Approximate Bayesian Computation for Biological Science
Ritabrata Dutta1* and Antonietta Mira1,2
1Institute of Computational Science, Universita della Svizzera italiana, Switzerland
2Department of Science and High Technology, Universita degli Studi degli Insubria, Italy
Submission: January 30, 2018; Published: July 09, 2018
*Corresponding author: Ritabrata Dutta, Institute of Computational Science, Universita della Svizzera italiana, Switzerland; Email: duttar@usi.ch
How to cite this article: Ritabrata D, Antonietta M. Approximate Bayesian Computation for Biological Science. Biostat Biometrics Open Acc J. 2018; 7(3): 555715. DOI: 10.19080/BBOAJ.2018.07.555715
Abstract
Approximate Bayesian computation (ABC) provides us a rigorous tool to perform parameter inference for models without an easily accessible likelihood function. Here we give a short introduction to ABC, focusing on applications in biological science. Furthermore, we introduce users to a Python suite implementing ABC algorithms, with optimal use of high performance computing facilities.
Keywords: Approximate Bayesian Computation; Biological science; ABCpy; Datasets; Markov chain Monte Carlo methods; Maximum likelihood; Bayesian methodology; Genomes; Statistical science; Parameters; Fundamental rejection; Fixed parameter; Minimize; Computationally expensive; Algorithms; Applications to biology; Network; Epidemics; Random forest; Posterior distribution; Uncertainty quantification
Introduction
DWith the recent innovations in biological science, we are increasingly facing large datasets of varied type and more realistic but complex models of natural phenomenon. This trend has led to a scenario where we do not easily have a likelihood function which is available in closed form and thus easy to evaluate at any given point (as required by most Monte Carlo and Markov chain Monte Carlo methods). Thus, traditional likelihood-based inference, as maximum likelihood or Bayesian methodology, is not possible. Still, if from the complex model, given values of the parameters that index it, we can forward simulate pseudo-dataset, a new methodology becomes available, namely Approximate Bayesian Computation (ABC). Models that have this possibility of forward simulating are known as simulator-based models and are becoming more and more popular in diverse fields of science [1-3], just restricting to the biological domain we can find many examples: evolution of genomes Marttinen et al. [4], numerical model of platelet deposition [5], demographic spread of a species among many [6]. Research in statistical science in the last decade or so, has illustrated how ABC can be a tool to infer and calibrate the parameters of these models.
The fundamental rejection ABC sampling scheme iterates between three step: First a pseudo-dataset, xsim is ,simulated from the simulator-based model M(φ) for a fixed parameter value of .φThen we compute a measure of the closeness between xsim and x0, the observed dataset, using a pre-defined discrepancy measure d(xsim,x0) Finally, based on this discrepancy measure, ABC accepts the parameter value φ when d(xsim,x0) is less than a pre-specified threshold value .∈
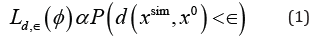
and, as a consequence, the accepted parameters follow the posterior distribution of φconditional on d(xsim,x0)<∈
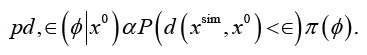
For a better approximation of the likelihood function, computationally efficient sequential ABC algorithms Marin et al. [7] Lenormand et al. [8], Albert et al. [9] decrease the value of the threshold ∈ adaptively while exploring the parameter space. The crucial aspect for a good ABC approximation to the likelihood function is the choice of the summary statistics, as we define the discrepancy measure between xsim and x0 through a distance between the extracted summary statistics from xsim and x0 Knowledge domain driven summary statistics are normally chosen keeping in mind that we want to minimize the loss of information on φ contained in the data through the choice of summary statistics. But one can also rely on automatic summary selection for ABC, thus removing a subjective component in this choice, as described in Fearnhead & Prangle [10], Pudlo et al. [11], Jiang et al. [12] and Gutmann et al. [13]. ABC provides a tool for statistical inference for simulator-based models, still, the necessity to simulate lots of pseudo-data, makes the algorithm extremely computationally expensive when data-simulation itself is costly. Further, the varied types of data sets available in different domain specific problems have hindered the applicability ABC algorithms to many applied science domains. Recently, Dutta et al. [14,15], have developed a High Performance Computing framework to efficiently parallelize different ABC algorithms which we believe will be extremely beneficial for inferential problems across different scientific domains. To highlight the versatility of ABC and ABCpy in diverse applied problems, we point the interested reader to two recent research paper of ours with applications to biology:
a. Estimation of parameters of a numerical platelets deposition model, where each forward simulation takes 10 minutes [16] and
b. Estimation of parameters of spreading processes on a network (e.g., epidemics on a contact network, but also fake news on a social network where the datasets are series of networks) [17].
