Co-occurrence of SARS-CoV-2 and Respiratory Pathogens in the Frail Elderly
Alan J Wolfe1*, David Baunoch3*, Dakun Wang2, Ryan Gnewuch3, Xinhua Zhao2, Thomas Halverson1, Patrick Cacdac3, Shuguang Huang2, Trisha Lauterbach4 and Natalie Luke3
1Department of Microbiology and Immunology, Loyola University Chicago, Maywood, IL
2Stat4ward, 711 Parkview Dr., Gibsonia, PA 15044
3Pathnostics, 17661 Cowan, Irvine, CA 92614
4Capstone Laboratory, 8601 Dunwoody Pl #444, Sandy Springs, GA 30350
Submission: June 24, 2020;Published: June 30, 2020
*Corresponding author:David Baunoch, Pathnostics, 17661 Cowan, Irvine, CA 92614 dbaunoch@pathnostics.com Alan J Wolfe, Department of Microbiology and Immunology, Loyola University Chicago, Maywood, IL awolfe@luc.edu Drs. Wolfe and Baunoch contributed equally to this article
How to cite this article: Wolfe AJ, Baunoch D, Wang D, Gnewuch R, Zhao Z, et al. Co-occurrence of SARS-CoV-2 and Respiratory Pathogens in the Frail Elderly. JOJ Urology & Nephrology, 2020; 7(3): 555711. DOI: 10.19080/JOJUN.2020.07.555711
Abstract
Background: Elderly SARS-CoV-2 participants are associated with higher hospitalization and mortality. Co-infection is critical in the severity of respiratory diseases. It is largely understudied for SARS-CoV-2.
Methods: Between March 24th and April 27th, 2020, nasopharyngeal and oropharyngeal swabs from 3,348 participants from nursing homes and assisted living facilities in 22 states in the US were tested by Capstone Healthcare for SARS-CoV-2, 24 other respiratory viruses, and 8 respiratory bacteria. Total nucleic acid was extracted with MagMAXTM Viral/Pathogen Ultra nucleic acid isolation kit. SARS-Co-V-2 was detected with the CDC 2019-novel coronavirus (2019-nCoV) diagnostic panel. Total nucleic acid was pre-amplified before analysis for other respiratory pathogens with Taqman OpenArrayTM Respiratory Tract Microbiota Plate.
Results: The participants mean age was 76.9 years. SARS-CoV-2 was detected in 1,413 participants (42.2%). Among them, 1,082 (76.6%) and 737 (43.7%) participants were detected with at least one bacterium or another virus, respectively. SARS-CoV-2-positive participants were more likely to have bacterial co-occurrences (76.6%) than SARS-CoV-2-negative participants (70.0%) (p<0.0001). The most common co-occurring bacteria were Staphylococcus aureus and Klebsiella pneumonia, detected in 55.8% and 40.1% SARS-CoV-2-positive participants, respectively. Staphylococcus aureus was associated with SARS-CoV-2, with higher detection rates in SARS-CoV-2-positive participants (55.8%) than SARS-CoV-2-negative participants (46.2%) (p<0.0001). Human herpes virus 6 (HHV6) also was common and associated with SARS-CoV-2, with higher detection rates in SARS-CoV-2-positive participants (26.6%) than SARS-CoV-2-negative participants (19.1%) (p<0.0001).
Conclusions: SARS-CoV-2-positive participants are more likely to be positive for certain respiratory bacteria and viruses. This observation may help explain high hospitalization and mortality rates in older people.
