Use of Systemic Antibiotics and Topical Nasal Treatments to Clear Mycoplasma ovipneumoniae from Lambs
Lauren E Christensen1, Janet E Williams1, Gavin G Cotterill2, Mark A McGuire1, Pedram Rezamand1, E Frances Cassirer2 and Denise E Konetchy1*
1Department of Animal, Veterinary and Food Sciences, University of Idaho, Moscow, USA
2U.S. Geological Survey, Northern Rocky Mountain Science Center, West Glacier, MT, USA
3Idaho Department of Fish and Game, Lewiston, USA
Submission: July 25, 2023; Published: August 07, 2023
*Corresponding author: Denise Konetchy, Department of Animal, Veterinary and Food Sciences, University of Idaho, 875 Perimeter Dr, Moscow, ID, 83844, USA. Email: konetchy@uidaho.edu
How to cite this article: Lauren E C, Janet E W, Gavin G C, Mark A M, Pedram R. Use of Systemic Antibiotics and Topical Nasal Treatments to Clear Mycoplasma ovipneumoniae from Lambs. Dairy and Vet Sci J. 2023; 15(5): 555924.DOI: 10.19080/JDVS.2023.15.555924
Abstract
Mycoplasma ovipneumoniae is commonly associated with chronic respiratory disease of domestic small ruminants worldwide. To assess the efficacy of systemic antibiotics with or without nasal flush treatments to clear M. ovipneumoniae in sheep. Suffolk lambs identified as positive for M. ovipneumoniae via nasal swab PCR were randomly assigned to one of eight different treatment groups and a positive control group (n=6 animals/group). Treatments groups included oxytetracycline (20 mg/kg SQ d1 or 10 mg/kg IM once daily for 5 d) with and without either a dilute betadine or chlorhexidine nasal flush SID for 5 d; or lincomycin (5 mg/kg IM on d 1, 3, and 5 with or without a dilute lincomycin nasal flush SID for 5d; or florfenicol (20 mg/kg IM on d 1, 3, and 5 with or without a dilute florfenicol nasal flush SID for 5d; and a positive control receiving no treatment. Least squares-means of PCR cycle threshold (Ct) values of each treatment were compared to Ct values of the control group over time to evaluate treatment effect. Oxytetracycline 10 mg/kg administered IM SID for 5 d was the only treatment to increase Ct values overall compared to the positive control (P = 0.003), indicating a decrease in M. ovipneumoniae DNA, however infection was not cured or maintained over time. Treatment with these bacteriostatic antibiotics did not clear M ovipneumoniae infection from lambs. The use of nasal flush therapies was unproductive and may have had a negative impact on the clearance of infection.
Keywords: antibiotics; Mycoplasma ovipneumoniae; sheep; oxytetracycline
Keywords: FARAD: Food Animal Residue Avoidance Databank; OXD: oxytetracycline injections
Introduction
Mycoplasma ovipneumoniae was described in 1974 in domestic sheep in New Zealand [1] and is commonly found in domestic sheep and goats worldwide. Often associated with a chronic cough, infected animals range from asymptomatic to severe polymicrobial pneumonia and death [2]. Infected lambs can have reduced efficiency in weight gain and carcass yield [3]. This organism can also cause severe disease in wildlife, including bighorn sheep (Ovis canadensis), mountain goats (Oreamnos americanus), and musk oxen (Ovibos moschatus) [4-8]. Characteristics of Mycoplasma spp. make it difficult to culture, contributing to limited antibiotic-based therapeutic options, and no vaccine is available to prevent infection [9]. M. ovipneumoniae is transmitted between animals in proximity via direct contact or aerosolized respiratory droplets.
Domestic sheep lambs (or naïve adult animals) are exposed to M. ovipneumoniae through contact with infected adults (many asymptomatic) in a flock that harbors the bacterium in their nasal passages [10]. Eighty-eight percent of 400 sheep operations and all range flocks in 22 states had at least one animal with a nasal swab positive for M. ovipneumoniae via PCR [11]. Multiple strains of M. ovipneumoniae as determined by multi-locus sequencing, have been identified in sheep flocks [7, 12] and within individual sheep [13], supporting the high prevalence of M. ovipnemoniae positive individuals and flocks in sheep operations.
