Genotypic Distribution of Hcv in Northeren India
Parveen Malhotra*, Vani Malhotra, Paramjeet Singh Gill, Isha Pahuja, Ajay Chugh and Sudhanshu
Department of Medical Gastroenterology, Gynecology & Obstetrics and Microbiology, India
Submission:December 27, 2018;Published: February 04 , 2019
*Corresponding author: Parveen Malhotra, Department of Medical Gastroenterology, Gynecology & Obstetrics and Microbiology, 128/19, Civil Hospital Road, Rohtak, Haryana, India
How to cite this article: Parveen M, Vani M, Paramjeet S G, Isha P, Ajay C et, al. Genotypic Distribution of Hcv in Northeren India. Adv Res Gastroentero Hepatol. 2019; 12(2): 555833. DOI: 10.19080/ARGH.2019.12.555833.
Abstract
Introduction: Viral hepatitis is recognized as a public health problem in India and globally. Various etiological agents (Hepatitis A, B, C, D, E and G virus) have been implicated that can lead to acute, chronic or acute on chronic infection. While Hepatitis A and E are often the cause of sporadic outbreaks of hepatitis, Hepatitis B and C can either clear spontaneously or can lead to chronic infection and thereafter sequel like cirrhosis and hepatocellular carcinoma (HCC).
Aims & objectives: To determine the genotyping of HCV infections among chronic hepatitis patients attending Department of Gastroenterology, PGIMS Rohtak.
Summary and conclusions: Our study confirms the finding of previous studies that in India Genotype 3 is most common genotype which is difficult to treat Genotype with oral antiviral therapy. Hence as per World Health Organization target of elimination of Hepatitis C till 2030 requires continuous, persistent and determined efforts at level of Center, State and society level.
Keywords: Chronic hepatitis C; Genotypes; HCV; RNA; Quantitative; Viral hepatitis; Hepatocellular carcinoma; Gastroenterology
Abbrevations: NPIs: Nucleoside Polymerase Inhibitors; SVR: Sustained Virologic Response Rate; PCR: Polymerase Chain Reaction; HCV: Hepatitis C Virus; HCC: Hepatocellular Carcinoma
Introduction
Viral hepatitis is recognized as a public health problem in India and globally. Various etiological agents (Hepatitis A, B, C, D, E and G virus) have been implicated that can lead to acute, chronic or acute on chronic infection. While Hepatitis A and E are often the cause of sporadic outbreaks of hepatitis, Hepatitis B and C can either clear spontaneously or can lead to chronic infection and thereafter sequel like cirrhosis and hepatocellular carcinoma (HCC) [1].
Hepatitis C Virus (HCV) is one of the major causes of death and morbidity and recent estimates showed Seroprevalence of 2.8%, corresponding to greater than 185 million infections worldwide [2]. In India there are about 6-12 million people with HCV infection with a mortality of 37,000 deaths in the year 2015 [1].
HCV is a single-stranded, positive sense RNA virus belonging to family Flaviviridae and genus Hepacivirus. The genome is approximately 10 kb in length, encoding a poly-protein product consisting of structural (capsid and envelope) and non-structural viral proteins. The HCV genome consists of seven functional regions- the core, the envelope, including the E1 and E2 regions, and the nonstructural region, including NS2, NS3, NS4, and NS5. Comparisons of sub-genomic regions, such as E1, NS4 or NS5, have allowed variants to be classified into major genotypes with many subtypes.
The term genotype refers to different genetic variations or strains of HCV. The variance in genetic differences is approximately one third between the different genotypes. There are six distinct major genotypes (genotype 1 to 6) and a minor genotype 7 and more than 50 subtypes within the genotypes of HCV have been identified. Within each genotype are further divisions called subtypes (for example 1a and 1b) and intra-genotypic variations are referred to as quasi-species and differ in sequence homology by only a few percent. As the HCV virus replicates rapidly, it constantly changes and mutates thus making it difficult to develop vaccine against HCV [3].
Genotyping is useful for investigating outbreaks and for understanding the epidemiology and virological features of this virus. Accurate classification of genotypes and subtypes of HCV is important for correct stratification of groups and accurate analysis of data related to efficacy and resistance of new HCV drugs. It is also essential for the implementation of therapeutic procedures, the production of effective vaccines, and the improvement of diagnostic tests [3]. A recent study shows that HCV genotypes 1 and 3 are the two most prevalent genotypes in India and globally, and account for about 76.3% cases of HCV in the world [4].
