The New Concepts for Leukemia Probably Especially in Chronic Myeloid Leukemia (CML) And Atypical CML (aCML)
Ahmad Reza Rahnemoon*
Department of Hematology, Iran University of Medical sciences, Iran
Submission: April 09, 2018; Published: May 03, 2018
*Corresponding author: Ahmad Reza Rahnemoon, Department of Hematology, Iran University of Medical sciences, Iran.
How to cite this article: Ahmad Reza Rahnemoon.The New Concepts for Leukemia Probably Especially in Chronic Myeloid Leukemia (CML) And Atypical CML (aCML).Open Acc Blood Res Trans J. 2018; 2(2): 555585. DOI: 10.19080/OABTJ.2018.02.555585
Abstract
While leukemia has been recognized as a genomic disease, we must attention in epigenetic mechanisms in these patients. Thus, the active epigenetic marks particularly in histone protein modification and in other biological phenomena can be important in injury of any progenitor or precursor cell particularly hematopoietic stem cell (HSC) at BM microenvironment. Totally, every oncogenic event with epigenetic modifier or any cancer event can be deregulated the cells of BM niche. Here, I try to submit the role of HSC and how subverted in these diseases..
Keywords: Chronic myeloid leukemia (CML); Atypical CML (aCML); Hematopoietic microenvironment; Hematopoietic stem cell (HSC)
Abbrevations: CML: Chronic Myeloid Leukemia; aCML: Atypical CML; HSC: Hematopoietic Stem Cell; MDS: Myelodysplastic Syndrome; BMT: BM Transplantation; HSCT: HSC Transplantation; m-BCR: Minor Breakpoint Region; MBCR: Major Breakpoint Region; LTHSC: Long-Term Hematopoietic Stem Cell; ALL: Acute Lymphoblastic Leukemia; TKI: Tyrosine Kinase Inhibitor; CP: Chronic Phase
Introduction
When the process of cell production is unbalanced, leading to an exacerbated and uncontrolled proliferation of blood progenitor cells, so leukemia may develop. In addition, leukemia is the consequence of stepwise genetic alterations that confer both proliferative and survival advantage, as well as self-renewal capacity to the malignant cells.
In CML, BCR-ABL fusion proteins can transform in hematopoietic progenitor in vitro. Nonetheless, the mechanism(s) by which P210 BCR-ABL1 promotes the transition from benign state to the fully malignant one is still unclear [1-3].
In mice, reduced BM and increased splenic LTHSC (long-term hematopoietic stem cell) in the effect of BCR-ABL fusion gene, suggest to altered LTHSC trafficking and/or niche requirements. Also, murine models have demonstrated that disruption of the hematopoietic microenvironment can initiate myeloproliferative disease and even leukemia.
Therefore, some researchers stated, restoration of normal niche interactions could play a role in leukemia control in the physiological setting.
In CML, BCR/ABL1 oncogene detected in some hematopoietic lineages, indicating that the cell of origin in thePSCs (pluripotent stem cells) with multi lineage differentiation potential as well. Also, malignant HSCs and the other progenitor cells reside not only on the bone marrow but also in extra sites like spleen that create a niche with a predominance of the neoplasm cells which contribute to the generation of malignant cells and go to the progression disease. In addition, many patients with hyper-cellular marrows, at least in part of physical replacement of normal marrow precursors by leukemic cells [2-4]. Thus, in hematologic malignancies, clonal neoplastic cells alter the hematopoietic microenvironment so that it becomes supportive of LSCs and becomes less supportive of normal HSCs, ultimately leading to decreased normal hematopoiesis. In addition, leukemia induced abnormalities in cytokine in CML bone marrow result in selective suppression of normal stem cell growth and enhanced growth of LSC.
Discussion
In CML, we know some of variant translocations involving additional genes and cryptic translocations requiring molecular detection occur. Most translocations involve a BCR breakpoint at the major breakpoint region (M-BCR) with resultant p210 protein, and occasional breakpoint at the u region results in a p230 protein. Minor breakpoint region (m-BCR) translocations with resultant p190 are usually associated with acute lymphoblastic leukemia (ALL), but rare m-BCR translocations in CML are associated with increase monocytosis, and a small amount of p190 may be found in standard CML. My question is:
i. In this case, we have two or three different lineages, different breakpoints but their proteins and their actions are the same. Is it possible?
ii. In these translocations, we have three different breakpoints and three types of proteins as well but their results are the same. Is it correct?