Keywords: SARS-CoV-2, COVID19, Respiratory Pathogens, Nursing Homes, Assisted Living Facilities, Long Term Care
Introduction
As of May 12, 2020, the SARS-CoV-2 pandemic has resulted in more than 3.5 million confirmed cases and 250,000 deaths [1]. In the US alone, there are more than 1.3 million confirmed cases, with an overall cumulative hospitalization rate of 50.3 per 100,000 [2]. In the US, the highest rates of hospitalization is in people 65 years and older (162.2 per 100,000), who contribute 80% of the 78,000 deaths [3]. In the ongoing battle against SARS-CoV-2, much effort has focused on screening and testing effective anti-viral drugs and developing vaccines. In contrast, co-infection is largely understudied. This is problematic, as co-infection has proven to be critical in the severity of other respiratory diseases and significantly impacts patient clinical management, infection control, and antimicrobial stewardship programs. Secondary bacterial pneumonia, a common form of co-infection, was a predominant cause of death in the 1918-1919 and the 1957-1958 influenza pandemics [4] Bacterial co-infection also was reported in approximately 20% of the 2009 H1N1 influenza participants and was associated with poor outcome [5,6]. Thus, it is reasonable to suspect significant levels of co-infection in the current pandemic. The level of co-infection is particularly important as it is reported that broad-spectrum empirical antibiotics are frequently prescribed (72%) in SARS-CoV-2 participants despite a paucity of evidence of bacterial co-infection [7]. A better understanding of co-infection levels in participants with SARS-CoV-2 will facilitate better evaluation of patient prognosis and allow development of more effective clinical management plans [8].
Co-infection with coronavirus, especially with its three most prominent members, SARS-CoV-1, MERS and SARS-CoV-2, has been reported only infrequently, due in part to the novel nature of this viral group [7]. For SARS-CoV-2, co-infection with bacteria, other respiratory viruses, and/or fungi has been described, with a mean co-infection rate of approximately 8% across all studies [8]. For example, Zhou et al. reported 15% secondary lung infection that existed in 50% of all fatal cases [9]. Chen et al. reported 4% co-infection of bacteria and fungi with SARS-CoV-2 [10]. However, most studies were case reports or based on scattered available clinical information, and not designed to systematically evaluate the prevalence or the characteristics of co-infections. Two studies were exceptions. Lin et al. used multiplex reverse transcription– polymerase chain reaction (RT-PCR) to simultaneously detect SARS-CoV-2 and 14 other respiratory viruses in sputum, nasal or throat swabs from 186 highly suspected COVID-19 cases between January 20th and February 1st, 2020 obtained from multiple sites in the city of Shenzhen, China. Among the 92 SARS-CoV-2-postive participants, co-infection with other viruses was only identified in 6 participants (3.2%), whereas 18 out of the 94 SARS-CoV- 2-negatvie participants (9.7%) had co-infections. Bacterial coinfection was not tested [11]. In a more recent study, Kim et al. analyzed 1217 nasopharyngeal swabs from 1206 symptomatic participants (with cough, fever, and/or dyspnea) received between March 3rd and March 27th, 2020 from multiple sites in northern California. These authors also used RT-PCR technology to test for SARS-CoV-2 and a panel of other pathogens, which included 12 other respiratory viruses or viral groups, as well as 2 bacteria species. A total of 116 of the 1217 specimens (9.5%) were identified as positive for SARS-CoV-2. They reported no statistically significant difference in co-infection rates with other respiratory pathogens in participants positive [24 of 116 (20.7%)] or negative [294 of 1101 (26.7%)] for SARS-Co-2 (difference 6.0%, 95% CI, -2.3% to 14.3%) [12].
Nursing homes and assisted living facilities are at high risk for SARS-CoV-2. Their residents are older adults living in close quarters; most have underlying chronic medical conditions. In this study, we used RT-PCR technology to investigate the cooccurrence of SARS-Co-V-2 with a large panel of other respiratory pathogens in 3,348 participants from nursing homes and assisted living facilities in the US.
Methods
Participant and specimens
Capstone Healthcare tested a total of 3,348 adult participants from 156 nursing homes and assisted living facilities from 22 states in the US for both SARS-CoV-2 and other respiratory pathogens. Specimens included in the study were collected via nasopharyngeal and oropharyngeal swabs between March 24th and April 27th, 2020, transported to Capstone Healthcare in molecular transport medium, and processed as described below. The study was exempted from Institutional Review Board (IRB) review by Western IRB.
