Characterization of Gut Microbiota in Lean and Overweight Children with Type 1 Diabetes Compared to Matched Healthy Control Subjects
Shashikala Gowda1*, Mahmoud Ghannoum2,3 and Ryan Farrell1
1Department of Pediatric Endocrinology, Case Western Reserve University, USA
2Intergated Microbiome Core and Center for Medical Mycology, Case Western Reserve University, USA/i>
3University Hospitals Cleveland Medical Center, USA
Submission: January 31, 2022; Published: February 08, 2022
*Corresponding author: Shashikala Gowda, Department of Pediatric Endocrinology, Case Western Reserve University, UH Rainbow Babies & Children’s Hospital, USA
How to cite this article: Shashikala G, Mahmoud G, Ryan F. Characterization of Gut Microbiota in Lean and Overweight Children with Type 1 Diabetes Compared to Matched Healthy Control Subjects. Curre Res Diabetes & Obes J 2022; 15(3): 555912.DOI: 10.19080/CRDOJ.2022.15.555912
Abstract
Background: The incidence of both type 1 Diabetes (T1D) and overweight/obesity is on the rise. Previous studies have shown altered gut microbiota in children with T1D. Other studies have also shown gut microbial differences between obese and lean children without diabetes. The purpose of this study was to characterize the microbial composition of lean and overweight children with T1D at disease onset in comparison to matched healthy controls.
Methods: We enrolled 23 children aged 3-18 years with T1D at disease onset including 13 lean and 10 overweight children & compared them with age, gender and BMI matched controls. Serum concentration of cytokines and Lipopolysaccharide (LPS) level were measured in subjects with T1D. Stool swabs were collected from cases and controls and DNA extraction was performed using Qiagen DNA mini kit. Bacterial regions v3, v4 were amplified and sequencing was completed on Ion Torrent S5 sequencer.
Results: Four most abundant phyla in both TID and control groups were Firmicutes, Proteobacteria, Bacteroidetes and Actinobacteria. Compared with healthy controls, children with T1D had a significantly increased abundance of Facklamia (P=0.034) at the genus level & Veillonella Dispar (P=0.040) at the species level. At the genus level, significant increases in Klebsiella aerogenes (Enterobacter aerogenes P=0.028) were observed in lean T1D compared to their overweight counterparts. Lipopolysaccharide levels were increased in lean T1D (P=0.035). There was no statistically significant differences in measured cytokines between lean T1D and overweight T1D.
Conclusion: Our study showed alterations in the gut microbiome in subjects with T1D at disease onset compared to healthy controls. Additionally, lean T1D exhibited different microbiome patterns compared to overweight T1D.
Keywords: Type 1 diabetes mellitus; Gut microbiota; Overweight; Lean
Background and Significance
Recent studies suggest that the incidence of type 1 diabetes mellitus (T1D) is increasing. T1D is an autoimmune disease characterized by insulin deficiency, accounts for 5-10% of all types diabetes and the patients are usually younger and leaner [1]. According to the International Diabetes Federation, the annual incidence is increasing by 3% [2]. While a SEARCH study, unadjusted estimated incidence rates of T1D increased by 1.4% annually [3] (from 19.5 cases per 100,000 youths per year in 2002–2003 to 21.7 cases per 100,000 youths per year in 2011–2012).
Simultaneously, the prevalence of overweight and obesity has rapidly increased. Increasingly, children with T1D are overweight at the beginning of the disease, hindering the differential diagnosis between type 1 and type 2 diabetes [4] Wilkin et al. [5] in 2001 postulated the idea of the “accelerator hypothesis” where in, overweight may also partly explain the increasing incidence of T1D because of the interplay of beta-cell apoptosis and insulin resistance. Youth with T1D are predisposed to cardio metabolic complications by having diabetes, regardless of weight status.6 Little is known about the role of obesity in the initiation and pathogenesis of T1D. Emerging evidence suggests that excess weight in youth with T1D increases the risk of cardio metabolic complications [6,7]. Several case studies have reported that overweight and obese youth with T1D and insulin resistance may be predisposed to additional long-term cardiovascular complications thus categorizing them as high-risk group. In addition to cardiovascular risk, increased insulin resistance can aggravate short and long-term complications, thus influencing the natural history of T1D and its treatment [8].
Recent studies have found an association between gut microbiota and development of T1D. In-vivo murine studies have reported a close link between gut microbiome and T1D in bio breeding diabetic prone rats [9] and non-obese diabetic mice [10]. Furthermore, these models showed that the composition of gut microbiota modulates the function of both innate and adaptive immunity and demonstrated that intestinal bacteria are regulators of autoimmune diabetes [11]. Associations between the intestinal microbiota alterations and B-cell autoimmunity have been described in HLA-DR/DQ genetically predisposed humans with positive GAD/IA2 autoantibodies and with T1D diagnosis [12-14] Murrie et al. [14] has documented marked differences in the composition of gut microbiota and reported low Firmicutes to Bacteroidetes ratio (F/B) in children with T1D. Similarly, Giaongo et al. in 2011 found more diverse and stable gut microbiome in healthy controls when compared to children who developed autoimmunity and eventually T1D. The study also demonstrated that Firmicutes levels were decreased and the Bacteroidetes levels were increased as children who developed T1D [12].