In CML therapy,
i. Before tyrosine kinase inhibitor (TKI), in chronic phase (CP) of CML, some patients after bone marrow transplantation, the survival of these patients were long and may be cured.
ii. Despite the undeniable success of TKI in treatment of BCR-ABL dependent leukemia, the issue of BCR-ABL positive stem cells that persist in the blood of patients with major molecular remission or complete molecular remission remains unresolved.
iii. Late treatment failures may be observed related to the persistence of leukemic stem cells.
So, we can say, although TKI in treatment of CML, corrects several abnormalities in cytokine and chemokine expression in the cells and reduces normal LTHSC inhibition by leukemic cells and facilities their regrowth, it does not completely reverse leukemia associated changes in the microenvironment. As we know, our data indicate that the frequency of BCR-ABL+, CD34+, CD38- cells rarely remained the blood of patients in remission. This is in agreement with the data showing that some of patients relapsed after TKI discontinuation. Regarding this point, It will be important to determine the mechanisms underlying these persistent the microenvironment changes and how leukemia related alterations in the hematopoietic microenvironment affect Leukemia stem cell (LSC) response to TKI treatment, regrowth of normal LTHSC, and persistence of LSC in CML patients on prolonged TKI treatment. Thus, if we want that some of patients will achieve deep molecular remission and in these cases TKI treatment, discontinue and for the problems as well as the questions above, We should be to understand how leukemic hematopoiesis disrupts the normal mechanisms controlling HSC function and blood production [1,4,5]. So, here in below, I have some points:
i. HSC are multi-potent stem cells defined by their ability to self renewal, differentiation and maintenance of all blood cell types in the hematological system during the entire lifetime of the organism. This physiological process called hematopoiesis is controlled by several complex interactions between genetic process in blood progenitor cells and bone marrow microenvironment. Some researchers stated that leukemic hematopoiesis turns the endosteal BM niche into a leukemic niche, which promotes LSC function and impairs the maintenance of normal HSCs that may be to contribute to myelofibrosis progression. These actions expand our understanding of the effects of leukemic hematopoiesis on the BM microenvironment and the contribution of the endosteal BM niche to MPN pathogenesis, which mainly affects normal HSCs, with minimal effects on transformed LSCs [1,4,6].
ii. Some investigators believed that, the loss of normal HSCs also probably contributes to the clonal dominance of BCR-ABL expressing LSCs in transplanted mice and patients with CML. Regarding we can say, in blood cancers, as CML, which are sustained by LSCs, that like normal HSCs, a range of biological characteristics that enable their long term survival, and accumulation of clonal myeloid cells that differentiate normally, that we know there is no consensus about the mechanisms by which the HSCs transformation occurs [4,7,8]. So, such differential retention provided by a remodeled BM microenvironment could explain clonal dominance in situations were transformed HSCs have impaired functions, such as in myelodysplastic syndrome (MDS) and BM failure syndrome [4,5,9].
iii. For more understanding, I state some points as follows
• In murine models, activation of the normal HSC niche improves recovery from radiation and chemotherapy injury and suppresses CML disease progression, impairing LSC maintenance in a syngeneic model.
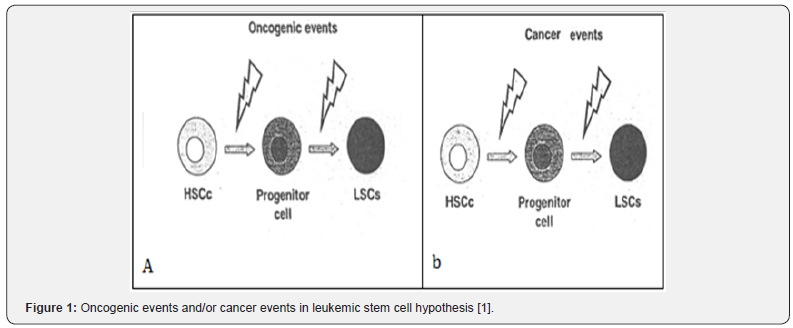
• Myelodysplastic syndrome (MDS) can be cured by allogeneic HSC transplantation and stromal cells remain of patient origin after this procedure, it has been postulated that alterations in MDS stroma must be secondary to interactions with clonal MDS cells and reversible upon their eradication [9,10]. Regarding this point, we know normal hematopoiesis depends on critical interactions that occur between stem cells and their microenvironment, so we can say, hematopoietic neoplasms are known or assumed to be clonal processes arising as the result of genetic errors. In fact, LSCs results from multiple genetic and epigenetic alterations within hematopoietic stem cells or progenitors that alter their normal self renewal, proliferation, differentiation and apoptotic pathways (Figure 1).
