A Mass Spectrometry Method for the Quantification of Heparan Sulfate
Mario DiPaola1*, Jing Li2 and Elaine Stephens3
1Charier River Laboratories, USA
2Currently at Ainyiam in Cambridge, USA
3Currently at Pfizer in Andover, USA
Submission: March 30, 2017; Published: April 13, 2017
*Corresponding author: Mario DiPaola, Charler River Laboratories, Woburn MA, USA, Tel: 617-234-0001; Email: Mario.Dipaola@crl.com
How to cite this article: DiPaola M, Li J, Stephens E. A Mass Spectrometry Method for the Quantification of Heparan Sulfate. Mod Appl Bioequiv Availab. 2017; 1(1): 555553.
Abstract
Heparan sulfate proteoglycans (HSPG) are present on the cell surface as well extracellular matrix, where they interact with a series of ligands [1]. Like most other macromolecules, HSPG are processed within the lysosome. However, in cases of lysosomal enzyme deficiency, HSPG are not properly digested leading to accumulation of these HSPG and disease state known as mucopolysaccharidoses [2]. We have developed a robust method for the analysis of heparan sulfate from cell extracts by measuring heparan lyase-digested disaccharides. For normalization, the samples were spiked with an internal standard to correct for losses during sample preparation which involved sample clean-up using hypercarb SPE columns. The disaccharide abundances were determined by LC-MS/MS using a triple quadruple mass spectrometer set-up in in selected reactions monitoring mode. We applied this method to measure the relative heparan sulfate disaccharide abundances in three mutants and three wild type cell extracts. The disaccharide amounts in the mutant cells were determined to be approx. 6-7 fold higher than those in wild type cells.
Abbreviations: HSPG: Heparan Sulfate Proteoglycan; LC: Liquid Chromatography; UPLC: Ultra-Pressure Liquid Chromatography; MS: Mass Spectrometry; TIC: Total Ion Chromatogram; SPE: Solid Phase Extraction; SRM: Selected Reaction Monitoring; MPS: Mucopolysaccharidoses; WT: Wild Type
Introduction
Mucopolysaccharidoses are recessive inherited disorders. These are characterized by the failure to degrade glycosaminoglycans, leading to lysosomal accumulation. These disorders are chronic and progressive with a number of clinical manifestations, including skeletal and joint abnormalities, hepatomegaly and splenomegaly, course facial features, neurological complications, hearing and vision loss and cardiovascular and respiratory disorders [3].
In recent years, a number of therapies have been introduced to address these complex diseases. Among the more successful therapies is enzyme replacement therapy (ERT) and hematopoietic stem cell transplantation [4,5]. Unfortunately, the optimization of these therapies has been adversely affected by the lack of reliable and measurable biomarkers. In this paper, we describe a mass spectrometry method that has been shown to specifically and accurately measure disaccharides generated from the enzymatic digestion of heparin sulfate.
Materials and Methods
Standards and sample preparation
Heparin disaccharides standards, I-A, II-A, III-A, IV-A, I-S,III- S and IV-S, were purchased from Galen Laboratories (CT,USA), The standards were weighed and dissolved in HPLC grade water (Fisher Scientific) to 1µg/mL prior to use. Each cell (cell culture not described in this paper) extract, GS1 through GS6 with GS1 through GS3 representing mutants (MPS1) [6] and GS4 through GS6 wild type, was diluted with 84µL of digestion buffer (50mM ammonium acetate and 1mM calcium acetate, pH 8.0), spiked with 10µL of 10µg/mL chondrosine (100ng) as internal standard (IS) and then digested with 4mU of heparinases I, II and III (Galent Laboratories, CT, USA). Digestion was allowed to proceed for 24 hours at 37 °C. Each sample was then cleaned up by passage through hypercarb SPE (Thermo Scientific, MA, USA) columns [7] with the aid of a vacuum manifold. Briefly, each hypercarb SPE column was wash with 2mL of 1M sodium hydroxide (Sigma, MO, USA), followed with 4 mL water, then with 2mL of 30% acetic acid (Sigma, MO, USA), and 2mL water Each SPE column was primed with 2mL of 50% acetonitrile (Honeywell, MA, USA) plus 0.1% trifluoroacetic acid (Thermo Scientific, MA, USA) in water (Solvent A), followed by 4mL of 5% acetonitrile plus 0.1% trifluoroacetic acid in water (Solvent B). A 100uL volume for each sample was loaded into its respective SPE column. washed with 2mL water, followed by 2 mL of Solvent B. The disaccharides were eluted with 4x0.4mL of Solvent A. The eluted disaccharides were dried using a speed-vac, re-dissolved in 100µL of 90% acetonitrile in water and loaded into auto sampler for analysis by LS-MS/MS.