Lipopolysaccharide (LPS) is a major component of the outer membranes of gram-negative bacterial species, known to play an important role in increasing the level of proinflammatory cytokines and impairing pancreatic beta-cell function, which may lead to diabetes. Therefore, LPS may act as a molecular link between gut microbiota, inflammation, and T1D [15]. Studies have shown obese children without T1D have elevated LPS levels and increased F/B ratio compared with lean children without T1D [16]. Researchers concluded that alteration in the gut microbiota is independently associated with obesity and T1D with evidence of increased systemic inflammation.
Clearly there is evidence of gut microbial differences that have been identified in children with T1D as well as children with obesity. Moreover, these gut microbe differences may have the capacity to influence disease processes and immune system responses. However, there are presently no published studies to date that have characterized the gut microbiota in children that are overweight with newly diagnosed T1D in comparison to the gut microbiome structure in newly diagnosed T1D in lean children. Identifying altered gut microbe composition in lean vs. overweight children with new onset T1D may play a role in disease progression, contribute to systemic inflammation, and potentially modulate risk for complication development over time.
Materials and Methods
Subjects enrolled into the study
In this cross-sectional case-control study, a total of 23 patients with T1D were enrolled from Rainbow Babies and Children’s Hospital between May 2019 and March 2020 under an IRB approved protocol (STUDY20190160). In addition, age, gender and BMI matched healthy controls were recruited from outpatient clinic. T1D cases were recruited at disease onset, when patients presented with clinical diagnosis of T1D with hyperglycemia, polyuria/polydipsia symptoms, with/without ketoacidosis, elevated hemoglobin A1c >6.5% and then confirmed the presence of at least one of the anti-islet autoantibodies including insulin (IAA), glutamic acid decarboxylase autoantibodies (GAD), tyrosine phosphatase autoantibodies (IA-2) and Zinc transporter 8 antibodies (ZnT8).
The parents/guardians of the participants were required to complete a questionnaire administered by the researcher in order to obtain following data: demographic information, mode of delivery, birth weight, breastfeeding, medication history, infections and antibiotic treatments prior to T1D diagnosis, diet and lifestyle habits. Informed written consent was obtained from all participants and their guardians. Inclusion criteria for recruitment included participants in the age group of 3-18 years, patients with clinical diagnosis of new onset T1D and controls without diabetes matched for age, gender and BMI. Participants were excluded if they were under 3 years and over 18 years of age, had prior use of probiotics, prebiotics or antibiotics in the 3 months prior to the study, had a history of international travel in the past month, or were pregnant. Controls with pre-existing autoimmune disease were also excluded. Patients with new onset T1D were recruited when they presented to the hospital for evaluation of their symptoms. Based on their BMI at presentation, they were categorized into lean if their BMI was <85th %ile and overweight groups if BMI >85th %ile. Recent anthropometric data from their primary care provider was obtained for patients with history of recent weight loss prior to presentation. Gender, age and BMI matched control subjects without Diabetes mellitus or any other autoimmune diseases ((12 lean and 10 overweight subjects) were recruited from outpatient endocrinology clinics at Rainbow Babies and Children’s Hospital, University Hospitals Cleveland Medical Center.
Laboratory measurements
Blood samples were collected from patients with T1D to measure serum concentration of cytokines, interleukin-1b (IL- 1b), IL-10, IL-6, IL-13, TNF- α and Lipopolysaccharide (LPS) level during their hospital stay. Samples were drawn by a research nurse and stored in at -80 C until ready for analysis. In patients with DKA, samples were obtained 24 hours after resolution of DKA. No blood samples were obtained from the controls.
Stool analysis
For patients with T1D, stool samples were collected during their hospital stay. For matched controls, stool sample from the participants were mailed to the research lab. Samples were processed in the Integrated Microbiome Core and microbiome composition was determined by NGS Ion Torrent technology. All data collected during the study was stored on secure UH server, a password-protected Excel document and paper records were stored in a locked cabinet at workplace which is only accessible by the PI, co-investigators, and approved study staff only.
Stool Analysis
DNA extraction
DNA was extracted using QIA amp Fast DNA Stool Mini kit (Qiagen GmpH, Hilden, Germany) according to manufacturer’s instructions. All swabs were transferred to tubes with 1 ml of InhibitEX lysis Buffer, incubated for 1 hour at 75°C and shaken using Fastprep 96 twice for 300 seconds each at a speed of 1800 RPM. Equal amounts of 100% ethanol and lysate were mixed in a collecting tube and passed through HiBind DNA Mini Columns (Omega Bio-tek, Georgia, USA) with the resulting DNA pellet eluted using 50 ul Molecular grade water.
PCR amplification
Amplifications of the 16S and 5.8S rRNA genes were performed using 16S-515 (5’-GGA CTA CCA GGG TAT CTA ATC CTG- 3’) and 16S -804 (5’-(TCC TAC GGG AGG CAG CAGT-3’) and ITS1 (5’-(TCC GTA GGT GAA CCT GCG G- 3’) and ITS4 (5’-TCC TCC GCT TAT TGA TAT GC- 3’) primers respectively. The PCR mixture was comprised of Q5 High-Fidelity Master Mix at a 1X concentration, along with a double volume of molecular grade water and 5μl 100mM of each primer. 100 ng of undiluted DNA was added to each 50 μl reaction. Thermo-cycling conditions consisted of an initial denaturation step (3 minutes at 98°C), followed by 30 cycles of denaturation (10 seconds at 98°C), annealing (10 seconds at 55°C for the 16S primers and 20 seconds at 58°C for the ITS primers), extension (10 seconds at 72°C) and a final extension step of 3 minutes 72oC. 10μl of each PCR product were separated using gel electrophoresis on 1.5% agarose gel (containing 7μg/ml ethidium bromide).