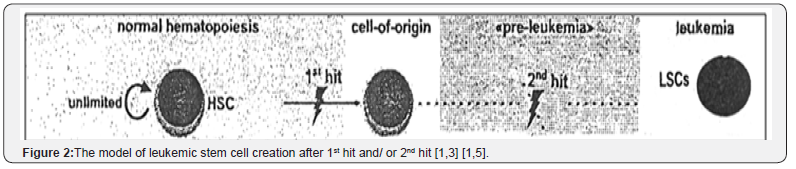
But the question is, how is the action of genetic error? Or how is the management of fusion gene? Before and after the fused? In leukemic stem cell model, after 1st hit to normal hematopoiesis, the cell of origin may be to change to LSC and after the clonal expansion (may be in 2nd hit) LSC enter to pre leukemia (unlimited phase) and in a pre-leukemic disease phase, genetically unstable, self-renewing LSCs clonally expand, facilitating the acquisition of further mutations and the development of different leukemic clones (Figure 2).
But the other question is, what is the agent of 1st hit and in the period of clonal expansion and change to leukemic progenitors and leukemic myeloid cells? What happen the period? As we know, the main bone marrow niche cells including: HSCs, adipocytes, MSCs, reticular cells, osteoclasts, and osteoblasts. In my discussion, the main role of bone marrow niche cells depends on HSCs. Regarding, I state some viewpoints of the researchers as follows:
i. Some rearrangements like ETV6/RUNX1 in neonatal could arise in a high proportion of developing fetuses but without the production of functional chimeric proteins, alternatively they could originate inappropriate cellular context but chiefly, I concentrated on the non-production of proteins. In addition, less than 1% of these children with ETV6/RUNX1 lastly go to ALL [8,11]; but why? Persistent infection may be helped to post natal genetic lesions.
ii. In order to produce a leukemic phenotype, these rearrangements should fulfill two conditions:
• The structure of the fusion gene must allow the production of a functional protein.
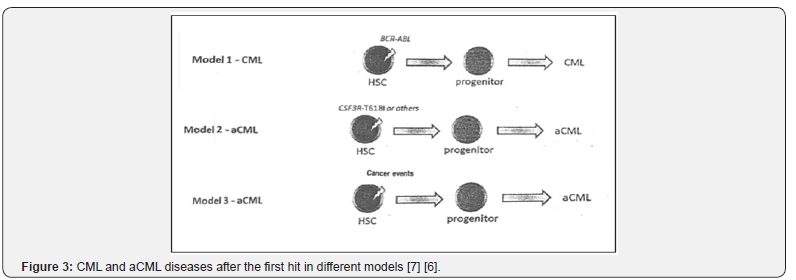
• The translocation must occur in early precursors with self-renewal capacity. Now, the question is, how is disrupt normal BM and impair the cell production? As we know, there are many genetic alterations or chromosomal aberrations in malignant diseases. Some of them, is permanent (BCR-ABL in CML, etc.), some of them is temporal (the translocations in AML or in ALL, etc. occasionally) and the other may be randomly (some of trisomy or monosomy, hyper-diploid and hypo-diploid in patients). For continue discussion, it is better to say about BCR-ABL fusion gene which seen in more than 95% of classic CML [12] patients as follows Figure 3.
The action of BCR-ABL producing a protein that destruct the normal cell and change the HSC to LSC (1st hit), After the clonal expansion and increased myeloid differentiation and may be in 2nd hit, we see the unlimited phase that leukemic myeloid cells increased [3,4]. So feedback of this model in leukemic bone marrow niche is decreased HSC maintenance. In other words,we think any action of genetic aberration can occur in first hit which follow in 2nd hit possibly, but perhaps the effect of action inany fusion gene and other agents like cancer events can be autonomous and simultaneously. My point is the start of first hit with cancer events possibly and after that a genetic alteration may be occurred in second hit. Thus, we accepted, CML is a malignancy resulting from the transformation of HSCs by the BCR-ABL fusion gene in most cases.