LC-MS/MS
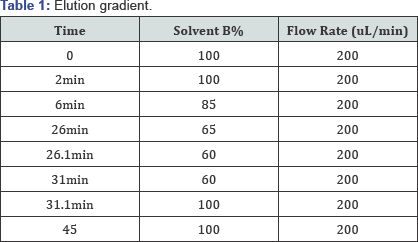
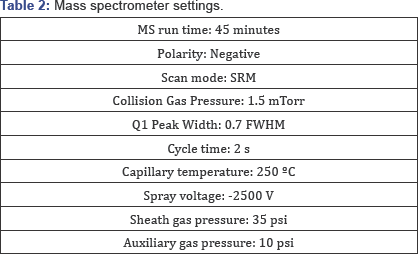
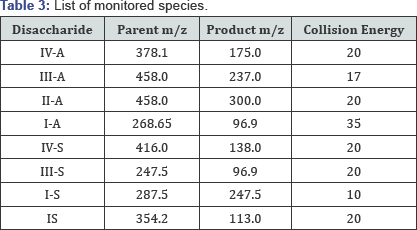
For the analysis by LC-MS/MS, each sample was loaded onto a UPLC BEH glycan column [8] (2.1x500mm), Milford MA, USA) and eluted with a gradient of mobile phases A (12.5mM ammonium format, pH 4.4) and B (90% acetonitrile with 10% mobile phase A) as specified in Table 1. The mass spectrometer (Vantage, Thermo Scientific MA, USA) conditions are shown in (Table 2) for monitoring the species listed in Table 3.
Results and Discussion
The heparinases digested cell extracts containing putative disaccharides (GS-1 to GS-6) were spiked with an internal internal standard (chondrosine) [9], to adjust fro losses during preparation. The disaccharides in each sample cleaned by The disaccharides in each sample were analyzed by LC-MS/MS, after clean-up by hypercarb SPE. The separation of the disaccharides was conducted with a UPLC BEH glycan column. The column eluate was analyzed by a triple quadrupole mass spectrometer set-up in selected reaction monitoring (SRM) [10], in negative mode. Duplicate injections were conducted for each sample to ensure injection reproducibility. Disaccharides standards, which constitute the major building blocks of heparan sulfate chains, were purchased from commercial sources. The monitored SRM transitions, listed in Table 3, were developed from these standards.
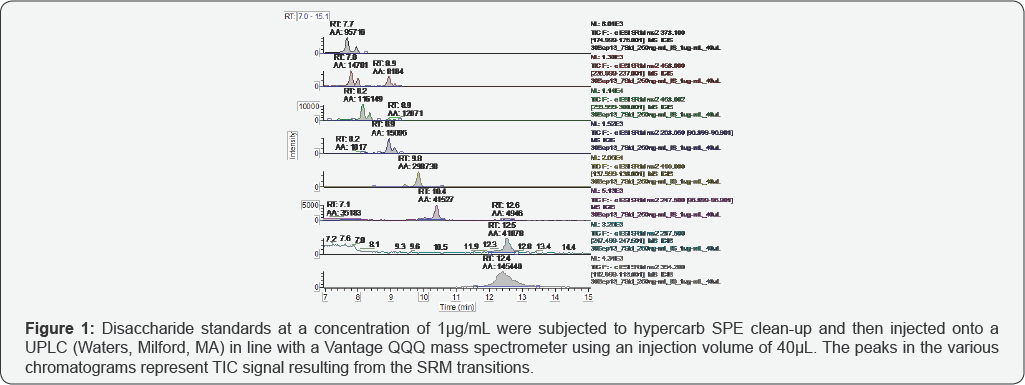
Figure 1 shows the representative standards separated by the glycan column and monitored by SRM. Anomers of each disaccharide were resolved by the glycan column and are observed as two peaks in the chromatograms. Integrated peak area (AA) and retention time (RT) are labeled above each peak. Disaccharides I-A, II-A, III-A and IV-A elute at approximately 8.9, 8.2, 7.8 and 7.7 minutes, respectively. Disaccharide I-A is prone to in-source fragmentation, which converts it to II-A and III-A; therefore, this species was observed in II-A and III-A chromatograms, as well. Disaccharides I-S, III-S and IV-S elute at approximately 12.5, 10.4 and 9.8 minutes, respectively. which converts it to II-S and III-s, thus it was observed in the III-S chromatogram, as well. The internal standard elutes at approximately 12.4 minutes.