Library preparation
Equal volumes of bacterial 16S rRNA and fungal ITS amplicons were pooled, cleaned with AMPure XP beads (Beckman Coulter, CA, USA) to remove unused primers, then exposed to end repair enzyme for 20 minutes at room temperature. Subsequent to an additional AMPure clean up, ligation was performed at 25°C for 30 minutes using Ion Torrent P1 and a unique barcoded ‘A’ adaptor per pooled sample. After AMPure removal of residual adaptors, samples were concentrated to 1/4 volume for 1 hour using Labconco Vacuum under heat. All separate barcoded samples were then pooled in equal amounts (10μl) and size selected for the anticipated 16S and ITS range (300-800 bp) using Pippin Prep (Sage Bioscience). The library was amplified for seven cycles and quantitated on StepOne qPCR instrument ahead of proper dilution to 300 pM going into IonSphere templating reaction on the Ion Chef. Library sequencing was completed on Ion Torrent S5 sequencer (ThermoFisher Scientific) and barcode-sorted samples analyzed in our custom pipeline based on Greengenes V13_8 and Unite database V7.2 designed for the taxonomic classification of 16SrRNA and ITS sequences, respectively. Downstream data analysis was performed using Qiime software.
Statistical Analysis
The difference of continuous measurements in survey data between T1D patients and healthy controls, and the disparity of microbiome between T1D patients and healthy controls were examined using t-test. The comparison of continuous survey data between lean and overweight patients with new onset T1D, and comparison of microbiome profiles between lean and overweight patients with new onset T1D were also examined using t-test. The association of two categorical variables was examined using Chi-square test. All tests are two-sided and P-value ≤ 0.05 were considered statistically significant.
Results
Clinical characteristics of T1D and healthy controls
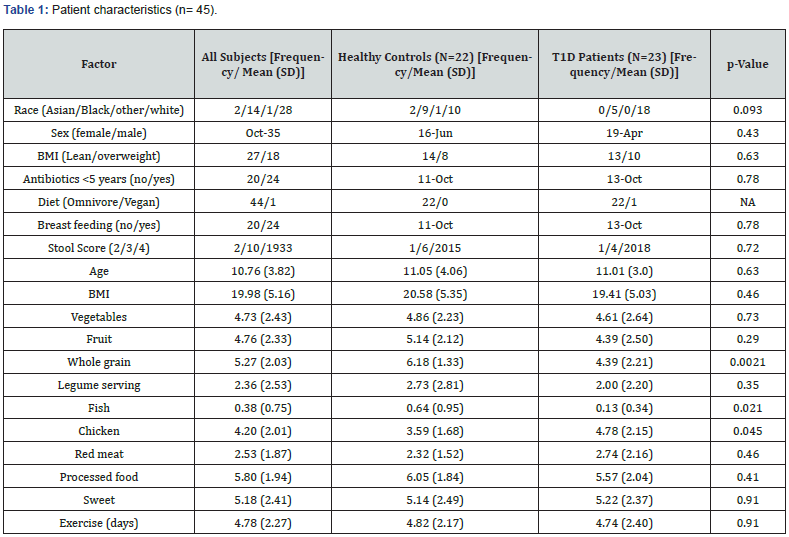
Twenty-three patients with new onset T1D and twenty-two age, gender and BMI matched controls completed the study. Participant characteristics are summarized in Table 1. The majority of the participants were male (78%) and white. Mean subject age at the time of diagnosis of T1D was 11.01 ± 3.0 years.
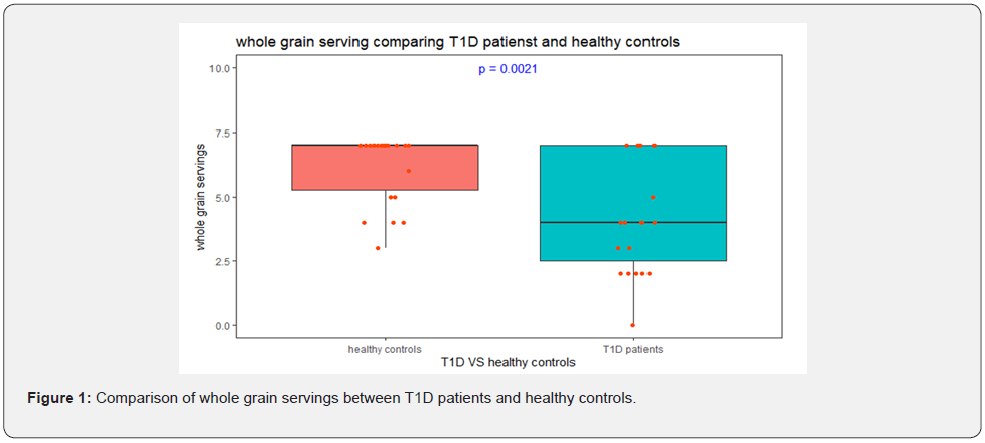
Of the 23 patients with new onset T1D, 43% were overweight and the remaining 57% were lean at diagnosis. Fifty six percent of them were breast fed and with similar percentage reported antibiotic use < 5 years of age. Only 1 participant out of 23 cases reported a vegan diet while the remaining 95% reported an omnivore diet. No statistically significant associations were noted between T1D and birth weight, breast feeding, early infection <5 years of age or antibiotic use. A significant association with consumption of whole grain was noted in control subjects when compared to T1D (P=0.002) (Figure 1) and also with consumption of fish in control subjects (P=0.021). No other significant differences were found between the two groups in the anthropometric variables.