Mycoplasma spp. are the smallest known bacteria, lack a cell wall and have a small genome (0.58-2.20 Mb) compared to other bacteria. Mycoplasma spp. are fastidious and difficult to culture from a diagnostic sample, as other bacteria present in the sample often outcompete mycoplasmas due to their slow growth rate and small colony size [14]. Serology can be used as a diagnostic aid or screening tool but is not definitive in diagnosis of active mycoplasma infections [15,16]. Domestic sheep have been found to have high amounts of M. ovipneumoniae in their nasal passages yet low antibody titers, suggesting an evolved tolerance of M. ovipneumoniae and the ability to shed the organism without incurring high rates of disease [17]. Polymerase chain reaction (PCR) is the current standard for laboratory diagnosis of Mycoplasma spp. in humans and animals. Compared to traditional bacterial culture, PCR is superior in sensitivity and specificity as well as efficiency [14]. We hypothesize that treating lambs with systemic antibiotics and a nasal flush will be effective in clearing M. ovipneumoniae from infected lambs. Based on this hypothesis, our objective is to identify an antibiotic therapy effective in clearing M. ovipneumoniae from lambs to develop recommendations for producers in reducing or eliminating infected animals in their flocks.
Materials and Methods
Animals, Treatments, and Experimental Design
All animals, treatments, and procedures were approved by the University of Idaho Animal Care and Use Committee (IACUC 2020- 63). The study took place December 2020 to February 2021 (n = 30 lambs, Cohort I) and December 2021 to February 2022 (n = 36 lambs, Cohort II).
Animals
A pool of sixty-six Suffolk ram and ewe lambs between 9-12 months of age were identified and screened for the presence of M. ovipneumoniae DNA via real time PCR (RT-PCR) using a deep nasal swab technique [5]. Animals were categorized based on the cycle threshold (Ct) results: ‘Detected’ (Ct ≤ 36), ‘Indeterminate’ (Ct = 36-40), or ‘Not Detected’ (Ct ≥ 40) [2]. In Cohort I, thirty animals (twenty ram and ten ewe lambs) were randomly selected from the pool of ‘Detected’ animals. Each treatment group in Cohort I included four ram lambs and two ewe lambs for a total of six animals per treatment. In Cohort II, thirty-six ewe lambs were randomly selected from a pool of available ‘Detected’ animals and randomly assigned to treatments with a total of six animals per treatment. In both Cohort I and Cohort II, animals were housed in groups of six, in corner pens separated from other groups by 2.9 meters. Each pen had a separate water source and feeder. Water was provided ad libitum and were fed a total mixed ration consisting of alfalfa, grass hay, corn, and barley to meet nutrient requirements for growth. Animals underwent a 1-week acclimation period prior to the start of the study and were returned to the University of Idaho sheep center following the conclusion of the study.
Treatment

Treatments started on a Monday (d 1 of treatment) and, if continued, were administered at 24- or 48-h intervals through Friday (d 5 of treatment) to conclude the treatment timeframe. Sampling occurred starting 6 d after the conclusion of the treatment timeframe (d 6 post-treatment). Treatments in Cohort I were as follows: oxytetracycline (Liquamycin LA-200, Zoetis Animal Health, Parsippany, NJ) 20 mg/kg subcutaneously (SQ) once on d 1 (OXO); oxytetracycline 10 mg/kg intramuscularly (IM) once daily for 5 d (OXD); oxytetracycline 10 mg/kg IM daily for 5 d with a dilute betadine (2% mixed with 0.9% sodium chloride to a 100ml volume) nasal flush daily for 5 d (OXB); oxytetracycline 10 mg/kg IM daily for 5 d with a dilute chlorhexidine (2% mixed with 0.9% sodium chloride to a 100 mL volume) nasal flush daily for 5 d (OXC); and positive control receiving no treatment (POS). In Cohort II, treatments were as follows: lincomycin (Lincomix 300, Zoetis Animal Health, Parsippany, NJ) 5 mg/kg IM on d 1, 3, and 5 (LIN); lincomycin 5 mg/kg IM on d 1, 3, and 5 with dilute lincomycin (lincomycin 0.5 mg/kg mixed with 0.9% sodium chloride to a volume of 100 mL) nasal flush daily for 5 d (LIF); florfenicol (Nuflor, Merck Animal Health, Madison, NJ) 20 mg/ kg IM on d 1, 3, and 5 (FLO); florfenicol 20 mg/kg IM on d 1, 3, and 5 with dilute florfenicol (florfenicol 2 mg/kg mixed with 0.9% sodium chloride to a volume of 100mL) nasal flush daily for 5 d (FLF); oxytetracycline 10 mg/kg IM daily for 5 d (OXD); and positive control receiving no treatment (POS). Antibiotic and nasal flush dosages were calculated based on individual animal weights obtained on the first day of the acclimation period. Nasal flushes were administered with a large volume low pressure human sinus irrigation bottle. Drug withdrawal for each antibiotic was referenced from the Food Animal Residue Avoidance Databank (FARAD) (Table 1).
