The Unveiled Genome and Cancer Management
*Giuseppa Quartarone MD, MSc
Independant Medical Affairs and Clinical Research Consultant, Milan Greater Area
Submission: June 01, 2016; Published: July 18, 2017
*Corresponding author: Giuseppa Quartarone MD, MSc, Email: giuseppina_quartarone@yahoo.com
How to cite this article: Quartarone G. The Unveiled Genome and Cancer Management. J of Pharmacol & Clin Res. 2017; 3(3): 555612. DOI:10.19080/JPCR.2017.03.555612
Keywords: Cancer; Check point inhibitors; PARP; PARP inhibitors; DNA; Trabedectin; Immunotherapy; Molecular testing; BRCA; Platinum sensitivity; Angiogenesis inhibitors; Bevacizumab; Immunological check point; Therapeutic vaccines; Solid tumours
Abbreviations: VEGF: Vascular Endothelial Growth Factor; BRCA; Breast-Cancer susceptibility gene 1-2; PARP: Poly ADP-Ribose Polymerase; PD1: Programmed Death1 Receptor; PD-L1: Programmed Death Ligand; PI3: Kinase Phosphoinositide 3-kinase
Mini Review
The Human Genome Unveiled
In 2001, I was a young clinical research in Clinical Oncology, when Nature would release a special issue devoted to the human genome unveiled: this special issue is always ready-to-use in my home library; it's a source of inspiration and hope because by that time many things have changed and continue to change. Translate what are the results in laboratory research into clinical research and even more in clinical practice, it's a leap, that's not easy and sometimes it's just a leap in the dark. In the same special issue of Nature, MR Stratton and co-workers [1], regarding mutations in oncogenes, stated that we might learn more about the mutations driving cancer if we were not too heavily influenced by past experience; and added that we should persevere in exploring every gene or protein, whatever its structure or putative function, as possible candidate. Regarding the genome sequencing in neoplastic cells, the Authors affirmed that the complexity of cancer, at the genomic level, would have required much more sequence data from cancer genomes which would have needed to be configured appropriately for the task at hand: to facilitate these analyses it would have been needed the finished sequence from which a structural framework would have form for a new generation of massive scale comparison of cancer cells and normal genome. They added that a new technology would have been required and there was no single technology, at that time, capable of detecting all types of abnormality, such as large deletions, rearrangements, base substitutions, small insertions or deletions, amplifications and epigenetic changes, such as methylation. Stratton and co-workers concluded that "sequencing of genomics libraries constructed from cancer genomes would come closest to this goal, but given the diversity of cancers and the effort and cost required to obtain reasonable coverage of human genome this is a daunting challenge".
Assessing new targets starting from the tumours cells
Nowadays the goals of cancer genomics are aimed at enhancing the gene sequencing technology to translate these tools in routine tests, in order to prevent or diagnose cancer, but also to discover new pharmacological targets, following the molecular pathogenesis of cancer that leads to find new classes of drugs or widen and rationalize the use of the existing ones. The role of Poly ADP-ribose polymerase (PARP) pathway in cancer pathogenesis has been explored since the beginning of the new Century; its inhibition is particularly interesting in a subset of patients with germline as well as in those with somatic BRCA [2,3] mutations (BRCAm) associated to breast and ovarian cancers. PARP inhibitors (PARPi's) are new drugs targeting PARP, exhibiting a cytotoxic activity in BRCAm (both germinal and somatic) carriers: BRCAm is a today considered a predictive biomarker for PARP i's response that, from a clinical standpoint, is correlated to platinum sensitivity, [4] a clinical feature that's shared with the so-called BRCAness phenotype, represented by defective HR, secondary to several mechanismsincluding hypermethylation of BRCA1 promoter, somatic mutation of BRCA1-2, loss of function mutations involving HR pathway genes as well as components of the Fanconi anemia repair pathway [5].