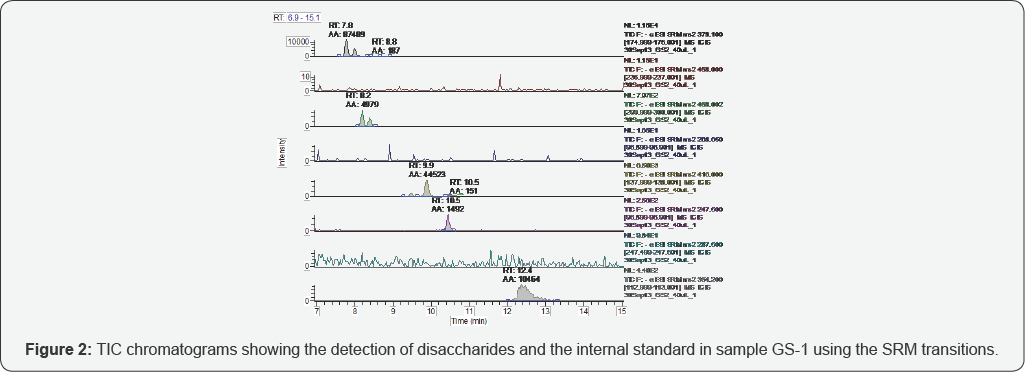
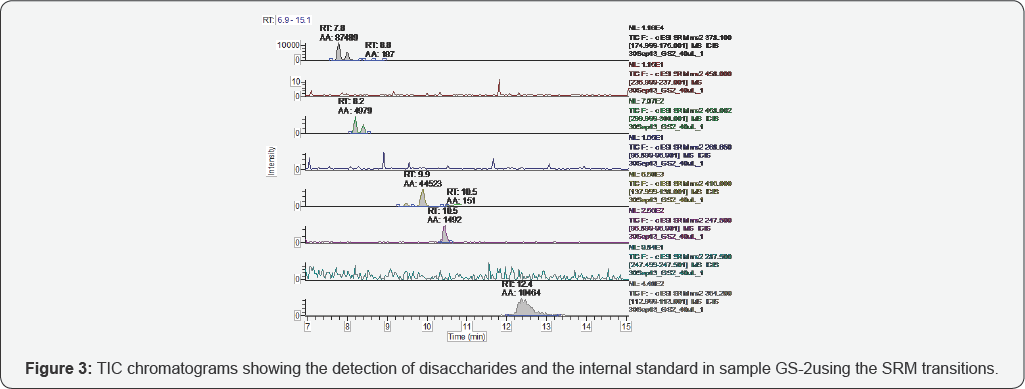
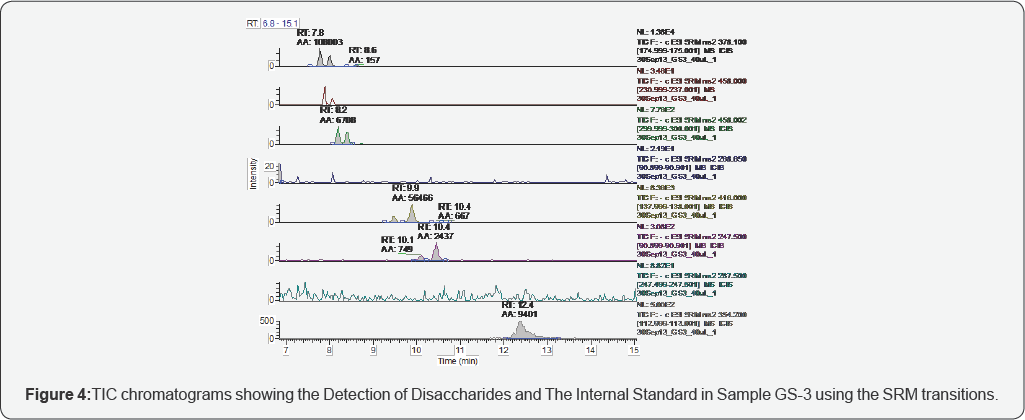
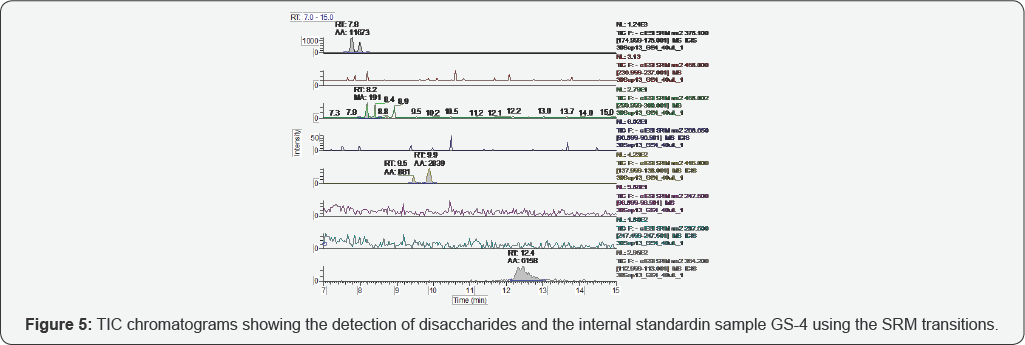
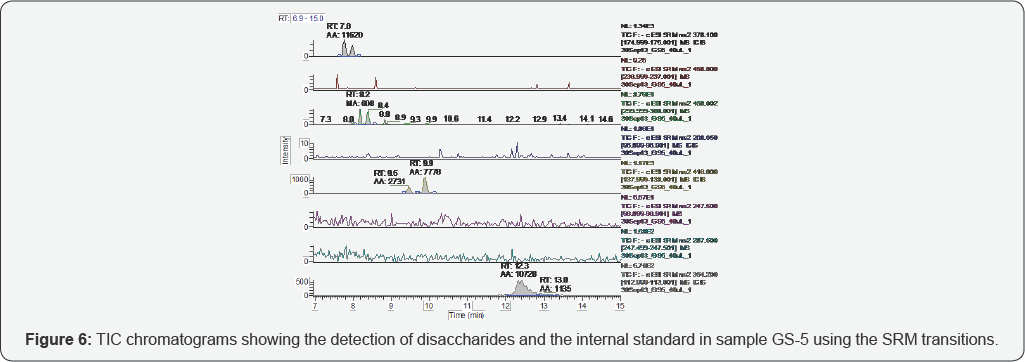
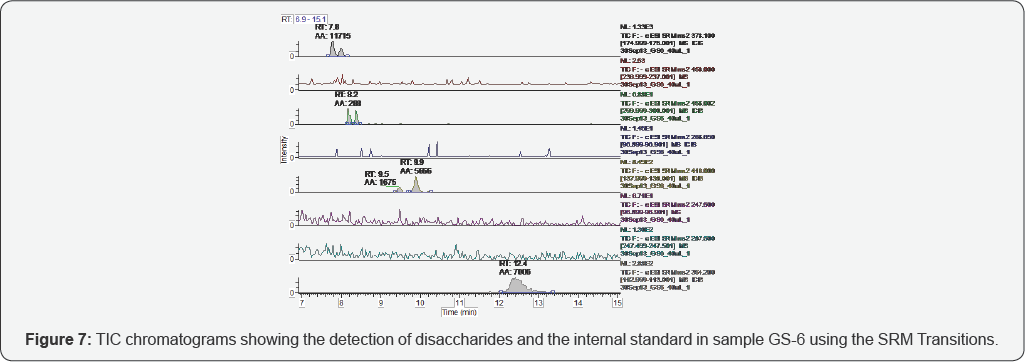
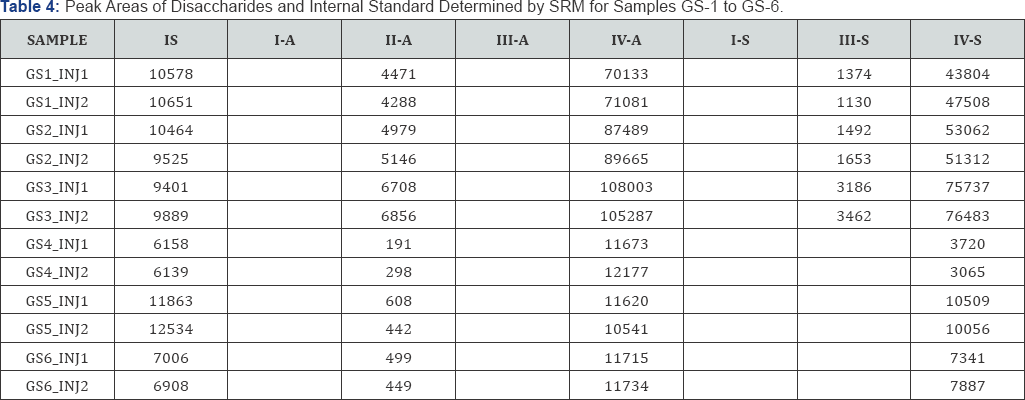
The representative TICs for disaccharides from samples GS-1 are shown in Figures 2-7. Four disaccharides II-A, IV-A, III-S and IV- S were observed in samples GS-1, GS-2 and GS-3 (mutants). Three disaccharides II-A, IV-A and IV-S were detected in samples GS-4, GS-5 and GS-6 (wild type). Disaccharides IV-A and IV-S were the dominant species in all samples, which agrees with the composition of heparan sulfate chains. The observed peak areas of each disaccharide and internal standards are summarized in Table 4.
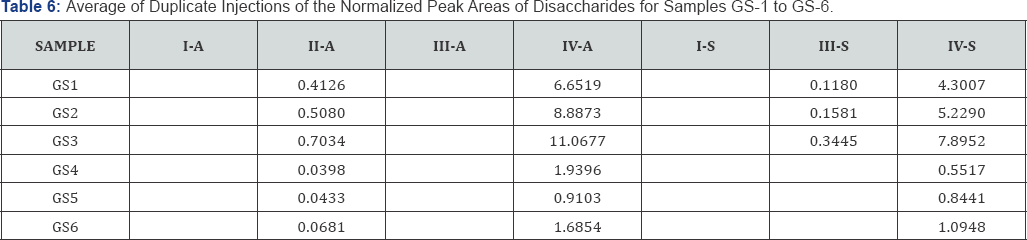
The observed peak areas of the disaccharides in each sample, divided by the peak area of corresponding internal standard (normalized peak areas) are summarized in Table 5. The average normalized peak areas of duplicate