Sample Collection
On d 7, 14, 21, and 28 post-treatments, all animals were nasal swabbed twice using sterile, polyurethane sponge-tipped swabs (Becton Dickson Co, Franklin Lakes, NJ) on a plastic shaft secured to a cap with a plastic transit tube. Swabs were labeled, kept in a cooler on ice and out of sunlight during transport, and stored within 1 h of collection at -20 °C until further analysis.
Nasal Swab Extraction
Swab tips were aseptically cut from the shaft and placed in a 1.5 mL microcentrifuge tube. Three hundred μL phosphate buffered saline was added and the sample was then pulse vortexed for 15 s to agitate the nasal mucus from the swab tip into suspension. The resulting suspension was then pipetted into a new microcentrifuge tube and DNA extracted using the QIAamp DNA Mini Kit (Qiagen, Germantown, MD) following manufacturer’s instructions. DNA was eluted in 100 μL of Buffer AE and either processed immediately in RT-PCR or stored at -20°C for later analysis.
Nasal Swab RT-PCR Analysis
Following DNA extraction of the nasal swabs, RT-PCR was conducted using the Applied Biosystems 7500 Fast Real Time PCR System using previously described methods [9] with primers 226Fnew (5′-GGGGTGCGCAACATTAGTTAGTTGGTAG-3′) and LMR1 (5′-GACTTCATCCTGCACTCTGT-3′), and probe Movip 253P (5′6-FAM-TTAGCGGGGCCAAGAGGCTGTABHQ-1-3’) [2]. Bovine serum albumin was added to the PCR master mix to increase PCR amplification yields in the presence of PCR inhibitors such as dirt and organic material that might be present on the nasal swabs [18].
Data Analysis
Individual Ct values were used in a mixed modeling framework with a random effect of individual animal to account for nonindependence of repeated measures, as well as a fixed effect of sex and a quadratic interaction effect of sample day. All statistical analyses were performed in R [19] using the lme4 package [20]. The emmeans package [21] was used to compare least squares means of Ct values between treatment and positive control groups overall and at each sample timepoint. Data were visually assessed for normal distribution. Significance was determined at P < 0.05.
Results
Detection of M. ovipneumoniae via PCR
Response to treatment was evaluated by comparing least squares means of PCR Ct values for each treatment to the least squares means of the control group Ct values over time. Least squares mean Ct values for POS were 25.3 (±1.21) overall and trended to decrease over time (d 7=29.1±1.34, d 14=26.0±1.16, d 21=25.2±1.16, and d 28=26.8±1.34; Figure 1). Least squaresmeans Ct values were higher in the OXD group when comparing POS and OXD overall (P = 0.0027) (Table 2) indicating a lower amount of M. ovipneumoniae DNA in the OXD group compared to the POS group. Least squares-means Ct values of OXD compared to that of POS were different on d 7 (P = 0.0039), d 14 (P = 0.0005), and d 21 (P = 0.0077), but not at d 28 (P = 0.5). Least squaresmeans Ct values for OXD were 31.4±1.21 overall and decreased over time, indicating an increase in M. ovipneumoniae genomic material (d 7=35.7±1.34, P = 0.0042, d 14=32.6±1.16, P = 0.0007, d 21= 30.6±1.16, P = 0.0096, and d 28= 29.6 ±1.34, P = 0.5148) (Figure 2).