Total nucleic acid extraction
Total nucleic acid was extracted in the BSL-3 laboratory at Capstone Healthcare with MagMAXTM Viral/Pathogen Ultra nucleic acid isolation kit, according to the manufacturer’s instruction (Cat. No. A42356, Thermo Fisher, Waltham, MA).
Detection of SARS-Co-V-2
Detection of SARS-Co-V-2 was performed with the CDC 2019-Novel Coronavirus (2019-nCoV) Real-Time RT-PCR Diagnostic panel according to CDC’s instruction (Cat. No. 2019-nCoVEUA-01, CDC, Atlanta GA). The one-step RT-PCR was performed with the total extracted nucleic acid as the template on a QuantStudio™12K Flex Real-time PCR system. SARS-CoV-2 status was determined based on RT-PCR results from 2019-nCoV markers N1 and N2, and RNase P, an extraction control, according to the CDC’s instruction.
Detection of other respiratory pathogens
The extracted nucleic acid was pre-amplified on a Veriti™ Thermal Cycler before analysis for presence of other respiratory pathogens on the QuantStudio™12K Flex Real-time PCR system with Taqman OpenArrayTM Respiratory Tract Microbiota Plate (Cat. No. A41237, Thermo Fisher, Waltham, MA), according to the manufacturer’s instructions outlined in the application guide for Respiratory Tract Microbiota Profiling Experiments (Pub. Num. MAN0017952). The OpenArrayTM Respiratory Tract Microbiota Plates included 24 respiratory viruses and 8 bacteria measured in triplicate for each specimen. The viruses include influenza A/B, influenza A/H1-2009, influenza A/H3, human herpesvirus 3/4/5/6, parainfluenza virus type 1/2/3/4, adenovirus, rhinovirus, human bocavirus, coronavirus HKU, coronavirus OC43, coronavirus NL63, coronavirus 229E, human metapneumovirus, respiratory syncytial virus A/B, human enterovirus, human enterovirus D68. The bacteria included Haemophilus influenzae, Legionella pneumophila, Klebsiella pneumonia, Staphylococcus aureus, Mycoplasma pneumonia, Bordetella bronchiseptica/ parapertussis/ Bordetella pertussis, Streptococcus pneumonia, and Chlamydophila pneumonia.
Statistical Analysis
Participant and facility characteristics were described by frequency and percentages. SARS-CoV-2 detection rate among the full sample, and the co-occurrence of respiratory bacteria and other respiratory viruses among SARS-CoV-2-positive participants were calculated and compared by participant characteristics using chi-square test, separately. The overall associations between SARS-CoV-2 with respiratory bacteria and other respiratory viruses were reported by crosstab and chi-square test. The three-way detection rate of SARS-CoV-2, respiratory bacteria, and other respiratory viruses were depicted in a bar chart. The infection rate of each individual respiratory bacterium and other respiratory virus was calculated overall and compared by SARS-CoV-2-positive/negative status. Multivariate logistic regression model with robust variance at facility level to account for clustering of participants within the same facility was used to assess whether SARS-CoV-2 was independently associated with the co-occurrence of respiratory bacteria. Lastly, sensitivity analyses were run by excluding the participants with multiple tests to assess the robustness of the study findings. The analyses were performed using SAS version 9.4 and StataSE 16.
Results
Participants characteristics
Capstone Healthcare tested a total of 3,586 nasopharyngeal and oropharyngeal specimens from 3,348 adult participants; 3,115 had one test, 228 had 2 tests, and 5 had 3 tests. Among those 233 participants with multiple tests, only 23 had results changed, including 8 participants changed from positive to negative and 15 changed from negative to positive. For the participants with multiple tests, results from the first test were included in the analysis with the following exception: for the 15 participants whose SARS-CoV-2 results changed from negative to positive, the second test set, associated with the positive SARS-CoV-2 result, was included in the analysis.