Composition of gut microbiota and differences of phyla between T1D and healthy controls
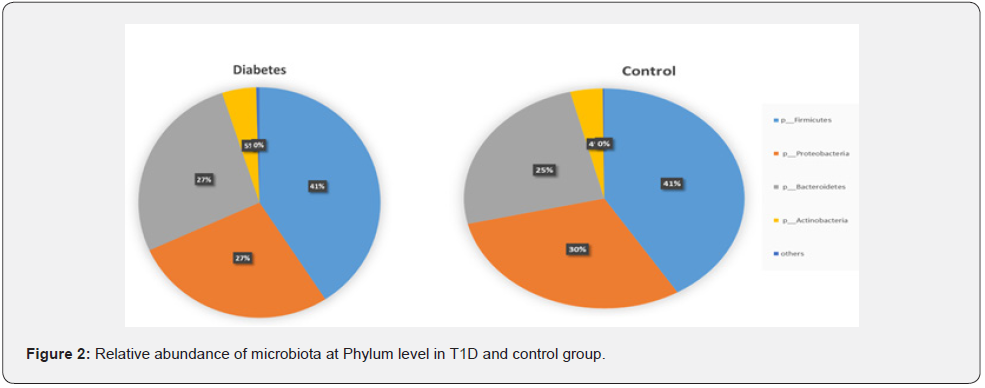
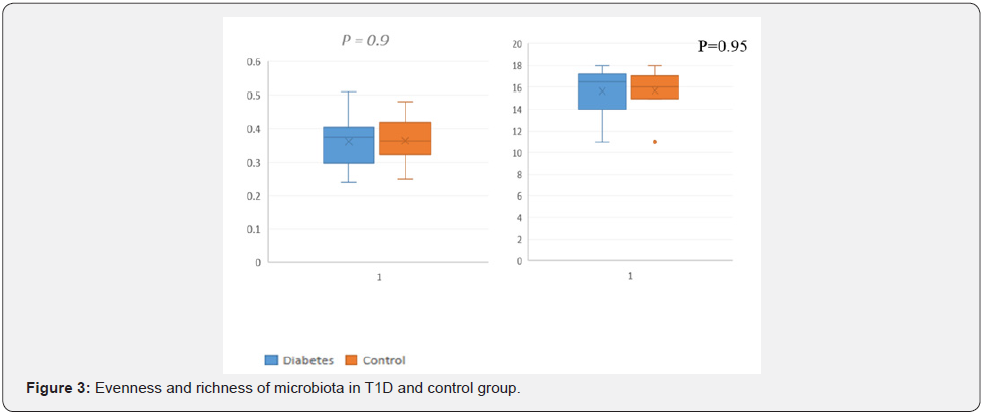
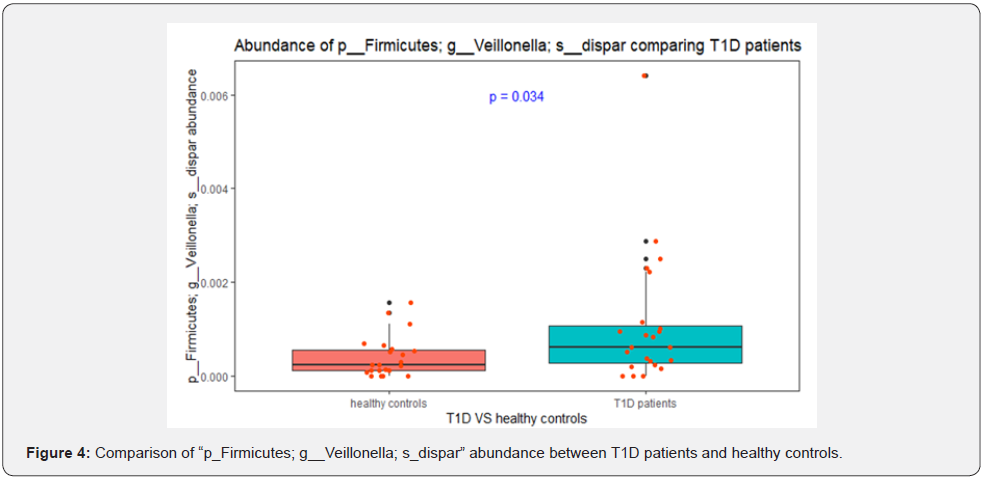
Our data showed that the four most abundant phyla in both T1D and control groups were Firmicutes, Bacteroidetes, Proteobacteria, and Actinobacteria (Figure 2). Dominant phyla were Bacteroidetes and Firmicutes, followed by Proteobacteria and Actinobacteria in T1D group with no differences between groups (P > 0.05), and the ratio of Firmicutes/ Bacteroidetes was not significantly different between the two groups (P=0.57). Results did not show any significant differences with regards to evenness, richness and alpha diversity (Figure 3). Compared with healthy controls, children with T1D were noted to have a significant increase in the abundance of Facklamia (P=0.034), a Gram-positive from the family Aeroccaceae, at the genus level and Veillonella Dispar (P=0.040), a Gram-negative anaerobic coccus, at the species level (Figure 4). Additionally, there was a significant increase in the pro-inflammatory bacteria Streptococcus in T1D individuals compared to controls (P=0.01).