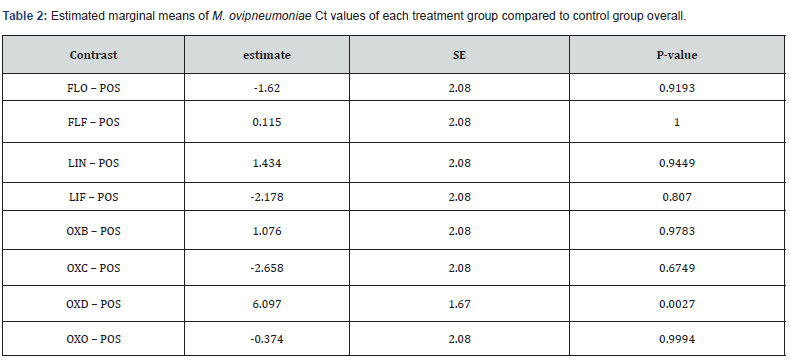
POS = positive control; FLO = Florfenicol injectable only; FLF = Florfenicol injectable with florfenicol nasal flush; LIN = Lincomycin injectable only; LIF = Lincomycin injectable with lincomycin nasal flush; OXB = Oxytetracycline injectable with betadine nasal flush; OXC = Oxytetracycline injectable with Chlorhexidine nasal flush; OXD = Oxytetracycline injectable daily for 5 days; OXO = Oxytetracycline single injection.

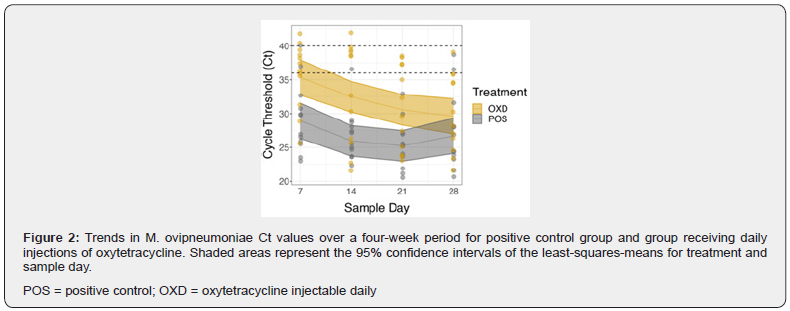
A difference in least squares means Ct values between POS and OXB was detected only on d 7 (P = 0.0339) where the least squares-means Ct values were higher in the OXB group (35.8±1.89) compared to the POS group (29.2±1.34; Figure 1), indicating a lower amount of M. ovipneumoniae DNA in the OXB group. There was no difference in mean Ct values between POS and OXO, OXB, OXC, LIN, LIF, FLO, or FLF overall nor at any time-point following treatment (Figure 1).
Discussion
Over the course of the study, all treatment cohorts were observed (Figure 1) to have Ct values trending lower over time, indicating an increase in M. ovipneumoniae DNA, presumably because of increased growth of M. ovipneumoniae in these animals over time. The presence, and increase, of bacteria could be related to re-infection from pen-mates which did not respond to treatment; re-colonization of an individual’s nasal passages with M. ovipneumoniae which remained present in the cornual sinus or lower airway [5], or incomplete response to treatment due to antibiotic susceptibility differences in strain type [22]. M. ovipneumoniae from the sheep in our study were not strain typed, though previous studies suggest the strong likelihood of multiple strains within individual study animals and variations related to strain type influencing short- and long-term treatment outcomes [12,13]. In a study by Kamath and others, 187 genetically different strains of M. ovipneumoniae were identified in 207 domestic sheep, with 77% of individual sheep having unique strains [7]. In bighorn sheep and domestic sheep, immunity to M. ovipneumoniae appears to be strain specific [9,23]. Oxytetracycline, a bacteriostatic antibiotic, administered daily appeared to produce a decline in M. ovipneumoniae DNA but failed to eliminate the bacteria DNA. Animals receiving oxytetracycline showed an increase in bacteria DNA presence over time once the bacteriostatic agent was no longer present systemically [24, 25].