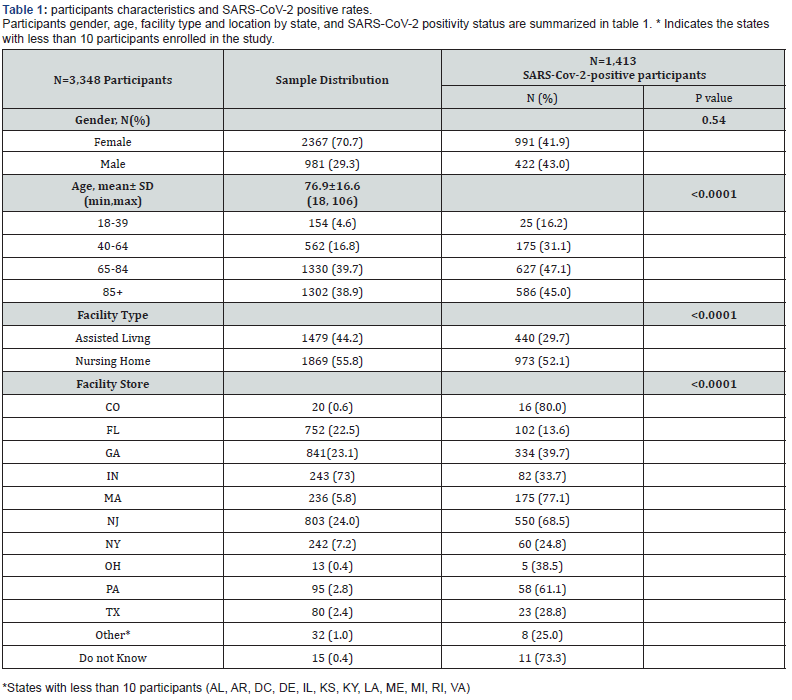
Table 1 provides detailed characteristics of the study subjects. All participants were from either nursing homes (55.8%) or assisted living facilities (44.2%). The mean age was 76.9 years old, with 78.6% subjects being 65 years or older. There were 2,367 (70.7%) females and 981 (29.3%) males. Participants were mainly from three states, FL (22.5%), GA (25.1%), and NJ (24.0%).
SARS-CoV-2 infection status
Of the 3,348 participants, 1,413 (42.2%) tested positive for SARS-CoV-2, with no significant difference between males and females (p=0.54) (Table 1). In contrast, older participants (≥ 65) tested positive more frequently than younger ones (p<0.0001). Test results also differed by facility type with more positive results from nursing homes than from assisted living facilities (<0.0001). They also differed by location (state) of the facility (<0.0001).
Co-occurrence of SARS-CoV-2 and other pathogens
Table 2 lists the co-occurrence of respiratory bacteria and other respiratory viruses with SARS-CoV-2, stratified by gender, age, facility type and facility location (by state). In total, of the 1,413 SARS-CoV-2-positive participants, 1,082 (76.6%) and 734 (52.2%) were detected with at least one respiratory bacterium or other respiratory virus, respectively. participants age (p=0.005) and facility type (<0.0001) were associated with co-occurrence of SARS-CoV-2 with respiratory bacteria, but not with other respiratory viruses. Facility location was more strongly associated with respiratory bacteria (p<0.0001) than other respiratory viruses (p=0.019).