HCV infection is usually acquired through infected syringes and needles, and transfusion of infected blood. Sexual transmission of HCV occurs infrequently in heterosexual couples. It is reported to be more common in HIV-positive persons, particularly in high risk group of male having sex with male (MSM). The risk of transmission of HCV from a mother to her child occurs in 4–8% of births to women with HCV infection, and in 10.8–25% of births to women with HIV and HCV co-infection. HCV causes both acute and chronic hepatitis. Acute hepatitis is often clinically mild and marked by fluctuating elevations of serum aminotransferase levels; >50% likelihood of chronicity, leading to cirrhosis in >20% [5].
Chronic infection with HCV is usually clinically silent and is only very rarely associated with life-threatening disease. Spontaneous clearance of acute HCV infection occurs within six months of infection in 15–45% of infected individuals in the absence of treatment. Almost all the remaining 60–80% of persons will harbor HCV for the rest of their lives (if not treated) and are considered to have chronic HCV infection. Left untreated, chronic HCV infection can cause liver cirrhosis, liver failure and HCC. Of those with chronic HCV infection, the risk of cirrhosis of the liver is 15–30% within 20 years. The risk of HCC in persons with cirrhosis is approximately 2–4% per year [6].
Currently highly effective DAAs have however changed the HCV treatment paradigm, leading to hope of elimination of this infection as a public health threat by the year 2030. WHO introduced global targets for the care and management of HCV including “a 90% reduction in new cases of chronic hepatitis C, a 65% reduction in hepatitis C deaths, and treatment of 80% of eligible people with chronic hepatitis C infection [7]. DAAs are the recommended first line treatment in India.
The combination of the DAAs and the duration of treatment will depend on presence or absence of cirrhosis and on the genotype of the virus and the duration of treatment will depend on the several situations such as, cirrhosis versus non-cirrhosis, presence of decompensation (ascites, variceal bleeding, hepatic encephalopathy) or infections. The four classes of DAAs are (a) protease inhibitors targeting NS3/5 protein, (b) nucleoside polymerase inhibitors (NPIs) targeting NS5B proteins, (c) nonnucleoside polymerase inhibitors (NNPIs) targeting NS5B protein, and (d) NS5A inhibitors. But treatment with combination therapy such as pegylated interferon in combination with ribavirin has been shown to offer better treatment responses than monotherapy. Thus, the ultimate goal of HCV treatment is to achieve significant sustained virologic response rate (SVR) by interfering with HCV replication [5].
Review of Literature
In 2016, Petruzziello A et al. [2] conducted a systematic study to quantify global HCV epidemiology, using the best available published data between 2000 and 2015 from 138 countries. Total global HCV prevalence was estimated to be 2.5% (177.5 million of HCV infected adults), ranging from 2.9% in Africa and 1.3% in Americas. HCV genotype 1 was the most prevalent worldwide genotype (49.1%), followed by genotype 3 (17.9%), 4 (16.8%) and 2 (11.0%). Genotypes 5 and 6 were observed to be responsible for the remaining < 5% of the study population.
In a study conducted by Samir RS the prevalence of HCV genotypes and subtypes in different geographic regions of India was assessed and the frequency and variation in patient genetic profiles among 500 patients enrolled at 19 centers, which were categorized into four geographic regions (North, South, East, and West) was determined. They observed that genotype 3 was the most common genotype overall (54%) followed by genotype 1 (24%) and genotype 4 (6%) [4].
An observational study was conducted by B. Debjyoti among chronic hepatitis patients visiting Calcutta National Medical College, Kolkata, India. Of the 350 chronic liver disease patients, 110 were positive for anti-HCV antibodies. Further, these patients were tested for the presence of HCV RNA and 66% patients were found to be HCV RNA positive. All HCV RNA positive samples were then subjected to genotype determination using RFLP and type specific PCR followed by direct sequencing. Genotype 3 was found to be most common genotype positive in 48 (73%) patients, of these 44 patients (67%) showed infection with subtype 3a while 4 patients (6%) had subtype 3b. Genotype 1a was seen in 18 (27%) patients [8].
A study was conducted in a North Indian hospital by Chakravarti A, et al. [9] to observe any emerging trend of HCV in high risk groups, transmission patterns, genotypic distribution of the virus in this geographical region and its correlation with viral load. Fifty-four anti-HCV positive patients with 31 positives for HCV RNA were included in the study. HCV genotyping was carried out by restriction fragment length polymorphism (RFLP) followed by direct sequencing of the core region. In 45/54 (83.3%) anti-HCV positive patients, iatrogenic procedures were found to be responsible for transmission, with blood transfusion alone responsible in 36/54 (67%) patients. Genotype 3 was found to be the commonest genotype present in all risk groups followed by type 1 and 2. Subtype 3b (35.5%) was found more prevalent than subtype 3a. A higher frequency of subtype 1b (19.4%) was also seen. Genotype 1 was associated with a significantly higher viral load as compared to genotypes 3 and 2.