All antibiotics evaluated (oxytetracycline, florfenicol, and lincomycin) are bacteriostatic antibiotics whose mechanism of action stalls bacterial cellular activity without directly causing bacterial death. Efficacy of bacteriostatic agents depends on the ability of the antimicrobial agent to suppress the bacteria long enough for the host immune system to respond to the pathogen, and for the host to recognize and respond to the presence of a pathogen [26]. Daily administration of oxytetracycline injections (OXD) resulted in higher Ct values (lower M. ovipneumoniae DNA concentrations) when compared to that for the POS group. A singledose therapy of oxytetracycline (OXO) did not achieve therapeutic levels of the antibiotic for a length of time to allow the host’s immune system to mount an effective response to the presence of the bacteria. Daily flush with either betadine or chlorhexidine paired with daily oxytetracycline injections should work synergistically and be the most efficacious in disrupting any M. ovipneumoniae by physically removing bacteria from the animal’s nasal passages. Betadine and chlorhexidine are bactericidal and bacteriostatic, respectively, with chlorhexidine being bactericidal at high concentrations but diminishing in efficacy in the presence of organic matter [27] such as nasal mucus. Lincomycin and florfenicol, labeled for use against infections due to Mycoplama spp. in swine and cattle, respectively, were each used as a nasal flush and anticipated to improve clearance of M. ovipneumoniae when used in combination with systemic lincomycin (LIF) and florfenicol (FLF) administration. Both, however, were unable to decrease bacterial presence through a local nasal flush application (Figure 1). In fact, LIF and FLF were among the least efficacious treatments, performing close to the control groups receiving no treatments. Increased serous nasal discharge was observed in the study animals receiving nasal flush treatments, regardless of flush material used. Nasal discharge could have diluted the nasal flush treatment and caused the treatment substance to be removed faster than the treatment could act locally in the nasal passages to remove or reduce bacterial load [28]. Both lincomycin and florfenicol have a basic pH of 4 and 5, respectively [29, 30], which would cause dehydration of nasal cilia, disrupt the ciliary beat, and reduce mucociliary clearance [31].
There are several limitations to studying. Differing nasal swab sampling techniques among individuals could result in variation in Ct values. Four individuals collected the samples during the study. However, all were trained to use a similar technique and depth of swabbing. There is also potential for variation between M. ovipneumoniae organism load between swabs when multiple swabs are taken from an individual animal at the same time.
Conclusion
The objective of this study was to find an antibiotic therapy effective in clearing M. ovipneumoniae from lambs, which could then be recommended to producers to use to treat M. ovipneumoniae positive animals in their flock. To be considered effective, a treatment is needed to clear M. ovipneumoniae from the animal and provide reasonable duration of effect. Although animals receiving daily oxytetracycline injections for 5 days had nasal swabs with increased Ct values, (reduced M. ovipneumoniae DNA), when compared to that for the control, it did not induce a complete cure over time in most animals. Our study findings indicate treatment with bacteriostatic antibiotics did not clear M. ovipneumoniae infection from most lambs and the use of nasal flush therapies did not aid in antibiotic efficacy and may have had a negative impact on clearance of infection in lambs. Further studies investigating strain-typing or utilizing segregation of test negative animals after treatment to prevent re-infection from a positive cohort might improve outcomes for clearing M. ovipneumoniae from lambs.
References
- Clarke JK, Alley MR (1974) Isolation and identification of mycoplasmas from the respiratory tract of sheep in New Zealand. New Zealand Veterinary Journal 22(7):117-121.
- Manlove K, Branan M, Baker K, Bradway D, Cassirer EF, et al. (2019) Risk factors and productivity losses associated with Mycoplasma ovipneumoniae infection in United States domestic sheep operations. Preventive Veterinary Medicine 168:30-38.
- Besser TE, Levy J, Ackerman M, Nelson D, Manlove K, et al. (2019) A pilot study of the effects of Mycoplasma ovipneumoniae exposure on domestic lamb growth and performance. PLoS ONE 14(2):1-9.
- Besser TE, Cassirer EF, Potter KA, Lahmers K, Oaks JL, et al. (2014) Epizootic pneumonia of bighorn sheep following experimental exposure to Mycoplasma ovipneumoniae. PLoS ONE 9(10).
- Besser TE, Cassirer EF, Potter KA, Foreyt WJ (2017) Exposure of bighorn sheep to domestic goats colonized with Mycoplasma ovipneumoniae induces sub-lethal pneumonia. PLoS ONE 12(6).
- Handeland K, Tengs T, Kokotovic B, Vikoren T, Ayling RD, et al. (2014) Mycoplasma ovipneumoniae - A primary cause of severe pneumonia epizootics in the Norwegian muskox (Ovibos moschatus) population. PLoS ONE 9(9).
- Kamath PL, Manlove K, Cassirer EF, Cross PC, Besser TE (2019) Genetic structure of Mycoplasma ovipneumoniae informs pathogen spillover dynamics between domestic and wild Caprinae in the western United States. Scientific Reports 9(1).
- Wolff PL, Blanchong JA, Nelson DD, Plummer PJ, McAdoo C, et al. (2019) Detection of Mycoplasma ovipneumoniae in pneumonic mountain goat (Oreamnos americanus) kids. Journal of Wildlife Diseases 55(1):206-212.