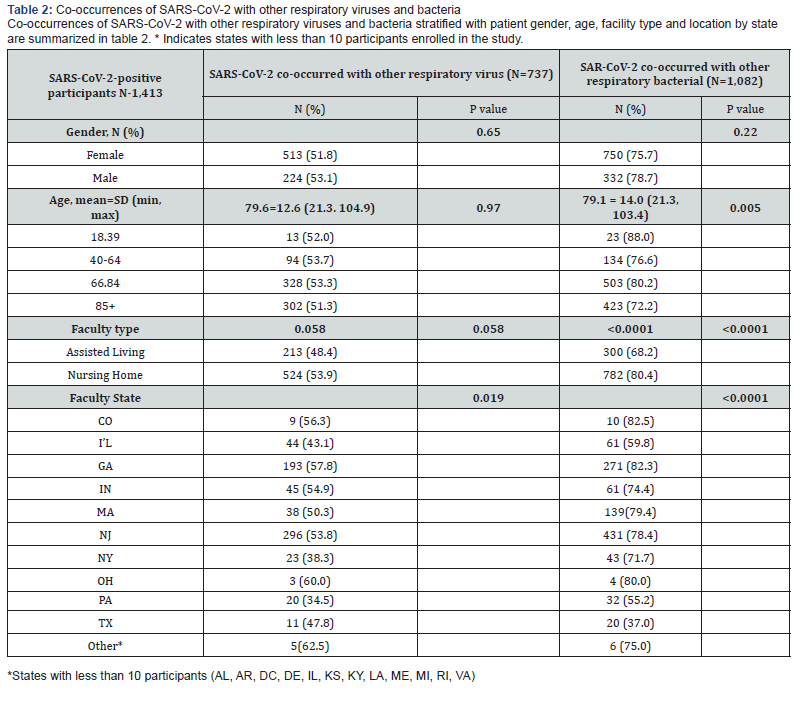
Table 3 compares SARS-CoV-2-positive and –negative participants. SARS-CoV-2-positive participants were more likely to test positive for at least one respiratory bacterium (1,082/1,413 [76.6%]) than SARS-CoV-2-negative participants (1,354/1,935 [70.0%]) (p<0.0001). No association was observed for other respiratory viruses (737/1,413 [52.2%] vs. 949/1,935 [49.0%] for SARS-CoV-2-positive and -negative participants, respectively [p=0.075]). Multivariate analysis showed that SARSCoV- 2 status was an independent factor associated with bacterial co-infection, regardless of participant age, gender, facility type and state location (odds ratio=1.26, 95% confidence interval: 1.04-1.54; p=0.02). More than 600 participants tested positive for SARS-CoV-2, at least one respiratory bacterium, and at least one other respiratory virus, which translates to 42.7% (603/1,413) co-occurrence with both a respiratory bacterium and another respiratory virus in SARS-CoV-2-positive participants (Figure 1).
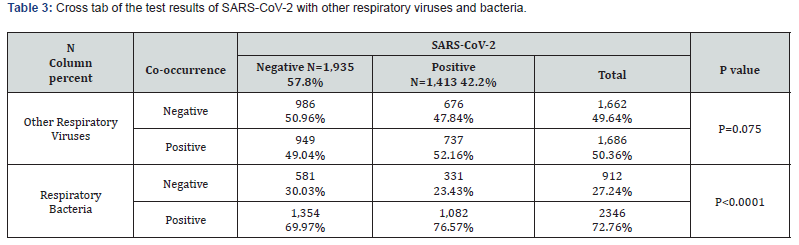
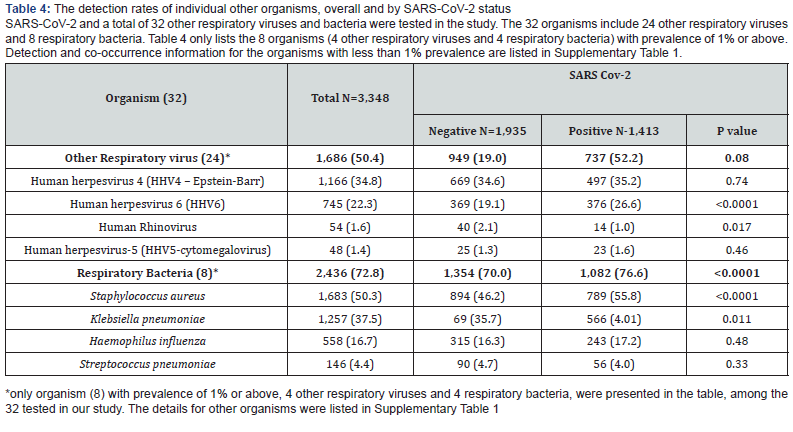
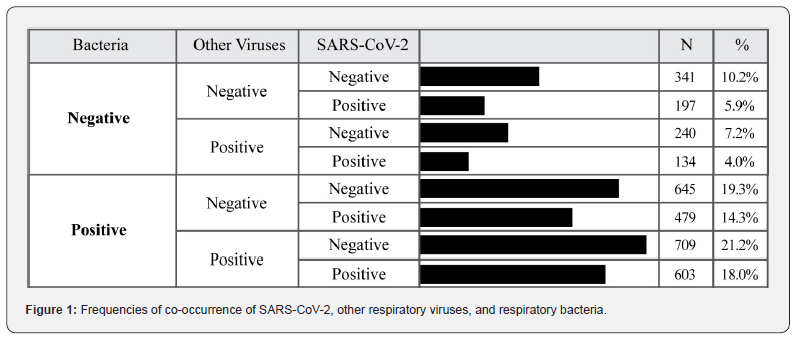
Table 4 shows the co-occurrence data for SARS-CoV-2, respiratory bacteria, and other respiratory viruses. All 24 other respiratory viruses and 8 bacteria were detected in at least one participants, with two exceptions: Chlamydophila pneumonia and Legionella pneumophila. The most common co-occurring bacterial species were Staphylococcus aureus, Klebsiella pneumoniae and Haemophilus influenza, detected in 55.8%, 40.1% and 17.2% of SARS-CoV-2-positive participants, respectively. More importantly, S. aureus was associated with infection of SARS-CoV-2, with detection rates of 55.8% and 46.2% (p<0.0001) in SARS-CoV-2- positive and -negative participants, respectively. K. pneumoniae was also associated with SARS-CoV-2. None of the other bacteria were associated with SARS-CoV-2. The most common co-occurring viruses were human herpesvirus 4 (HHV4 - Epstein-Barr Virus) and human herpesvirus 6 (HHV6). The majority of the viral detections (22/24, 91.7%) were not associated with detection of SARS-CoV-2, with the exception of HHV6 and rhinovirus. The detection rates of HHV6 in SARS-CoV-2-positive and SARSCoV- 2-negative participants were 26.6% vs. 19.1% (p<0.0001), respectively. A