injections are summarized in Table 6. It can be seen that the mutant samples (GS1, GS2, GS3) have overall much higher disaccharide abundance than the wild type (GS4, GS5, GS6).
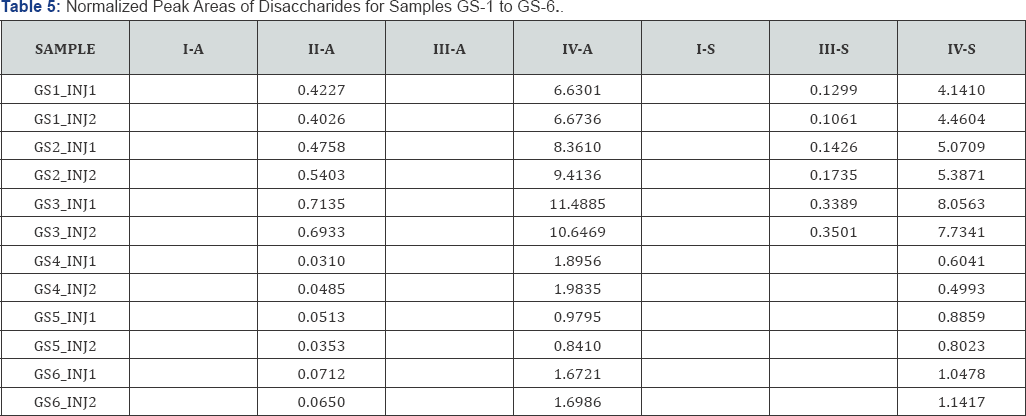
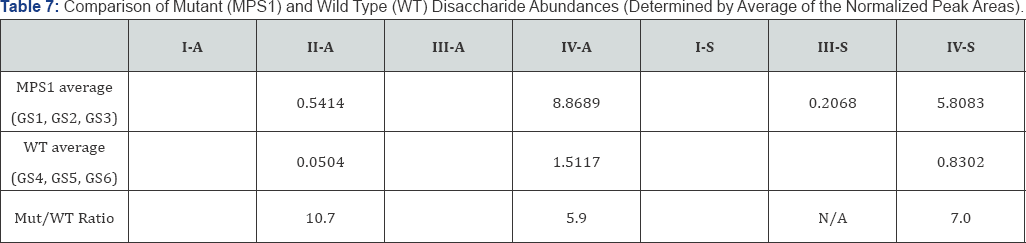
The average peak areas of all three mutants (MPS1) samples and all three wild type (WT) samples, respectively, are listed in Table 7. The MPS1/WT ratios are determined to be 10.7 for II- A, 5.9 for IV-A and 7.0 for IV-S. The disaccharide II-A is in much lower abundance than IV-A and IV-S. Quantization of the low abundance species by SRM is not likely to be as accurate as those in high abundance (IV-A and IV-S). Therefore, the MPS1/WT ratios determined by IV-A and IV-S are likely to be more accurate.
Conclusion
A robust method has been developed for the analysis of disaccharides derived from heparan sulfate from cell extracts. The samples were spikedwith and internal standard, digested and cleaned up using hypercarb SPE columns. The disaccharide abundances present in each of the cell extracts