A government initiated cross-sectional study was conducted by Verma R, et al. [10] in the year 2012 to determine the incidence of HCV infection in the Ratia block of the Fatheabad district, Haryana, India. This study was carried out by house-tohouse visits over 2 weeks (13 February to 25 February 2012). After obtaining the written consent and collecting patients’ clinical details and risk factors, blood sample was collected from suspected cases of hepatitis. From a total of 7533 samples, anti-HCV Abs were found to be positive in 25.3 % (1912/7533) cases by using ELISA test. Maximum number of HCV cases, 486 (29.8 %), were found in the age group 31–40 years and only 0.8 % of cases were found in the 0–10-year age group. A total of 1202/1630 (74.0 %) were infected with HCV in the age group 20–60 years. This was the first and the largest government-aided study to be conducted in Haryana state of India, with important observations that there were some highly endemic areas in Haryana for HCV infection with a very high prevalence. Following this survey, the Haryana government launched the State-wide program of free treatment and testing for HCV in the year 2014 and is being successfully running in the state.
Keeping the observations of the study conducted by Verma R et al. [10] the present study has been planned to assess the prevalence of HCV infection with respect to the genotyping and its correlation with clinical spectrum in chronically infected hepatitis patients in Northern India.
Aim & Objective
To determine the genotyping of HCV infections among chronic hepatitis patients attending Department of Gastroenterology, PGIMS Rohtak.
Material and Methods
The present study was undertaken in department of Gastroenterology, PGIMS (Rohtak), over a time period of five year. A total of 10000 Anti HCV-Ab positive patients was included in this study.
Inclusion Criteria - Patients above 18 years reporting to the Gastroenterology department with Anti HCV antibody positive and willing to participate in current study by giving informed consent, were included in the study.
Exclusion criteria
The study excluded:
a) Patients below 18 years of age, Pregnant women
b) Co-infection with HIV, Patients not willing to participate.
Blood sample collection
Patients who fulfill the inclusion criteria of the current study and gave informed consent to participate in the study were enrolled in the study. The relevant data (personal information, clinical details and history of risk factors) of the patients were collected on patient proforma. Then 5 ml of blood sample of these patients were collected by venipuncture in vacutainer (red capped) after taking all aseptic measures [11]. The collected blood sample were kept at room temperature for 2 hours for clot formation and then sample were centrifuged at 3000 rpm for 5 min to separate the serum.
The serum was then subjected to ELISA for HCV antibody detection using HCV-Ab ELISA kit following manufacturer’s instruction (the ELISA kit was selected having the sensitivity and specificity of 100%). The samples positive for anti-HCV Ab were further tested by real time-polymerase chain reaction (rt-PCR) test for molecular confirmation and genotyping. HCV RNAs from the serum sample were extracted using Qiamp mini kit (Qiagen) on Qiacube automated extraction system following manufacturer’s instructions. The extracted RNA was then subjected to rt-PCR test. Viral load of the samples was assessed using HCV rt-PCR kit on ROTOR Gene Q rt-PCR system, following manufacturer’s instructions. The samples positive for HCV-RNA by rt-PCR were further tested for genotyping of HCV using HCV genotyping kit following instruction provided by the manufacturer.
Disposal of waste
All the biomedical waste generated during this study in the laboratory was discarded after proper disinfection or sterilization as per the Biomedical Waste Management and Handling rules 2016 guidelines, and Bio-Medical Waste Management (Amendment) Rules 2018 [12,13].
Observations & Results
All the ten thousand patients who were found to be Anti HCV Antibody positive and fulfilled Inclusion criterion including willful consent were enrolled in study and their blood samples were tested for baseline HCV RNA Quantitative and Genotyping.
Out of ten thousand patients enrolled in the study, sixty percent (60%) were males and forty percent (40%) were females. Maximum number of patients i.e. 45% were in younger age group i.e. from 20-40 years of age. Majority of patients (62%) had rural background in comparison to 38% of patients belonged to urban areas.
Out of total of ten thousand patients, most common Genotype was 3 which was seen in 65% (13,000 patients) followed by Genotype 4 which was present in 15.88 % (3176 patients). Genotype 1 was seen in 11% (2200 patients), Genotype 5 was seen in 0.06% (12 patients), Genotype 6 in 0.05 % (10 patients) and Genotype 2 in 0.01% (2 patients). The Genotype could not find to be in 1 % (200 patients) due to low viral load i.e. Indeterminate group. There were 7% (1400 patients) in whom baseline HCV RNA Quantitative, no virus was found and hence Genotype test was also negative (Figure 1).