Concerning in the model, MSC activated and perhaps OBCs involved. So, in malignant pathway, BM microenvironment should be disrupted partially and impaired the normal cell production [13]. This is the CP-CML. After chemotherapy and BM transplantation (BMT) in CP, some patients had a long survival and may be cured (before TKI). So what happen for the CP- CML patients after BMT? Also after BMT in MDS? Some patients with MDS [6], and patients with other leukemia like CML and ALL may be cured. Thus, after BMT, BM niche is rebuilt or completely repaired. I believe, the structure of bone marrow niche must intact, and after the impair of hematopoiesis, should be changed ,which means BMT or HSC transplantation (HSCT) can repair BM structure that maintain HSCs chiefly and the other cells of BM niche as well. But the question is about the role of genetic aberration in aCML [14,15]. In aCML, we have BCRABL negative fusion gene that clinical features resembles with BCR-ABL positive CML patients but has a worse overall survival. This is a kind of CML without BCR-ABL or no production of the protein. So what happen in aCML? In response, the stem cell or progenitor cell is upset directly, transformed and change to the LSC (unexpected result in stem cell) (Figure 4).
This subject is resemble with some of leukemias without any known fusion gene or translocation. In this CML, may be another genetic aberration or transient genetic abnormality (temporal or randomly or accidentally), but its result is the same. In other words, we have a CML with the same clinical feature and cell morphology. For more understanding, I would like to state several viewpoints as follows which some of them may sparsely because of I must explain the subject totally:
i. In the study pointed out the positive interphase nuclei which in using interphase - FISH technique was higher than the translocation of FISH positive metaphase nuclei [16,17].
ii. Dysfunctional nuclear membrane components can be important in some disease like leukemia [18]. In fact some researchers believed nuclear protein complex bind super enhancers which regulatory structures that drive the key genes expression for identify of specify cell and they mentioned nucleus of cell can be as a driver of gene expression and perhaps a key point in some disease like leukemia.
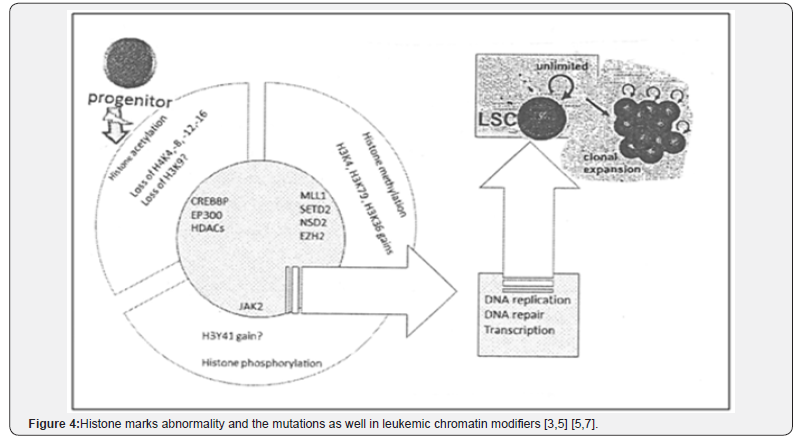
iii. In the study point out, the necessary role of histone protein modifications including acetylation, methylation and phosphorylation in hematologic cancers particularly in altered histone marks and histone modifiers. Regarding in this research the role of histone modifiers is embossed [5,8] (Figure 5).
iv. In stem cell dynamics, in the symmetric and asymmetric divisions and in expansion and maintenance of the cells, some of these cells feedback to quiescent stage and eventually in this long period, perhaps do any mistake in the divisions particularly in feedback the cells to quiescence stage and then my question is, what happen? [11,19] (Figure 6,7).
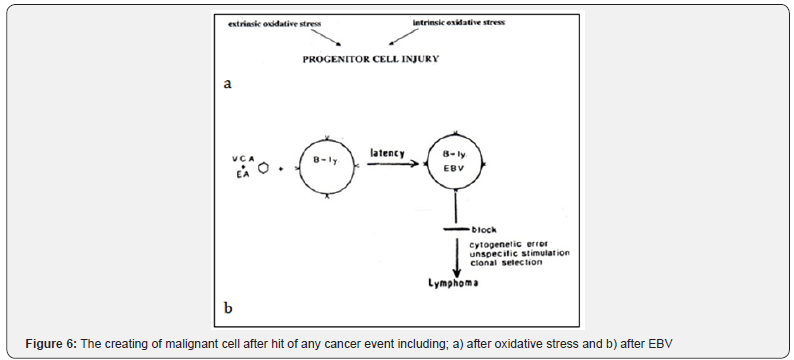
v. What is the agent of change in EBV (Epstein Barr virus) to malignancy? Viruses or transient genomic instability (or accidentally genetic error)? In response to the question, seizing in hematopoietic microenvironment pathway control and their cellular functions and diverse and deregulate of the pathway and its microenvironment. Therefore, induce both lytic and latent infections that result in some diseases including lymphoproliferative disorders. In other words, the exact role of EBV is unclear in this area but we know after the infection and latency and possibly a block or hit in EBV disease pathway, cytogenetic error and clonal selection is occurred and the disease change to lymphoma [19-21].
vi. As we know, some leukemic fusion genes detected in healthy individuals. How? [26] In response to the question, I mention it in my discussion and Figure 6 & 7 as well.