were then detected and quantified by LC-MS/MS. Using this method, we measured the relative disaccharide abundances in three mutant and three wild type cell extracts. The heparin sulfate disaccharide amounts in the mutant cells were determined to be 6-7 fold higher than those in wild type cells, as predicted. These results validate to potential use of this method for quantitative measurement of heparin sulfate as a biomarker for disease state and also as a surrogate endpoint for therapeutic optimization.
References
- Sarrazin S, Lamanna WC, Esko JD (2011) Heparan Sulfate Proteoglycan. Cold Spring Harb Perspect Biol 3(7): a004952.
- Lawrence R, Brown JR, Al Mafraji K, Lamanna WC, Beitel JR, et al. (2012) Disease-Specific Non-Reducing End Carbohydrate Biomarkers for Mucopolysaccharidoses. Nat Chem Biol 8(2): 197-204.
- Neufeld EF, Muenzer J (2001) The Mucopolysaccharidoses. Metabolic and Molecular Basis of Inherited Diseases. San Fransisco: MacGraw- Hill 3: 3421-3452.
- Beck M (2010) Therapy for Lysosomal Storage Disorder. IUBMB Life 62: 33-40.
- Clarke LA (2008) The Mucopolysaccharidoses: a Success of Molecular Medicine. Expert Rev Mol Med 10: el.
- Danes BS, Bearn AG (1967) Cellular Metachromasia, a Genetic Marker for Stdying the Mucopolysaccharidoses. The Lancet 289(7484): 241243.
- Grass J, Pabst M, Kolarich D, Polti G, Leonard R, et al. (2011) Discovery of Structural Characterization of Fucosylated Oligomannosidic N-Glycans in Mushrooms. J Biol Chem 286(8): 5977-5984.
- Ahn J, Bones J, Yu YQ, Rudd PM, Gilar M, et al. (2010) Separation of 2-Aminobenzamide Labeled Glycans Using Hydrophobic Interaction Chromatography Columns Packed with 1.7µm Sorbent. J Chromatogr B 878: 403-408.
- Tomatsu S, Shimada T, Mason RW, Kelly J, LaMarr WA, et al. (2014) Assay for Glycosaminoglycans by Tandem Mass Spectrometry and its Applications. J Anal Bioanal Tech (Suppl 2): 006.
- Yadlin WA, Hautaniemi S, Lauffenburger DA, White FM, (2007) Multiple Reaction Monitoring for Robust Quantitative Proteomic Analysis if Cellular Signaling Networks. Proc Natl Acad Sci USA (14): 5860-5865.






