- Ziegler JC, Lahmers KK, Barrington GM, Parish SM, Kilzer K, et al. (2014) Safety and immunogenicity of a Mycoplasma ovipneumoniae bacterin for domestic sheep (Ovis aries). PLoS ONE 9(4).
- Brogden KA, Rose D, Cutlip RC, Lehmkuhl HD, Tully JG (1988) Isolation and identification of mycoplasmas from the nasal cavity of sheep. American Journal of Veterinary Research 49(10):1669-1672.
- USDA (2014) Sheep 2011 Part IV: Changes in Health and Production Practices in the U.S. Sheep Industry 1996-2011.
- Harvey ME, Morrical DG, Rosenbusch RF (2007) Sheep flock infections with Mycoplasma ovipneumoniae involve multiple strains. Small Ruminant Research 73(1-3): 287-290.
- Lonas G, Clarke JK, Marshall RB (1991) The isolation of multiple strains of Mycoplasma ovipneumoniae from individual pneumonic sheep lungs. Veterinary Microbiology 29(3-4): 349-360.
- Parker AM, Sheehy PA, Hazelton MS, Bosward KL, House JK (2018) A review of mycoplasma diagnostics in cattle. Journal of Veterinary Internal Medicine 32(3): 1241-1252.
- Maes D, Sibila M, Kuhnert P, Segalés J, Haesebrouck F, et al. (2018) Update on Mycoplasma hyopneumoniae infections in pigs: Knowledge gaps for improved disease control. Transboundary and Emerging Diseases 65(S-1):110-124.
- Maunsell FP, Woolums AR, Francoz D, Rosenbusch RF, Step DL, et al. (2011) Mycoplasma bovis infections in cattle. Journal of Veterinary Internal Medicine 25(4):772-783.
- Cassirer EF, Manlove KR, Almberg ES, Kamath PL, Cox M, et al. (2018) Pneumonia in bighorn sheep: Risk and resilience. Journal of Wildlife Management 82(1): 32-45.
- Lekang K, Thompson EM, Troedsson C (2015) A comparison of DNA extraction methods for biodiversity studies of eukaryotes in marine sediments. Aquatic Microbial Ecology 75(1).
- R Core Team (2020) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria.
- Bates D, Maechler M, Bolker B, Walker S, (2015) Fitting Linear Mixed-Effects Models Using lme4. Journal of Statistical Software 67(1):1-48.
- Lenth RV (2022) emmeans: Estimated Marginal Means, aka Least-Squares Means.
- Maksimović Z, Bačić A, Rifatbegović M (2020) Antimicrobial susceptibility of caprine and ovine Mycoplasma ovipneumoniae Microbial Drug Resistance 26(10): 1271-1274.
- Cassirer EF, Manlove KR, Plowright RK, Besser TE (2017) Evidence for strain-specific immunity to pneumonia in bighorn sheep. Journal of Wildlife Management 81(1): 133-143.
- Aktas İ, Yarsan E (2017) Pharmacokinetics of conventional and long acting oxytetracycline preparations in kilis goat. Frontiers in Veterinary Science 4(229).
- Craigmill AL (2003) A physiologically based pharmacokinetic model for oxytetracycline residues in sheep. Journal of Veterinary Pharmacology and Therapeutics 26(1): 55-63.
- Murray P, Rosenthal K, Pfaller M (2020) Medical Microbiology (9th) Elsevier.
- Mcdonnell G, Russell AD (1999) Antiseptics and disinfectants: Activity, action, and resistance. In Clinical Microbiology Reviews 12(1):147-179.
- Gizurarson S (2015) The effect of cilia and the mucociliary clearance on successful drug delivery. Biological and Pharmaceutical Bulletin 38(4):497-506.
- Chen T, Zhu Z, Zhang H, Shen X, Qiu Y, et al. (2019) Enhanced removal of veterinary antibiotic florfenicol by a Cu-based fenton-like catalyst with wide pH adaptability and high efficiency. ACS Omega 4(1): 1982-1994.
- Czarniak P, Boddy M, Sunderland B, Hughes JD (2016) Stability studies of lincomycin hydrochloride in aqueous solution and intravenous infusion fluids. Drug Design Development and Therapy 10: 1029-1034.
- Bustamante-Marin XM, Ostrowski LE (2017) Cilia and mucociliary clearance. Cold Spring Harbor Perspectives in Biology 9(4).






