The loss of DNA repair in the presence of PARPi’s has led to consider them in combination therapy with cytotoxic agents that provoke DNA damage as well as in monotherapy. In addition, PARP i's are being investigating in combination with other targeted agents such as PI3-kinase or angiogenesis inhibitors: the VEGF monoclonal antibody bevacizumab, has been shown to induce hypoxia in the tumour microenvironment which might contribute to genomic instability and increase sensitivity to PARPi's. Platinum sensitivity [6] is a clinical feature that identifies responder to trabectedin too: this agent might represent a valuable alternative option in patients in whom platinum agents are contraindicated. The activity of trabedectin is related to altered function and expression of DNA repair genes, such as BRCA1-2.
Role of the healthy cells in tumours formation and progression
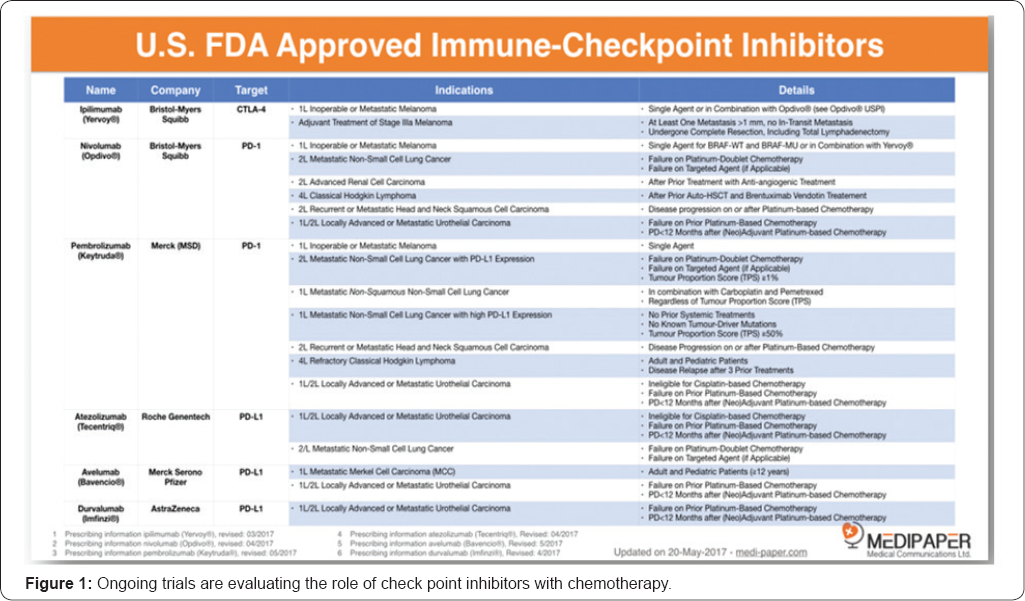
As members of scientific community, it's once again required not to be heavily influenced by the past experience and to look beyond; one of the new questions is what role the "healthy" cells play in cancer progression. Immune System certainly plays a fundamental role: with the molecular identification of human tumour antigens in the early 1990's, the opportunity to sensitize immune cells against tumour antigens is another way to be explored: [7] for instance, vaccination is a possible adjuvant treatment post-surgery, when tumour-induced immune suppression is minimal; we have examples in pancreatic and breast cancer. Next steps will include the production of targeted vaccines, patient-specific, to be used in combination with other treatments. Immune- check point inhibitors have shown promising [8] activity in several solid tumours and have demonstrated a favourable toxicity profile. Data from early clinical trials, utilizing PD1 and PD-L1 inhibitors, showed encouraging results. Ongoing trials are evaluating the role of check point inhibitors with chemotherapy (Figure 1).
Changing perspective
We need to change perspective and consider that in studying and treating cancer we should go beyond the histotypes, the clinical presentation and looking at the molecular targets, the genetic markers that are shared by apparently different diseases. This makes time gains in the applied pharmacological research and wide new solutions for incurable patients. It's the horizon of the future but seems to get closer and closer. [2] Accumulating evidence suggests that, for instance, PARP i's may have a wider application in other malignancies, where the genetic signature is a defective DNA damage repair pathway, such as pancreatic cancer, prostate cancer ad Ewing sarcoma. Several PARPi's are currently in phase I/II clinical investigation, as single agents and /or combination therapy in these solid tumours.