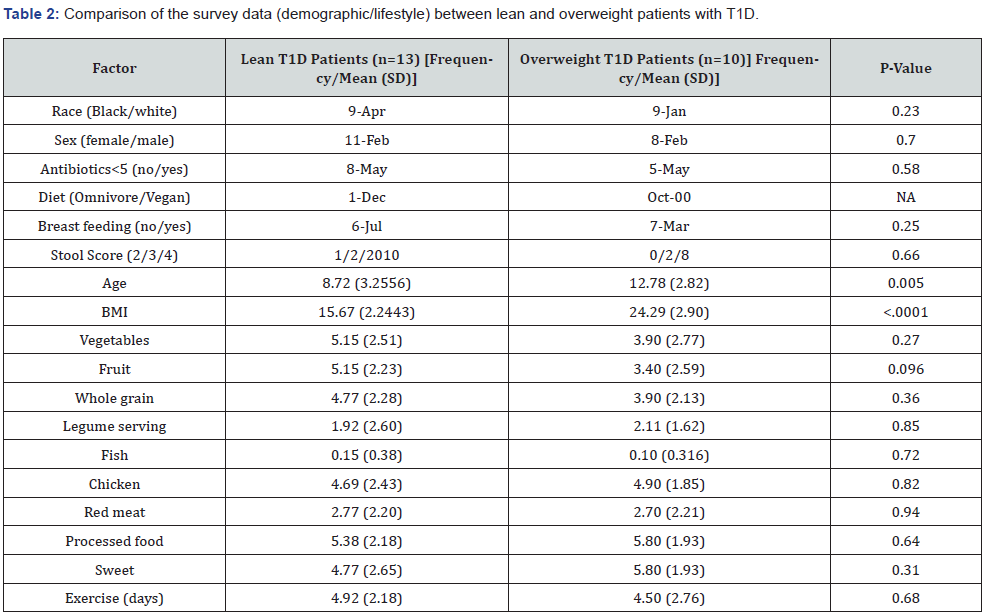
Comparison of clinical characteristics of lean with overweight T1D subjects
Overweight children with BMI>85th%ile diagnosed with T1D were older compared to lean subjects (BMI <85%ile) with T1D. The age at diagnosis of T1D in overweight group was 12.7 ± 2.8 years (P=0.005). No other significant differences were found between the two groups in the demographic, dietary and variables. In addition, the subjects of the two groups showed similar pattern in the breastfeeding time and early infections and antibiotics use < 5 years of age (Table 2).
Comparison of microbial difference between lean and overweight T1D subjects
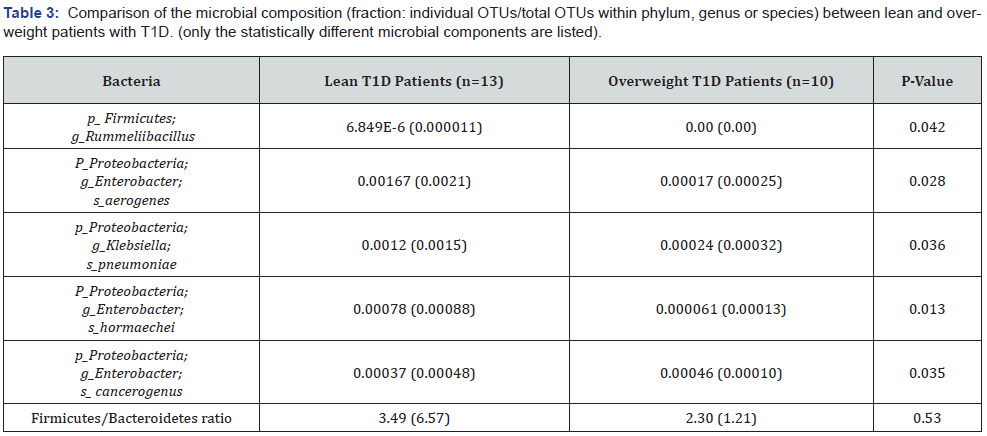
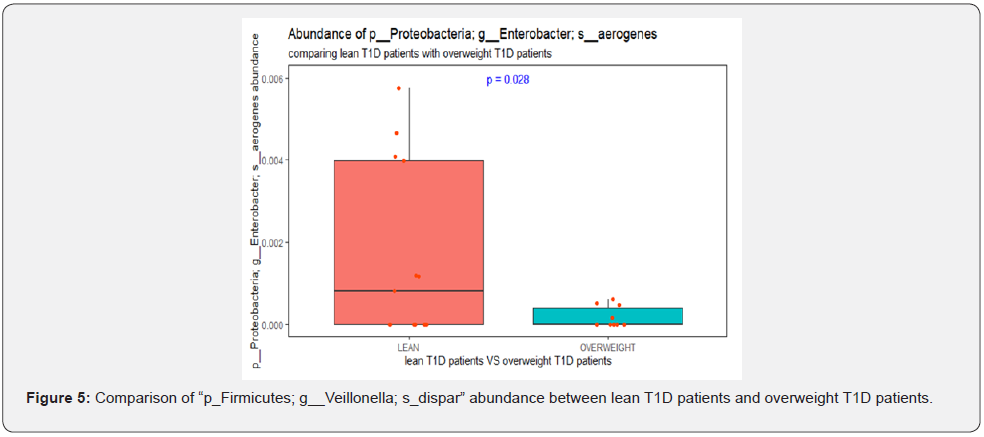
At the genus level, there was a significant increase in Klebsiella aerogenes (previously known as Enterobacter Aerogenes) (P=0.028) observed in lean T1D compared to their overweight counterparts (Figure 5). The Firmicutes/Bacteroidetes ratio was not different in overweight T1D compared to lean T1D (P=0.53) (Table 3).