small number of participants (0.1% to 0.4%) tested positive for both influenza (A/B or other subtypes) and SARSCoV- 2. Co-occurrence of other coronaviruses with SARS-CoV-2, such as human coronavirus OC43 (0.1%), HKU1 (0.5%), 229E (0.0%), and NL63 (0.5%), were also low (Supplementary Table 1). The results were very similar from the sensitivity analysis by excluding those 233 participants with multiple tests.
Discussion
The SARS-CoV-2 pandemic is a major challenge to healthcare systems. Proper diagnosis and management of elderly people are critical due to their high hospitalization and mortality rates. Experiences from past influenza pandemics demonstrated the important role of bacterial co-infection in the severity of respiratory diseases [4-6]. Our observation that SARS-CoV-2- positive participants were more likely also to be positive for certain respiratory bacteria and viruses may help explain high hospitalization and mortality rates in older participants. The rates of co-occurrence in our study were considerably higher than those from the Lin et al. and Kim et al. studies. This difference may be due to several reasons. First, more pathogens were included in this study. A total of 24 respiratory viruses and 8 respiratory bacteria were tested in this study, compared with 12 viruses in the former study, and 12 viruses and viral groups, and 2 bacteria (Chlamydophila pneumoniae and Mycoplasma pneumoniae) in the latter study. In this study, Chlamydophila pneumoniae was not detected in any participants and Mycoplasma pneumoniae was only detected in 3 participants, and none of these participants were SARS-CoV-2-positive. Second, the participants included in our study were much older, with mean age of 76.9 years and only 21.4% younger than 65 years. The mean age in the Kim et al. study was 45.1 years. Likewise, 78% of participants were <65 years in the Lin et al. study. Age, suboptimal medical condition and immune-compromised status, congregated living environment and other factors may make these elderly participants more susceptible to respiratory infections. Third, all the participants in our study were from nursing homes and assisted living facilities. Fourth, the specimens were collected primarily in April for our study, but January and March for the other two studies [10,11].
