Isolation of Bacillus spp. from Soil for Antimicrobial Production and Antibiotic Resistance
Pushpendra Singh1*, Rajesh Sharma2, Ashish Kumar Shukla1 and Ravindra Singh1
1 Department of Biological sciences, Mahatma Gandhi Chitrakoot Gramodaya Vishwavidhyalaya, India
2Department of Biotechnology, Veer Bahadur Singh Purvanchal University, India
Submission: December 05, 2017; Published: February 28, 2018
*Corresponding author: Pushpendra singh, Department of Biological sciences, Mahatma Gandhi Chitrakoot Gramodaya Vishwavidhyalaya, Chitrakoot, Madhya Pradesh, India.
How to cite this article: Pushpendra S, Rajesh S, Ashish K S, Ravindra S. Isolation of Bacillus spp. from Soil for Antimicrobial Production and Antibiotic Resistance. Adv Biotech & Micro. 2018; 8(4): 555741. DOI: 10.19080/AIBM.2018.08.555741
Abstract
Members of the Bacillus genus are generally found in soil and most of these bacteria have the ability to disintegrate proteins, namely proteolytic activity. Protease enzymes not only have important industrial uses, but also the proteases of these microorganisms play an important role in the nitrogen cycle, which contributes to the fertility of the soil. Bacillus spp. show higher level of resistance in supernatant and methanol. Ethyl acetate showed more effective against E. coli MTCC-1687. The result of this study showed that antimicrobial agent resistance was present in almost all isolates of Bacillus spp. isolated from different sample of soil. Isolates show higher level of resistance or reduced susceptibility to, Ampicillin and Ceftriaxone as there was less effective zone of inhibition observed. This study showed the distribution of antimicrobial agent resistance in Bacillus spp. isolates from a variety of sample and analysis of such pattern of resistance may prove to be useful beyond simple description.
Keywords: Isolation of bacillus; Antibiotic resistance; Antimicrobial production; Ampicillin and ceftriaxone
Introduction
The bacteria are the most abundant group usually more numerous than the four combined. Soil bacteria can be rod, (bacilli) cocci (spherical) spirilla (spirals), of these, Bacillus are more numerous than the others. They are one of the major groups of soil bacteria population and are very widely distributed [1]. The number and type of bacteria present in a particular soil would be greatly influenced by geographical location such as soil temperature, soil type, soil pH, organic matters contents, cultivation, aeration and moisture content [2]. Members of the Bacillus genus are generally found in soil and most of these bacteria have the ability to disintegrate proteins, namely proteolytic activity. Protease enzymes not only have important industrial uses, but also the proteases of these microorganisms play an important role in the nitrogen cycle, which contributes to the fertility of the soil. In paddy field soil, most of the nitrogen source is stored as biomass protein and decomposes slowly to low molecular weight amino acids by the activity of soil protease. Soil protease is thought to be mainly supplied by soil microorganisms. There are numbers of bacteria having potential to produce antibiotic example of which is Bacillus species which produce antibiotic like bacitracin, pumulin and gramicidin which are active against Gram positive bacteria such as Staphylococci, Streptococci, Corynebacter, Streptomyces species which produce antibiotic like tetracycline, chloramphenicol, vancomycin, gentamycin which are active against Gram negative bacteria. Antibiotic resistance is a specific type of drug resistance virtually evolves via natural selection acting upon random mutation, but it can also be engineered by applying an evolutionary stress on a population.
The use of antibiotics along with immunization and sterile techniques in hospitals has significantly decreased the number of lethal bacterial infections. However, antibiotics once seen as miracle drugs are now becoming useless in treating various bacterial diseases. The rapid emergence of resistance to antibiotics amongst pathogen generates visions of the “potential post antibiotics era threating present and future medical advances” [3] the emergence of bacteria resistant to antibiotics is common in areas where antibiotics are used, but antibiotic resistant bacteria are also increasingly occurs in aquatic environment. The widespread use of antibiotics in medicine and in intensive animal husbandry is indicative of selection pressure exerted on bacteria. Intensive animal husbandry causes resistant bacteria to enter the environment directly from liquid manure and muck. There are several reports have also documented the presence, for example, of vancomycin resistant Enterococci (VRE) in the stools of asymptomatic individuals who have neither recently also been found in sewage, from stool of healthy farm animals and animal products, but also in surface water as well as soil.
Materials and Method
Collection and preparation of soil sample
In systematic screening program for isolation of bacteria 20 soil samples were collected at different suburb of Delhi NCR were collected from upper layer where most of the microbial activity take place and thus where most of the bacteria population is concentrated. Soil samples (approximately 10g) were collected using clean dry and sterile container along with sterile spatula. The isolation and identification of Bacillus spp. was done by the technique as per IS 5887 (Pt-6) [4], in test samples. Out of 20 soil samples 18 Bacillus spp. were isolated and confirmed by biochemical test.
