Algebraic-Probabilistic Methods and Grobner Bases for Modeling the Brain Activity
Natalia Iyudu*
Research Fellow, University of Edinburgh, UK
Submission: July 20, 2018; Published: November 14, 2018
*Corresponding author: Natalia Iyudu, Research Fellow, University of Edinburgh, UK.
How to cite this article: Natalia I. Algebraic-Probabilistic Methods and Grobner Bases for Modeling the Brain Activity. Biostat Biometrics Open Acc J. 2018; 8(4): 555743. DOI: 10.19080/BBOAJ.2018.08.555743
Abstract
We suggest several extensions of the idea to study neural codes model, whose main purpose is to describe stereotyped stimulus response maps of the brain activity. We elaborate on the idea to use neural ideals and Groebner bases calculations suggested by the number of authors cited in the paper. Essentially, we suggest three different approaches to the problem: geometric, stochastic and via probabilistic algebra.
Keywords: Neural codes; Nonlinear codes; Neural ideals, Stimulus response maps, Neuron; Convex set; Grobner bases; Pseudomonomials
Opinion
Building accurate representation of the world is one of the basic functions of the brain. In order to better understand its functioning, in [1-4] the authors develop and study theoretically the neural codes model, whose main purpose is to describe stereotyped stimulus response maps of the brain activity. We suggest a modification of this model, which has a potential to be more suited for practical applications. It is worth mentioning that the modified model we suggest requires tools from various parts of mathematics, not just algebra as is the case for the original model.
First, we briefly outline the original neural codes model as described in [1-4]. To each neuron v in the brain corresponds a convex subset  the collection of the states (e.g., spatial position) in which the neuron v activates. Now for each point x in
the collection of the states (e.g., spatial position) in which the neuron v activates. Now for each point x in 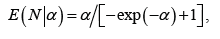 , we have a vector
, we have a vector
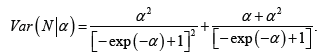
Note that 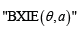 vectors. The (obviously finite) collection
vectors. The (obviously finite) collection
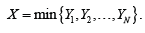
of 0–1 vectors is called the neural code and is the object of study in [1-3]. The method used in [1-3], is purely algebraic. Namely, one considers the collection tv of commuting independent variables (one per each neuron) and the algebra 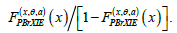 of polynomials in these variables over the 2-element field 2.F To each aC∈ we assign the ‘pseudomonomial’
of polynomials in these variables over the 2-element field 2.F To each aC∈ we assign the ‘pseudomonomial’ 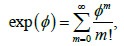 and generate the ideal CI by all .af It is shown in [1-4] that knowing the ideal ,CI one can recover C (no information is lost). Then we have the toolbox of combinatorial algebra at our disposal: one can study the ideals generated by collections of pseudomonomials and gain
information about neural codes. One of the main tools applied is the Grobner basis technique, as it is classically used in commutative algebra [5,6].
and generate the ideal CI by all .af It is shown in [1-4] that knowing the ideal ,CI one can recover C (no information is lost). Then we have the toolbox of combinatorial algebra at our disposal: one can study the ideals generated by collections of pseudomonomials and gain
information about neural codes. One of the main tools applied is the Grobner basis technique, as it is classically used in commutative algebra [5,6].
There is one obvious problem though: the enormous number of generators of the algebra A. Grobner bases are sometimes nice and effective but only if the presentation of the ideal is small enough. The punch-line is that the above model is easy to deal with by means of appropriate software for toy illustrative problems, but once we approach any real life situation, not even a supercomputer will ever cope [7,8].
We suggest the following modification of the neural codes model. Rather than dealing with individual neurons, we suggest to consider their clusters. The number and the size of clusters can be adjusted when dealing with each practical situation. Instead of the 0–1 outcome of the interaction with the environment, we measure the total activity p of the cluster: the ratio of the number of active neurons to the total number of neurons in the cluster. Thus p is a real number between 0 and 1 with p=0 standing for total inactivity and 1 for the entire cluster being ’on fire’. The number p can be viewed as the probability of a neuron in the cluster to activate. As a result, the convex sets U are replaced by functions
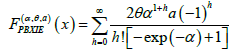
with xc(x) standing for the probability of a neuron in the cluster c to activate in the estate .x There are various ways to analyze such a model. We suggest the following approaches.