Association between cytokines, LPS and microbiome
Lipopolysaccharide levels were increased in lean T1D when compared to overweight T1D (P=0.035) (Figure 6). Out of 13 lean patients 7 had DKA at presentation (54%) as opposed to 4 patients out of 10 overweight T1D.
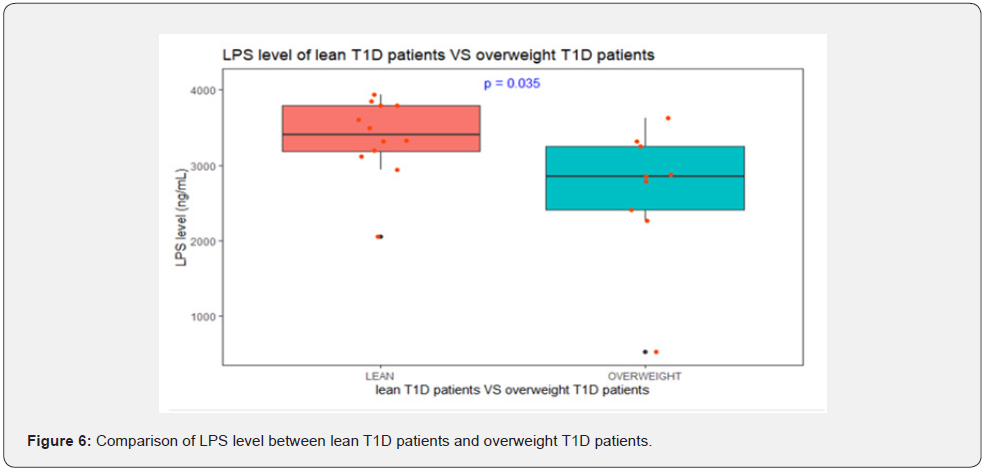
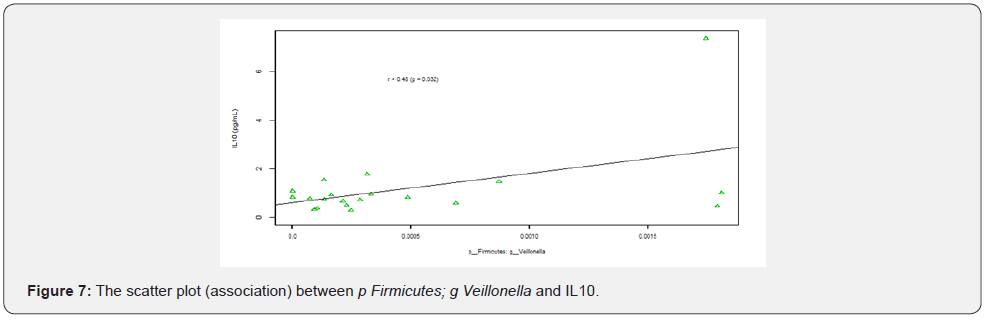
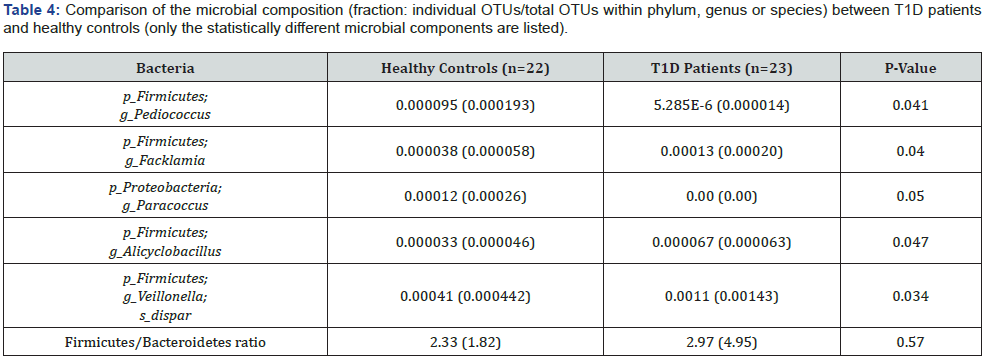
There were no statistically significant differences of other measured cytokines between lean T1D patients and overweight T1D patients. Among the microbial compositions that were significant between cases and healthy controls (Table 4) and between lean and overweight patients with T1D (Table 3), p_ Firmicutes; g_ Veillonella was the only bacteria that was associated with IL-10 (r = 0.48, P = 0.032, (Figure 7).