Conclusion
We would like to stress here the fact that ABC inference scheme provides not only a point estimate of the parameters of interest but also their entire (approximated) posterior distribution thus allowing for uncertainty quantification: the higher the variability of the posterior distribution the higher the uncertainty inherent in the inferential scheme. Via the ABC approximated posterior one can then construct credible intervals and perform hypothesis testing. Furthermore, ABC allows to compare possible alternative models by simply adding, to the three steps ABC scheme illustrated above, an additional initial layer where first a model index is sampled from the model prior distribution and then, once a model has been selected a regular ABC scheme within that model is performed. For details on ABC model selection via random forest approach [11].
References
- Martinez EA, Muschik CA, Schindler P, Nigg D, Erhard A, et al. (2016) Real-time dynamics of lattice gauge theories with a few-qubit quantum computer. Nature 534(7608): 516-519.
- Turchin P, Currie TE, Turner EA, Gavrilets S (2013) War, space, and the evolution of old world complex societies. Proc Natl Acad Sci U S A 110(41): 16384-16389.
- Joop S, Robert AC, Richard G, Michelle F, Matthieu S, et al. (2015) The EAGLE project: simulating the evolution and assembly of galaxies and their environments. Monthly Notices of the Royal Astronomical Society 446(1): 521-554.
- Marttinen P, Croucher NJ, Gutmann MU, Corander J, Hanage WP (2015) Recombination produces coherent bacterial species clusters in both core and accessory genomes. Microb Genom 1(5): e000038.
- Chopard B, de Sousa DR, Lätt J, Mountrakis L, Dubois F, et al. A physical description of the adhesion and aggregation of platelets. R Soc Open Sci 4(4): 170219.
- Excoffier L, Dupanloup I, Huerta-Sánchez E, Sousa VC, Foll M (2013) Robust demographic inference from genomic and snp data. PLoS genetics 9(10): e1003905.
- Jean M, Pierre P, Christian P, Robin J (2012) Approximate Bayesian computational methods. Statistics and Computing 22(6):1167-1180.
- Maxime L, Franck J, Guillaume D (2013) Adaptive approximate Bayesian computation for complex models. Computational Statistics 28(6): 2777-2796.
- Carlo Albert, Hans RK, Andreas S (2015) A simulated annealing approach to approximate Bayesian computations. Statistics and Computing 25: 1217-1232.
- Paul F, Dennis P (2012) Constructing summary statistics for approximate Bayesian computation: semi-automatic approximate Bayesian computation. Journal of the Royal Statistical Society: Series B (Statistical Methodology) 74(3): 419-474.
- Pierre P, Jean M, Arnaud E, Jean Ma, Mathieu G, et al. (2015) Reliable ABC model choice via random forests. Bioinformatics 32(6): 859-866.
- Bai J, Tung W, Charles Z, Wing H (2015) Learning summary statistic for approximate Bayesian computation via deep neural network. arXiv preprint arXiv:1510.02175.
- Michael UG, Ritabrata D, Samuel K, Jukka C (2017) Likelihood-free inference via classification. Statistics and Computing 1: 1-15.
- Dutta R, Schoengens M, Onnela JP, Antonietta M (2017) ABCpy: A user-friendly, extensible, and parallel library for approximate Bayesian computation. In Proceedings of the Platform for Advanced Scientific Computing Conference. ACM.
- Ritabrata D, Marcel S, Avinash U, Jukka P, Antonietta M (2017) Abcpy: A high-performance computing perspective to approximate Bayesian computation. arXiv preprint arXiv:1711.04694
- Ritabrata D, Bastien C, Jonas L, Frank D, Karim Z, et al. (2017) Parameter estimation of platelets deposition: Approximate bayesian computation with high performance computing. arXiv preprint arXiv:1710.01054.
- Ritabrata D, Antonietta M, Jukka O (2017) Bayesian inference of spreading processes on network. arXiv preprint arXiv:1709.08862.






