In fact, normal blood cell development is regulated by transcription factors and other internally regulators and also growth factors and cellular interactions in the hematopoietic microenvironment as well. Thus, transcription factor is one of these regulators and cytokines as well as interactions of bone marrow cellular are the other important factors in the delineating of leukemia processing [23-25].
At the present time, it is better to give a more activated role for HSCs niche or hematopoietic microenvironment in the malignant diseases, particularly how the cells are subverted and do to drive toward a malignancy (Figure 6 & 7). Finally, in CML, it is important to determine that how do the alterations in the hematopoietic microenvironment in response to TKI therapy, regrowth of normal HSCs, and how do the persistence of LSC on prolonged TKI treatment? In response, if we concentrate to role of HSC and HSC maintenance in BM niche with the consideration of fusion gene, may solve it (like the role of BMT in CP-CML) [26- 29].
Conclusion
CML is a uncontrolled clonal proliferation of pluripotent stem cells in the bone marrow. Advances in leukemic genomics have revealed the spectrum of mutations that give rise to CML and aCML. But in some cases, the mutations are not the first step in the start of leukemia possibly. In other words, according to the models in my discussion and as we know the hematopoietic microenvironment is important on tumor growth dynamics, but in a few cases cancer events may be the leukemic first step(1st hit) with or without oncogenic events (Figures 1-3) which can be as a 2nd hit possibly. Thus, in this case HSCs or other progenitor cells can be subverted by cancer events and after upset the cells (Figure 4-7), BM microenvironment is remodeled initially which with clonal dominance in the situation where transformed to injury progenitor cell (from cell membrane to inside of cell particularly in nuclear membrane components), then impaired function with additional abnormality that create a self-reinforcing leukemic BM niche which the malignant HSC is embossed in this condition. At any rate expansion of precursor cell populations, resulting in progression of disease through combinational effects including any event like cancer event or oncogenic event, additional mutations, resulting drive to leukemia in the process of leukemic clonal evolution. Here, we can understand the role of fusion genes like BCR/ABL, ETV6/ RUNX1 and MLL in normal individuals and we understand the skill and ability of malignant HSCs to relation with molecular analysis mostly and the hematopoietic microenvironment as well in heterogenous cell populations. In other words, fused genes can be in healthy individuals without any subvert in HSCs or other progenitor cells and without following additional abnormal features in BM niche cells and lastly without any malignant action and leukemic effect as well.



References
- Bydlowski SP, e Lara Janz F (2012) Hematopoietic stem cell in acute myeloid leukemia development. In: PelayoR (Eds.) Advances in hematopoietic stem cell research. In Tech Europe pp. 261-276.
- Wetzler M (2012) Acute and chronic myeloid leukemia. In: Longo DL, Fauci AS, et al. (Eds.) Harrison’s principles of internal medicine (18th edn). McGraw- Hill Company, Newyork, USA 1: 905-918.
- Riether C, et al. (2015) Regulation of hematopoietic and leukemic stem cells by the immune system. Cell Death and Differentiation 22(2): 187- 198.
- SchepersK, Pietras EM, Reynaud D, Flach J, Binnewies M, et al. (2013) Myeloproliferative neoplasia remodels the endosteal bone marrow niche into a self-reinforcing leukemic niche. Cell 13(3): 285-299.
- Janczar S, et al. (2017) The role of histone protein modifications and mutations in histone modifiers in pediatric B- cell progenitor acute lymphoblatic leukemia. Cancers 9(1): 1-14.
- Zhang B, Ho YW, Huang Q, Maeda T, Lin A, et al. (2012) Altered microenvironmental regulation of leukemia and normal stem cells in chronic myelogenous leukemia. Cancer Cell 21(4): 577-592.
- Dao KHT, Tyner JW (2015) What’s different about atypical CML and chronic neutrophilic leukemia? Hematology pp. 264-271.
- Grassinger J, Mueller G, Zaiss M, Kunz-Schughart LA, Andreesen R, et al. (2006) Differentiation of hematopoietic progenitor cells towards the myeloid and B- lymphoid lineage by hepatocyte growth factor(HGF) and thrombopoietin(TPO) together with early acting cytokines. Eur J Haematol 77: 134-144.