Discussion
Our study showed alterations in the gut microbiome in subjects with T1D at disease onset. The four most abundant phyla in both T1D and control subjects were Firmicutes, Proteobacteria, Bacteroidetes and Actinobacteria. There was no association with mode of delivery, breast feeding, birth weight or antibiotic use. There were statistically significant associations with consumption of whole grain and fish in control subjects. Microbial differences were noted amongst T1D patients –lean and overweight. Although not statistically significant, Firmicutes/ Bacteroidetes ratio was lower in overweight T1D compared to lean T1D. Lipopolysaccharide level was increased in lean T1D. Previous studies have suggested that perturbations to the normal composition of commensal communities in the gut may cause dysbiosis which in turn breaks the gut homeostasis and may contribute to the development of autoimmune diseases including T1D. Specific to T1D, the common gut microbiome shift associated with T1D development is the decreased microbial diversity compared to healthy controls. Our study showed that there were significant differences in the gut microbiota in subjects with T1D at disease onset when compared with healthy controls. We did not find any statistically significant decrease in the microbial diversity. Additionally, there was no association between T1D and breast feeding, early infections or antibiotic use < 5 years of age. Our study results also found a significant association between consumption of whole grain in healthy controls. A previous study in adults documented significant prebiotic effects, with increase in bacterial diversity, Actinobacteria, Bifidobacteria and Lactobacilli in people with increased consumption of whole grains [17]. Fermentation of dietary polysaccharides produces short chain fatty acids (e.g., acetate, butyrate, and propionate), the most important one being butyrate, which signals to host genes and affect lipid and glucose metabolism through the liver. The ability of short chain fatty acids (SCFAs) to affect insulin sensitivity and energy metabolism is now well documented. Production of SCFAs with signaling effects to stimulate secretion of GLP-1 and GLP-2 and PYY which increases gut mobility, satiety and insulin sensitivity. Microbes with prebiotic effects are butyrate producing bacteria where butyrate is the main energy source of the gut epithelium and also associated with increased insulin sensitivity and energy balance.
Our study population had most abundant phyla in both groups were Firmicutes, Bacteroidetes, Proteobacteria and Actinobacteria which is in agreement with many studies conducted in developed countries. From the phylum Firmicutes, at genus level Alicyclobacillus, Facklamia and V. dispar were significantly increased in T1D group compared to healthy controls. V. dispar is associated with production of SCFAs including acetate and propionate which are reported to play a role in gluconeogenesis. However, the Firmicutes/Bacteroidetes ratio was not significantly decreased in T1D group when compared to control subjects and this finding was not consistent with most studies in children with T1D where they had documented decreased Firmicutes/ Bacteroidetes ratio in T1D compared to healthy controls. Some studies showed a decrease in the F/B ratio in children with T1D when compared to healthy controls while others did not [18]. Many variables could have affected this ratio such as dietary differences, poor appetite, weight loss secondary to dehydration and the disease process just prior to initial presentation. In our study, participants were enrolled at disease onset and stool samples were obtained during their hospital stay, at least within 1 week of their diagnosis.
Amongst T1D patients, our data showed that overweight children diagnosed with T1D were older at presentation. BMI difference noted in these patients could have confounded the age of presentation in our study. Younger patients were leaner and older patients were overweight. There were significant microbial differences noted between lean and overweight T1D groups and our study is the first one to report this finding. When we compared the microbial composition at different taxonomy levels (phylum, genus or species) between lean and overweight patients with T1D, lean T1D group had increased abundance of Proteobacteria including Enterobater, Klebsiella Pnuemoniae which is a proinflammatory microbe, has LPS on the outer surface and can release inflammatory cascade in the host. Other microbes present in lean T1D including Enterobacter aerogenes, Enterobacter hormaechei and Enterobacter cancerogenes known for their proinflammatory properties and opportunistic infections.
Lipopolysaccharide level was increased in lean T1D when compared to overweight T1D. This was a paradoxical finding in our study since previous studies had reported the opposite in which LPS levels elevated in overweight individuals. Our study patients who presented in DKA at diagnosis of T1D, had their labs drawn for LPS and cytokines 24 hours post resolution of DKA. Dalton et al. reported in their study that LPS levels were elevated in patients with DKA even in the absence of infection however, levels down trended by 24 hours after resolution of DKA [19]. Our study has several strengths. This is the first study to characterize gut microbiota of overweight T1D in comparison to lean T1D at disease onset. These patients were clinically and immunologically confirmed T1D cases and enrolled at disease onset. T1D group was compared with age, gender and BMI matched healthy controls.
We also recognize several limitations. This was a small study. We had limited time frame which may have resulted in lack of varied demographics. Our study was a cross sectional study which cannot establish causation. These results are limited to assess association only. Larger prospective studies are needed to prove causation. Diet history of our study population was not collected using a validated questionnaire.