Detection and isolation of bacillus from soil sample
A. Step1: Total Bacterial Count from soil sample (Table 1)
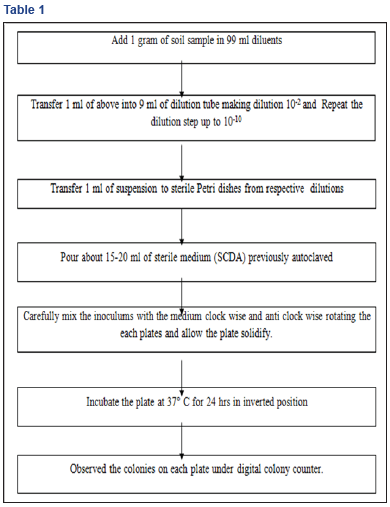
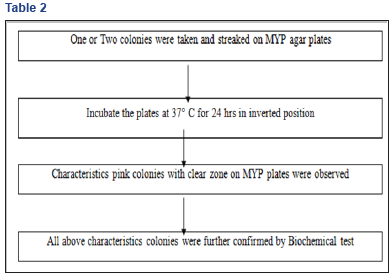
B. Step2: Isolation and Identification of bacillus spp. By MYP Agar Selective media (Table 2)
After biochemical confirmation by standard biochemical tests, confirmed Bacillus spp. isolates were streaked on NA slants and tested for antibiotic producing against E. coli MTCC 739, E. coli MTCC 1687 and S. aureus MTCC 96 and antibiotic sensitivity test against four different antibiotics (Ampicillin, Meropenem, Erythromycin stearate and Ceftriaxone) by agar well diffusion assay method.
Extraction for antimicrobial compound from isolated Bacillus strains
24hrs old cultures were harvested by sterile 0.85% normal saline solution and inoculate 2ml culture in 100ml sterile nutrient broth media flask. Incubate and shake NB flask orbital incubator shaker 37 °C and 160rpm for five days. After 5 days cultures were taken in sterile centrifuge tube from NB flask for centrifugation at 5000 rpm for 15 minutes. After centrifugation supernatant were collected in flasks. For solvent extraction 20ml ethyl acetate and 20ml methanol were added into the remaining biomass in centrifuge tube respectively. After centrifugation at 5000rpm for 15 minutes ethyl acetate and methanol were collected in test tubes and solvent dried by nitrogen gas.
Inoculum preparation
E. coli MTCC 739, E. coli MTCC 1687 and S. aureus MTCC 96 were sub cultured on non selective nutrient agar slants. The bacterial cultures were incubated overnight at 37 °C. 0.5 McFarland density of bacterial isolates was adjusted in normal saline (0.85% NaCl) using densitometer to get bacterial population of 1.0x108 cfu/ml.
Agar well diffusion assay (Zone of inhibition evaluation)
Antimicrobial activity of isolated Bacillus strains was evaluated by agar well diffusion assay. 100μl of each of the adjusted cultures were mixed into separate 100ml of sterile, molten, cool MHA, mixed well and poured into sterile petri plates. These were allowed to solidify and then individual plates were marked for each individual culture. Each plate was punched to make wells of 6 mm diameter with the help of sterile cork borer at different sites of the plates. 100μl of supernatant and ethyl acetate, methanol solutions were pipette into the well in assay plates. Plates were incubated overnight at 37 °C. Following incubation, petri-plates were observed for the inhibition zones, diameters of which were measured by using Vernier Calipers.
Method for Antibiotic Sensitivity Test
Inoculum preparation
All 18 isolated Bacillus cereus culture were sub cultured on non selective nutrient agar slants. The bacterial cultures were incubated overnight at 37 °C. 0.5 McFarland density of bacterial isolates was adjusted using normal saline (0.85% NaCl) using densitometer to get bacterial population of 1.0x108 cfu /ml.
Agar well diffusion assay (zone of inhibition evaluation)
Antibiotic susceptibility and resistance were evaluated by agar well diffusion assay. 100μl of each of the adjusted cultures were mixed into separate 100ml of sterile, molten, cool MHA, mixed well and poured into sterile petri plates. These were allowed to solidify and then individual plates were marked for each individual Bacillus isolates. Each plate was punched to make wells of 6mm diameter with the help of sterile cork borer at different sites of the plates. 100μl of respective antibiotic solutions were pipette into the well in assay plates. Plates were incubated overnight at 37 °C. Following incubation, petri-plates were observed for the inhibition zones, diameters of which were measured by using Vernier Calipers.
Results
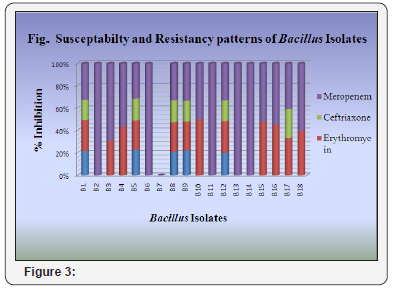
Antibiotic resistance and susceptibility patterns of Isolated Bacillus spp. were checked against four different commonly prescribed antibiotics namely Ampicillin, Erythromycin, Meropenem, and Ceftriaxone by agar well diffusion assay method. Third part was to check antimicrobial activity of isolated strains against E. coli MTCC 739, E. coli MTCC 1687 and S. aureus MTCC 96. Table 3 represented the zone of inhibition in millimeter (mm) of isolated Bacillus strain against test four different antibiotics. Results of antimicrobial activity against E. coli MTCC 739, E. coli MTCC 1687 and S. aureus MTCC 96 were shown in Table 4 (Figure 1 & 2).