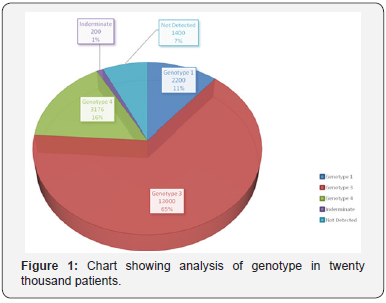
It has been already reported that in India there is North- South divide i.e. Genotype 3 is predominant in North India whereas Genotype 1 & 4 is common in South India. Our study also confirms the same but normally in North India Genotype 1 is more common than Genotype 4 but in our study it is reverse and majority of patients belonging to Genotype 4 belong to one pocket of Ratia and Fatheabad which borders Punjab. The second significant observation is regarding 7% of patients who had baseline anti HCV antibody test positive but had negative pre-therapy HCV RNA quantitative test, hence never required treatment for Chronic hepatitis C. This 7 % figure is significantly higher than average 2 % of patients who had Anti HCV antibody positivity but in whom HCV RNA Quantitative is not detectable. This can be due to either lesser accuracy of rapid diagnostic test kit which are used to test baseline anti HCV antibody test or due to repeated testing got done by patients in screening camps who had already been treated in past for Chronic Hepatitis C.
Conclusion
There are six distinct major genotypes (genotype 1 to 6) and a minor genotype 7 and more than 50 subtypes within the genotypes of HCV have been identified. As the HCV virus replicates rapidly, it constantly changes and mutates thus making it difficult to develop vaccine against HCV. Genotyping is useful for investigating outbreaks and for understanding the epidemiology and virological features of this virus. Accurate classification of genotypes and subtypes of HCV is important for correct stratification of groups and accurate analysis of data related to efficacy and resistance of new HCV drugs. It is also essential for the implementation of therapeutic procedures, the production of effective vaccines, and the improvement of diagnostic tests. Our study confirms the finding of previous studies that in India Genotype 3 is most common genotype which is difficult to treat Genotype with oral antiviral therapy. Hence as per World Health Organization target of elimination of Hepatitis C till 2030 requires continuous, persistent and determined efforts at level of Center, State and society level.
References
- (2018) National Viral Hepatitis Control Program operational guidelines. Ministry of Health and Family Welfare (MOHFW) Government of India.
- Petruzziello A, Marigliano S, Loquercio G, Cozzolino A, Cacciapuoti C (2016) Global epidemiology of hepatitis C virus infection: An up-date of the distribution and circulation of hepatitis C virus genotypes. World J Gastroenterol 22(34): 7824-7840.
- (2018) National Laboratory Guidelines for Testing of Viral Hepatitis Control. Ministry of Health and Family Welfare (MOHFW) Government of India.
- Shah SR, Rao PN, Sarin SK, Chowdhury A, Bhatia S, et al. (2016) Chronic hepatitis C virus infection in India. Regional demographics and distribution of viral genotypes. Ind J Gastroenterol 35(6): 469-477.
- (2018) National Guidelines for Diagnosis and Management of Viral Hepatitis. Ministry of Family and Health Welfare (MOHFW) Government of India.
- (2018) WHO. Hepatitis C.
- (2018) WHO. Combating Hepatitis B and C to Reach Elimination by 2030.
- Bhattacharjee D, Mukherjee K, Chakroborti G, Ghosh R, Mandal N (2015) HCV genotype and viral load. Journal of Clinical and Diagnostic Research 9(5): 15-17.
- Chakravarti A, Ashraf A, Malik S (2013) A study of changing trends of prevalence and genotypic distribution of hepatitis C virus among high risk groups in North India. Indian J Med Microbiol 31(4): 354-359.
- Verma R, Behera BK, Jain RB, Arora V, Chayal V, et al. (2014) Hepatitis C: A silent threat to the community of Haryana, India: A community-based study. AMJ 7(1): 11-16.
- Colle JG, Miles RB, Watt B (2014) Specimen collection, culture containers and media. In: Collee JG, Fraser AG, Marmion BP, et al. (Eds), Practical medical microbiology. 14th (edn), New Delhi: Churchill Livingstone, Elsevier, pp. 95-112.
- (2018) Ministry of Environment and Forests: draft biomedical waste (management and handling) rules. 2016, New Delhi, Govt. of India.
- (2018) Ministry of Environment and Forests: G.S.R 234(E) [16-03- 2018]. The Bio-Medical Waste Management (Amendment)Rules, 2018. New Delhi, Govt. of India.






