Challenges and change management of patients
Since there is currently no gold standard method of testing new classes of drugs response and no universal guidelines are in place on how to incorporate biomarker testing into routine clinical diagnostics, we are required to challenge the status quo, both at the level of clinical research, new drugs approval, looking at precision medicine, that means changing our routine, both at diagnostic and clinical level, improving the use of diagnostic techniques and modulating therapeutic algorithms. This is not easy to achieve: it is a challenge, which must be accepted if knowledge about the unveiled genome is to be gained, seeing that technology now allows faster, more accurate sequencing, including single cell sequences, full length gene sequencing, more accurate specimen easy-to-obtain such as circulating tumour cells detection, both to diagnose the disease and to monitor therapy-resistant clones.
High accuracy and new standards are required in labs where the specimen is processed, acquiring the right number of cells to be used for genome analysis; avoiding DNA contamination (different specimen from different patients)by using disposable equipment (one microtome per specimen, clearing the cutting area); choosing the right chip /test based on the kind of request coming from the clinicians that's diagnosis or risk assessment (e.g.: hereditary breast /ovarian cancer).Changing management of patients is required, not only in clinical trials but also in surgery or hospital, where the clinicians should give the right weight to genetic markers, addressing the patients to the suitable upfront line and subsequent lines of therapy, communicating properly the need for a genetic test to their patients, without feeding false hopes regarding therapy or, based on genetic test results, choosing the right way for prophylaxis.
The next challenges come from increasing knowledge on mechanisms of action of the single new drug, including the assessment of the principal and secondary molecular targets, defining the basic of pharmacology, i.e. pK, PD, CYP variants' interactions, dose-range, chemoresistance, possible associations with more than one class of drugs (i.e.: antiangiogenetics + PARPi's) and their efficacy in patients with advanced metastatic disease.
The unveiled genome today
I have a dream: that soon the genome of cancer cells brings us more and more rationally and consciously to the human being - centric rather than tumour - centric vision: for some types of cancer such as breast tumours, for instance, studies on family trees have saved human lives, the ones of mutations carriers who have not yet clinically developed the disease, increasing knowledge in Human Population Genetics. I make my own the words of C. Dennis [9] in the special issue of Nature dedicated to the human genome, stated: humans are much more than simply the product of a genome, but, in a sense, we are, both collectively and individually, defined within genome. The mapping, sequencing and analysis of the human genome is therefore a fundamental advance in self-knowledge, it will strike a personal chord with many people and application of this knowledge will, in time, materially benefit almost everyone in the world.
References
- Futreal PA (2001) Cancer and Genomics. Nature 409(6822): 850-852.
- O Sullivan C (2014) PARP inhibitors for BRCA mutation-associated and BRCA-like solid tumors. Cancer Molecular Targets and Therapeutics 4: p. 1-42.
- George A, Banerjee S, Kaye S (2017) Olaparib and somatic BRCA mutations. Oncotarget 8(27): 43598-43599.
- Lim D, Ngeow (2016) Evaluation of the methods to identify patients who may benefit from PARP inhibitor use. Endocrine Related Cancer 23(6): R267-R285.
- Konecny G, Kristeleit RS (2016) PARP inhibitors for BRCA1/2-mutated and sporadic ovarian cancer: current practice and future directions. British Journal of Cancer 115(10): 1157-1173.
- Lorusso D, Scambia G, Pignata S, Sorio R, Amandio G, et al. (2016) Prospective phase II trial of trabectedin in BRCA-mutated and/or BRCAness phenotype recurrent ovarian cancer patients: the MITO 15 trial. Ann Oncol 27(3): 487-93.
- Dodson LF, Hawkins WG, Goedeggeburre p (2011) Potential targets for pancreatic cancer immunotherapeutics. Immunotherapy 3(4): 517-37.
- Hardwick N, Frankel PH, Cristea M (2016) New Approaches for Immune Directed Treatment for Ovarian Cancer. Curr Treat Options Oncol 17(3): 14.
- Dennis C (2001) The Human Genome unveiled. Nature 409: 814-816.






