- Delforge M (2003) Understanding the pathogenesis of myelodysplastic syndromes. The hematology Journal 4: 303-309.
- Balderman SR, Calvi LM (2014) Biology of BM failure syndromes: role of microenvironment and niches. Hematology p. 71-76.
- Gawad C, Koh W, Quake SR (2014) Dissecting the clonal origins of childhood acute lymphoblastic leukemia by single cell genomics. PNAS 111(50): 17947-17952.
- Babashah S, Azodi MZ, Saki N (2012) Chronic myeloid leukemia as a stem cell-derived malignancy. Journal of Paramedical Sciences 3: 43- 55.
- Jaganthan BG, Tisato V, Vulliamy T, Dokal I, Marsh J, et al. (2010) Effects of MSC co-injection on the reconstitution of aplastic anemia patient following hematopoietic stem cell transplantation. Leukemia 24(10): 1791-1795.
- Giri S, Pathak R, Martin MG, Bhatt VR (2015) Characteristics and survival of BCR/ABL negative chronic myeloid leukemia: a retrospective analysis of the surveillance, epidemiology and end results database. Ther Adv Hematol 6(6): 308-312.
- Wang SA, Hasserjian RP, Fox PS, Rogers HJ, Geyer JT, et al. (2014) Atypical chronic myeloid leukemia is clinically distinct from unclassifiable myelodysplastic/myeloproliferative neoplasms. Blood 123(17): 2645-2651.
- Yakut T, Ali R, Egeli U, Ozkalemkas F, Ercan I, et al. (2004) Comparison of genetic changes between interphase and methphase nuclei in monitoring CML and APL treatment using DC- FISH technique. Cancer Biology and Therapy 3: 858-863.
- Khajehmarjany MS, Rahmani SA1, Chavoshi SH2, Esfahani A2, Movassaghpour Akbari AA (2015) Reliability evaluation of fluorescence in situ hybridization (FISH) and G-banding on bone marrow and peripheral blood cells in chronic myelogenous leukemia patients. Cell Journal 17(1): 171-180.
- Ibarra A, Benner C, Tyagi S, Cool J, Hetzer MW (2016) Nucleoprotein mediated regulation of cell identity genes. Genes Dev 30(20): 2253- 2258.
- Martinez- Agosto JA, et al. (2007) The hematopoietic stem cell and its niche: a comparative view. Genes and Development 21(23): 3044- 3060.
- Cai Q, Chen K, Young KH (2015) Epstein Barr virus positive T/NK cell lymphoproliferative disorders. Exp Mol Med 47: e133.
- Wolf H, et al. (1984) Benign and malignant disease caused by EBV. The journal of Investigative Dermatology 83(Suppl 1): 88s-95s.
- Brassesco MS (2008) Leukemia/Lymphoma associated gene fusions in normal individuals. Genetics and Molecular Research 7: 782-790.
- Swaminathan S, Klemm L, Park E, Papaemmanuil E, Ford A, et al. (2015) Mechanisms of clonal evolution in childhood acute lymphoblastic leukemia. Nat Immunol 16(7): 766-774.
- Deshpande AJ, Bradner J, Armstrong SA (2012) Chromatin modifications as therapeutic targets in MLL rearranged leukemia. Trends immunol 33(11): 563-570.
- Luc S, Orkin SH (2013) Delineating the mixed lineage leukemia gene expression network in hematopoietic stem cells. PNAS 110: 11670- 11671.
- Hutchison RE (2017) Leukocytic disorders. In: McPherson RA, Pincus MR (Eds.) Henry’s clinical diagnosis and management by laboratory methods (23rd edn). printed in china, Elsevier; pp. 606-658.
- Cross NCP, White HE, Colomer D, Ehrencrona H, Foroni L, et al. (2015) Laboratory recommendations for scoring deep molecular responses following treatment for chronic myeloid leukemia. Leukemia 29(5): 993-1003.
- Valent P (2007) Imatinib-resistant chronic myeloid leukemia (CML): Current concepts on pathogenesis and new emerging pharmacologic approaches. Biologics: Targets and Therapy 1(4): 433-448.
- Delforge M, Boogaerts MA, et al. (1999) BCR/ABL- CD34+ HLA DRprogenitor cells in early chronic phase, but not in more advanced phases, of chronic myelogenous leukemia are polyclonal. Blood 93(1): 284-292.






