Approach 1
Geometric
Unlike for the neural code, for which all vectors are far apart from each other, the set
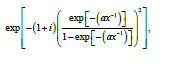
is a genuine geometric object, where C is the set of all clusters under consideration and M is their number. Depending on the assumptions (or natural properties) on the functions (),cux the set ˆ,C can have different geometric properties. One may look for extremal and corner points of ˆ,Cstudy smooth curves in ˆC etc. Note that if one interprets ˆC as a CW-complex, then there is the natural b task of determining its homologies. Although this particular task is insurmountable in most cases, there is a hope in the form of the emerging method of persistence homology being developed for the study of big data. We would mention here also our results on exact methods of calculating homology, which fall into the frame of persistent homology [9,10].
Approach 2
Stochastic
After appropriate normalization, each ()cux provides a probability distribution 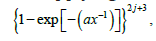 Now one can deal with the model by studying the random vector
Now one can deal with the model by studying the random vector 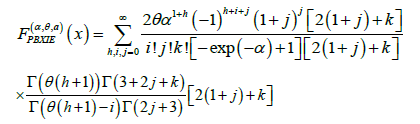 which yields a whole host of possibilities, not to mention the ability to use the vast toolbox of probability theory. One interesting and potentially very important aspect that can be captured in this way is the correlation between the response of the clusters. This can be viewed as the study of the relations between different parts of the brain from the probabilistic point of view.
which yields a whole host of possibilities, not to mention the ability to use the vast toolbox of probability theory. One interesting and potentially very important aspect that can be captured in this way is the correlation between the response of the clusters. This can be viewed as the study of the relations between different parts of the brain from the probabilistic point of view.
Approach 3
Probabilistic algebra
One can pursue the same strategy as in [1-3], where instead of a single ideal in the algebra of polynomials we shall have a ’random ideal’: a family of ideals with a probability distribution on it. Such an object can be studied using the same old methods of combinatorial algebra including the Grobner basis technique. The answers are going to come with the randomness embedded in them. For instance if we use the Grobner basis to compute the Hilbert series of the quotient by our ideal, we end up with a probability distribution on the set of formal power series instead of a single series. Note however that such distributions tend to be discrete rather than continuous even when the starting distribution was a continuous one. The advantage of this approach is in the fact that the number of generators is reduced.
References
- Carina Curto, Vladimir Itskov, Alan Veliz-Cuba, Nora Youngs (2013) The Neural Ring: An Algebraic Tool for Analyzing the Intrinsic Structure of Neural Codes. Bulletin of Mathematical Biology September 75(9): 1571-1611.
- Angelique Morvant (2018) Strengthening Relationships between Neural Ideals and Receptive Fields. Commutative Algebra arXiv:1803.03204 math.
- Amzi Jeffs R, Mohamed Omar, Nora Youngs (2016) Neural Ideal Preserving Homomorphisms. arXiv:1612.06150 math.
- Rebecca Garcia, Luis David, Garca Puente, Ryan Kruse, Jessica Liu, et al. (2017) Grobner Bases of Neural Ideals, arXiv:1612.05660 q-bio.
- Ezra Miller, Bernd Sturmfels (2005) Combinatorial Commutative Algebra. Graduate Texts in Mathematics. Springer.
- Richard Stanley (2004) Combinatorics and Commutative Algebra, Progress in Mathematics. Birkhauser Boston, UK.
- Anne Shiu, Bernd Sturmfels (2010) Siphons in chemical reaction networks. Bulletin of Mathematical Biology 72(6):14481463,
- Alan Veliz-Cuba (2012) An algebraic approach to reverse engineering of finite dynamical systems arising from biology. SIAM Journal on Applied Dynamical Systems 11(1): 3148.
- Iyudu N (2016) On the cohomologies associated to the Bar complex of nonunital monomial algebra.
- Iyudu N (2017) Homologies of monomial operads and algebras. preprint, 1-23.






