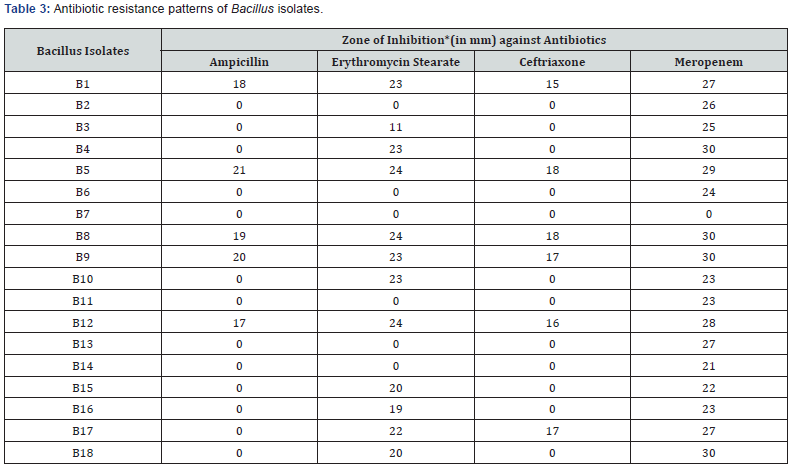
*Zone of inhibition in mm. Diameter including well diameter of 6.0mm.

*Zone of inhibition in mm. Diameter including well diameter of 6.0mm.
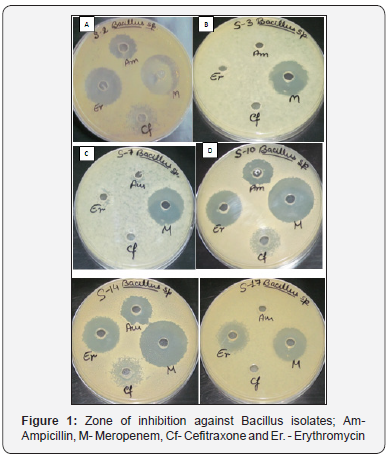
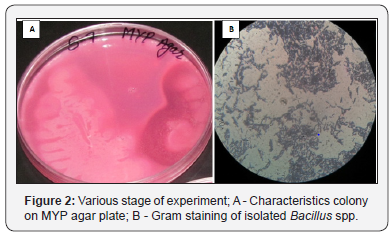
Conclusion
The result of this study shows that many strains of the collection of Bacillus isolated from soil have strong antimicrobial activity against positive culture. Bacillus spp. show higher level of resistance in supernatant and methanol. Ethyl acetate showed more effective against E. coli MTCC-1687. The result of this study showed that antimicrobial agent resistance was present in almost all isolates of Bacillus spp. isolated from different sample of soil. Isolates show higher level of resistance or reduced susceptibility to, Ampicillin and Ceftriaxone as there was less effective zone of inhibition observed. The Bacillus spp. isolates showed maximum susceptibility to Meropenem as the observed zone of inhibition has more effective diameter of 30mm. Isolates showed enter effective susceptibility to Erythromycin. The zone of inhibition measured is Erythromycin 24mm.
This study showed the distribution of antimicrobial agent resistance in Bacillus spp. isolates from a variety of sample and analysis of such pattern of resistance may prove to be useful beyond simple description. As concern about water quality and environmental contamination by human and agricultural waster have increased it has become increasingly important to develop low cost screening tools that can be used to identify the most probable contamination. The distinct pattern of antimicrobial agent resistance may prove to be a valuable tool for the development of multivariate statistical techniques for bacterial sources identification. Based upon these studies, it was found that most of the isolates in the present study show multiple tolerances to antibiotics.
Acknowledgement
The research was supported by Department of Biological sciences, Mahatma Gandhi Chitrakoot Gramodaya Vishwavidhyalaya, Chitrakoot satna (M.P.) India.
References
- Bhagabati A, Dillar T, Grisel N, Sladic Radez G, Mandic Mulec (2004) The influence of Bacillus substilis protein Degu, sin R and sin IR on biosynthensis in Bacillus licheinformis. Biotechniske falk V Iybijani, Knetistro, 200 Technics, 72: 37-42.
- Davies C, Williams B (1999) Genus Bacillus. In: Sneath PH (Ed.), Bergeys Manual of Systematic Bacteriology. Williams and Wikins Company, Baltimore, USA.
- Wise R (2008) The worldwide threat of antimicrobial resistance. Curr Sci 95: 181-187
- Reaff (2005) Identification, Enumeration and confirmation of Bacillus cereus. IS 5887(Pt-6):1999.






























