting clinically significant prostate cancer. Urol Oncol 1078-1439(20): 30059-30064.
- Donovan M J, M Noerholm, S Bentink, S Belzer, J Skog, et al. (2015) A molecular signature of PCA3 and ERG exosomal RNA from non-DRE urine is predictive of initial prostate biopsy result. Prostate Cancer Prostatic Dis 18(4): 370-375.
- McKiernan J, Michael J Donovan, Vince O'Neill, Stefan Bentink, Mikkel Noerholm, et al. (2016) A novel urine exosome gene expression assay to predict high-grade prostate cancer at initial biopsy. JAMA Oncol 2(7): 882-889.
- McKiernan J, Michael J Donovan, Eric Margolis, Alan Partin, Ballentine Carter, et al. (2018) A prospective adaptive utility trial to validate performance of a novel urine exosome gene expression assay to predict high-grade prostate cancer in patients with prostate-specific antigen 2-10ng/ml at initial biopsy. Eur Urol 74 (6): 731-738.
- McKiernan J, Mikkel Noerholm, Vasisht Tadigotla, Sonia Kumar, Phillipp Torkler, et al. (2020) A urine-based exosomal gene expression test stratifies risk of high-grade prostate cancer in men with prior negative prostate biopsy undergoing repeat biopsy. BMC Urology 20(1): 138.
- Tutrone R, Michael J Donovan, Phillipp Torkler, Vasisht Tadigotla, Tom McLain, et al. (2020) Clinical utility of the exosome based ExoDx Prostate(IntelliScore) EPI test in men presenting for initial Biopsy with a PSA 2-10 ng/mL. Prostate Cancer Prostatic Dis 23(4): 607-614.
- Salami S S, Folke Schmidt, Bharathi Laxman, Meredith M Regan, David S Rickman, et al. (2013) Combining urinary detection of TMPRSS2:ERG and PCA3 with serum PSA to predict diagnosis of prostate cancer. Urol Oncol 31(5): 566-571.
- Cornu J N, Cancel Tassin G, Egrot C, Gaffory C, Haab F, et al. (2013) Urine TMPRSS2: ERG fusion transcript integrated with PCA3 score, genotyping, and biological features are correlated to the results of prostatic biopsies in men at risk of prostate cancer. Prostate 73(3): 242-249.
- Tomlins S A, John R Day, Robert J Lonigro, Daniel H Hovelson, Javed Siddiqui, et al. (2016) Urine TMPRSS2: ERG Plus PCA3 for Individualized Prostate Cancer Risk Assessment. Eur Urol 70(1): 45-53.
- Carneiro A, Paulo Priante Kayano, Álan Roger Gomes Barbosa, Marcelo Langer Wroclawski, Carolina Ko Chen, et al. (2018) Are localized prostate cancer biomarkers useful in the clinical practice? Tumour Biol 40(9): 1010428318799255.
- Rigau M, Mireia Olivan, Marta Garcia, Tamara Sequeiros, Melania Montes, et al. (2013) The present and future of prostate cancer urine biomarkers. Int J Mol Sci 14(6): 12620-12649.
- Fujita K, Nonomura N (2018) Urinary biomarkers of prostate cancer. Int J Urol 25(9): 770-779.
- Moldovan P C, Thomas Van den Broeck, Richard Sylvester, Lorenzo Marconi, Joaquim Bellmunt, et al. (2017) What is the negative predictive value of multiparametric magnetic resonance imaging in excluding prostate cancer at biopsy? A systematic review and meta-analysis from the European Association of Urology Prostate Cancer Guidelines Panel. Eur Urol 72(2): 250-266.
- Nassir A M (2020) A piece in prostate cancer puzzle: Future perspective of novel molecular signatures. Sau J Biol Sci 27(4): 1148-1154.
- American Urological Association (2018) Early Detection of Prostate Cancer.
- Mohler J L, Emmanuel S Antonarakis, Andrew J Armstrong, Anthony V D Amico, Brian J Davis, et al. (2019) Prostate Cancer, Version 2.2019, NCCN Clinical Practice Guidelines in Oncology. J Natl Compr Canc Netw 17(5): 479-505.
- European Association of Urology (2020) Prostate cancer.
- US Food and Drug Administration. Summary of safety and effectiveness data (SSED). Progensa PCA3 Assay.
- Zapała P, Dybowski, B, Poletajew, S, Radziszewski P (2018) What can be expected from prostate cancer biomarkers? A clinical perspective. Urol Internat 100(1): 1-12.
- Couñago F, Fernando López Campos, Ana Aurora Díaz Gavela, Elena Almagro, Esaú Fenández Pascual, et al. (2020) Clinical applications of molecular biomarkers in prostate cancer. Cancers 12(6): 1550.
- Wei J T (2015) Urinary biomarkers for prostate cancer. Curr Opin Urol 25(1): 77-82.






