The high bacterial co-occurrence rates in our study may support the clinical use of antibiotics in the management of elderly participants with SARS-CoV-2. The majority of study participants were positive for at least one respiratory bacterium, and the detection rate was higher in SARS-CoV-2- positive participants. The most commonly detected bacterial species in SARS-CoV-2-postive participants was S. aureus, which also was positively associated with SARS-CoV-2 infection. This Gram-positive bacterium is commonly carried on the skin or in the nose of healthy people [13]. It also is a recognized cause of influenza-associated communityacquired pneumonia [14, 15]. It was a relatively minor cause of 1918 influenza fatalities but was predominant in the 1957–1958 influenza pandemic [4,14,16,17]. It also was one of the most common bacteria co-infected with the 2009 H1N1 influenza virus, in addition to Streptococcus pneumoniae and Streptococcus pyogenes [6]. In the current study, S. aureus was detected in 56.2% of SARS-CoV-2-positive participants, compared with 46.2% of SARS-CoV-2-negative participants (10% difference, p<0.0001). Clinically, some S. aureus strains are antibiotic-resistant, such as methicillin-resistant S. aureus (MRSA), and vancomycin-resistant S. aureus (VRSA). These findings may help increase the awareness of the potential presence of hard-to-treat bacterial infections, such as MRSA or VRSA in individuals with SARS-CoV-2. The other two commonly detected bacteria were the Gram-negatives Klebsiella pneumonia and Haemophilus influenza, but the former was weakly associated, whereas the latter was not associated.
The high rate of S. aureus co-occurrence with SARS-CoV-2 may support the clinical use of broad-spectrum antibiotics in SARSCoV- 2 participants. Results from this study also provide pertinent information for healthcare facilities to adapt their antibiotic stewardship programs, in response to the pandemic. Whether cooccurrences with other pathogens result in clinically relevant coinfections, and whether they indeed contribute to the severity and the outcome of SARS-CoV-2 are important and intriguing clinical questions that warrant further investigations.
Results from the current study also confirmed previous reports that suggest the importance of testing participants with respiratory infectious diseases with both SARS-CoV-2 and other respiratory viruses, and that identification of other respiratory virus cannot exclude SARS-CoV-2, and vise versa. The cooccurrence rates of influenza viruses were low in both SARS-CoV- 2-positive and - negative participants, presumably due to the fact that the specimen collection period occurred at the end of the flu season and lockdown policies were commonly in place due to the pandemic. The two most commonly detected viruses identified in our study were HHV4 (Epstein-Barr virus) and HHV6. Both viruses have been identified as risk factors for cognitive deterioration and progression to Alzheimer’s disease in elderly persons [18]. Many people become infected with HHV4 and HHV6 in childhood. The viruses then became latent, during which time they may replicate in the salivary glands and be secreted in saliva without inducing any obvious pathology. In immune-compromised persons, reactivation of virus from latency or reinfection may occur [19,20]. However, in this study, only HHV6, not HHV4, was associated with SARS-CoV-2 positivity, which may be due to selective reactivation of HHV6 in SARS-CoV-2-positive participants. HHV6 is known to have broad tropism [21]. This virus may be present, whereas HHV4 may not, in pneumocytes and/or angiotensin-converting enzyme 2 (ACE2)-positive cells that are targeted by the SARSCoV- 2 virus. The exact underlying mechanisms of the selective association with SARS-CoV-2 of HHV6, but not HHV4, and the clinical significance of this co-occurring virus in disease severity or response to potential treatment in participants with SARSCoV- 2 should be investigated further.
In summary, this co-occurrence study of SARS-CoV-2 and 32 other respiratory pathogens in a large cohort of participants from multiple nursing homes and assisted living facilities in 22 states in the US demonstrated that SARS-CoV-2-positive participants are more likely to be positive for certain respiratory bacteria and viruses. This observation may help explain high hospitalization and mortality rates in older individuals.
Acknowledgements
This work was supported by Pathnostics.