Conclusion
Our study showed alterations in the gut microbiome in subjects with T1D at disease onset, with significant increase in pro-inflammatory microorganisms. Lean T1D exhibited different microbiome patterns compared to overweight T1D. LPS level was increased in lean T1D and there was no significant difference of cytokines between lean T1D and overweight T1D subjects. Although we report microbial differences in T1D group from our cross-sectional study, we cannot differentiate if bacterial differences are potential causative of T1D or reflection of gut changes secondary to hyperglycemia.
Future Directions
Larger studies are needed to confirm association. We need longitudinal studies to assess microbiome/inflammatory markers at baseline associated with complication rates, cardiovascular disease, insulin dosing and A1c over time. Exploring these differences will advance our understanding of the potential significance of this knowledge gap about altered CV risk, altered insulin resistance and altered natural history of T1D which may point to unique therapeutic interventions in the future.
Acknowledgement
Authors are grateful to Dr. Sarah MacLeish MD and Dr. Ali Khalili MD for their encouragement. We would like to thank Iman Salem MD, Mauricio Retuerto, Shufen Cao PhD, and Pingfu Fu PhD who helped us with data analysis.
Funding Sources
This study was supported by the University Hospitals Cleveland Medical Center Fellowship Research Award Program and Friedman fellowship program.
References
- (2017) American Diabetes Association Classification and diagnosis of diabetes. Sec 2 In: Standards of Medical Care in Diabetes. Diabetes Care 40(Suppl 1): S11-S24.
- Whiting DR, Guariguata L, Weil C, Shaw J (2011) International Diabetes Federation. Diabetes Atlas. (5th), Brussels IDF 94(3): 311-321
- Elizabeth JMD, Dabelea D, Lawrence JM (2017) Incidence Trends of Type 1 and Type 2 Diabetes among Youths, 2002– N Engl J Med 376: 1419-1429.
- Libman IM, Pietropaolo M, Arslanian SA, LaPorte RE, Becker DJ (2003) Changing prevalence of overweight children and adolescents at onset of insulin-treated diabetes. Diabetes Care 26(10): 2871-2875.
- Wilkin TJ (2001) The accelerator hypothesis: Weight gain as the missing link between type I and type II diabetes. Diabetologia 44(7): 914-922.
- Krishnan S, Short KR (2009) Prevalence and significance of cardiometabolic risk factors in children with type 1 diabetes. Journal of the Cardiometabolic Syndrome 4(1): 50-56.
- Van Vliet M, Vander HJC, Diamant M, Rosenstiel IAV, Schindhelm RK, et al. (2010) Overweight is highly prevalent in children with type 1 diabetes and associates with cardiometabolic risk. J Pediatr 156(6): 923-929.
- Liu LL, Lawrence JM, Davis C, Liese AD, Pettitt DJ, et al. (2010) Prevalence of overweight and obesity in youth with diabetes in USA: The SEARCH for Diabetes in Youth study. Pediatric Diabetes 11: 4-11.
- Roesch LF, Lorca GL, Casella G, Giongo A, Naranjo A, et al. (2009) Culture independent identification of gut bacteria correlated with the onset of diabetes in a rat model. ISME J 3(5): 536-548.
- Kriegel MA, Sefik E, Hill JA, Wu HJ, Benoist C, et al. (2011) Naturally transmitted segmented filamentous bacteria segregate with diabetes protection in nonobese diabetic mice. Proc Natl Acad Sci USA 108(28): 11548-11553.
- Vaarala O (2013) Human intestinal microbiota and type 1 diabetes. Curr Diab Rep 13(5): 1-7.
- Giongo A, Gano KA, Crabb DB, Mukherjee N, Novelo LL, et al. (2011) Toward defining the autoimmune microbiome for type 1 diabetes. ISME J 5(1): 82-91.
- De Goffau MC, Luopajärvi K, Knip M, Ilonen J, Ruohtula T, et al. (2013) Fecal Microbiota Composition Differs Between Children with β-Cell Autoimmunity and Those Without. Diabetes 62(4): 1238-1244.
- Mora M, Leiva I, Zumaquero JMG, Tinahones JF, Cardona F, et al. (2013) Gut microbiota in children with type 1 diabetes differs from that in healthy children: a case-control study. BMC Medicine 11: 46.
- Pussinen W, Havulinna AS, Lehto M, Sundvall J, Salomaa V (2011) Endotoxemia is Associated with an Increased Risk of Incident Diabetes. Diabetes Care 34(2): 392-397.
- Bervoets L, Hoorenbeeck KV, Kortleven I, Noten CV, Hens N, et al. (2013) Differences in gut microbiota composition between obese and lean children: a cross-sectional study. Gut Pathogens 5(1): 10.
- Angie J, Katie A (2019) The Effects of Intact Cereal Grain Fibers, Including Wheat Bran on the Gut Microbiota Composition of Healthy Adults: A Systematic Review. Front Nutr 6: 33.
- He Z, Lin S, Siwen Z, Xue Z, Xiaokun G, et al. (2020) Evaluating the Causal Role of Gut Microbiota in Type 1 Diabetes and Its Possible Pathogenic Mechanisms. Frontiers in Endocrinology 11: 125.
- Rory D, Hoffman WH, Passmore GG, Martin SLA (2003) Plasma C - reactive protein Levels in Severe Diabetic Ketoacidosis. Ann Clin Lab Sci Fall 33(4): 435-442.






