Author Contributions
Concept and design: Baunoch, Wolfe
Acquisition, analysis, or interpretation of data: Lauterbach, Gnewuch
Drafting of the manuscript: Wang
Critical revision of the manuscript for important intellectual content: Wolfe, Baunoch, Luke, Huang, Zhao, Halverson, Cacdac
Statistical analysis: Zhao, Huang
Administrative, technical, or material support: Wang, Luke
Supervision: Luke
Conflict of Interest Disclosures
None reported.
References
- Coronavirus disease (COVID-19) Situation Report–108. Data as received by WHO from national authorities by 10:00 CEST, 7 May 2020. World Health Organization.
- https://www.cdc.gov/coronavirus/2019-ncov/cases-updates/cases-in-us.html
- https://data.cdc.gov/NCHS/Provisional-COVID-19-Death-Counts-by-Sex-Age-and-S/9bhg-hcku.
- Morens DM, Taubenberger JK, Fauci AS (2008) Predominant role of bacterial pneumonia as a cause of death in pandemic influenza: implications for pandemic influenza preparedness. J Infect Dis. 198(7): 962-970.
- MacIntyre CR, Chughtai AA, Barnes M, Ridda I, Seale H, et al. (2018) The role of pneumonia and secondary bacterial infection in fatal and serious outcomes of pandemic influenza a(H1N1) pdm09. BMC Infect Dis. 18(1): 637.
- https://www.cdc.gov/mmwr/preview/mmwrhtml/mm5838a4.htm.
- Rawson TM, Moore LSP, Zhu N, Ranganathan N, Skolimowska K, et al. (2020) Bacterial and fungal co-infection in individuals with coronavirus: A rapid review to support COVID-19 antimicrobial prescribing. Clin Infect Dis. pii: ciaa530.
- Wu CP, Adhi F, Highland K (2020) Recognition and management of respiratory coinfection and secondary bacterial pneumonia in patients with COVID-19. Cleve Clin J Med. pii: ccc015.
- Zhou F, Yu T, Du R, Fan G Liu Y, Liu Z, et al. (2020) Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet. 395(10229): 1054-1062.
- Chen N, Zhou M, Dong X, Qu J, Gong F, et al. (2020) Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. 395(10223): 507-513.
- Lin D, Liu L, Zhang M, Hu Y, Yang Q, et al. (2020) Co-infections of SARS-CoV-2 with multiple common respiratory pathogens in infected patients. Sci China Life Sci. 63(4): 606-609.
- Kim D, Quinn J, Pinsky B, Shah NH, Brown I (2020) Rates of Co-infection Between SARS-CoV-2 and Other Respiratory Pathogens. JAMA.
- https://www.cdc.gov/flu/symptoms/flustaph.htm
- Martin CM, Kunin CM, Gottlieb LS, Finland M (1959) Asian influenza A in Boston, 1957– 1958. II. Severe staphylococcal pneumonia complicating influenza. Arch Intern Med 103: 532-542.
- Hageman JC, Uyeki TM, Francis JS, Jernigan DB, Wheeler JG, et al. (2006) Severe community-acquired pneumonia due to Staphylococcus aureus, 2003-04 influenza season. Emerg Infect Dis. 12(6): 894-899.
- Oswald NC, Shooter RA, Curwen MP. Pneumonia complicating Asian influenza. Br Med J. 1958 Nov 29;2(5108):1305-11.
- Robertson L, Caley JP, Moore J (1958) Importance of Staphylococcus aureus in pneumonia in the 1957 epidemic of influenza A. Lancet. (7040): 233-236.
- Carbone I, Lazzarotto T, Ianni M, Porcellini E, Forti P, et al. (2014) Herpes virus in Alzheimer's disease: relation to progression of the disease. Neurobiol Aging. 35(1): 122-129.
- https://wwwnc.cdc.gov/eid/article/5/3/99-
- https://www.cdc.gov/epstein-barr/about-ebv.html.
- Jia HP, Look DC, Shi L, Hickey M, Pewe L, et al. (2005) ACE2 receptor expression and severe acute respiratory syndrome coronavirus infection depend on differentiation of human airway epithelia. J Virol. 79(23): 14614-14621.






























