ACE2 and Covid-19 Spike Protein Docking in Family Pets
Atlas Mashayekhi Sardoo1, Alyssa M Torres2, Claude Krummenacher1,2 and Yong Chen1*/h3>
1Department of Molecular and Cellular Biosciences, Rowan University, Glassboro NJ 08028, USA
2Department of Biological Sciences, Rowan University, Glassboro NJ 08028,
Submission:December 13, 2021; Published:January 18, 2022
*Corresponding author: Yong Chen, Department of Molecular and Cellular Biosciences, Rowan University, Glassboro NJ 08028, USA
How to cite this article: Sardoo AM, Torres AM, Krummenacher C, Chen Y. ACE2 and Covid-19 Spike Protein Docking in Family Pets. Dairy and Vet Sci J. 2022; 15(1): 555905.DOI: 10.19080/JDVS.2022.15.555905
Abstract
COVID-19 infection is caused by Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). COVID-19 infection has resulted in hundreds of thousands of human deaths since the announcement of the Coronavirus pandemic in 2020. Emerging evidence supports the fact that SARS-CoV-2 infection happened not only in humans but in animals as well. Given the high frequency f of pets in family homes in many countries, COVID-19 infection in pets poses a potential threat to other domestic and wild animals, and humans. In this paper, we review the infection cases in pets world-wide and investigate the possible infection chain by an in-silico docking analysis of the SARS-CoV-2 spike protein with ACE2 proteins amongst 27 species including many pet animals. High docking scores indicate high interacting affinities between the SARS-CoV-2 spike protein and the cellular ACE2 receptor from these pet species. The phylogenetic analysis of human and pet ACE2 receptors show high sequence identities, especially for several functional sequence regions. These results suggest that the SARS-Cov-2 may have similar ability to penetrate cells from these different species and use similar molecular interactions to initiate infection of pet and human cells. Considering the high occupancy rates of pets in American and world-wide, the results suggest that pets should be considered in strategies to control COVID-19.
Keywords:SARS-CoV-2; Family pets; Human ACE2 receptor; Protein–protein docking
Abbreviations:WHO: World Health Organization; COVID-19: Coronavirus Disease 2019; SARS-CoV-2: Severe Acute Respiratory Syndrome Coronavirus 2; TEMPRSS2: Transmembrane Serine Protease 2; ACE2: Angiotensin I Converting Enzyme 2; RT-PCR: Reverse Transcription-Polymerase Chain Reaction; NCBI: National Center for Biotechnology Information
Introduction
Early in 2020, the World Health Organization (WHO) announced Coronavirus Disease (COVID-19) a pandemic [1]. This infection is caused by a pathogen named Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). This existing pandemic has currently been reported in 215 countries and territories with about 250 million proven cases and more than 5 million deaths globally [2]. It is now proven that SARS-CoV-2 is able to infect not only in humans but also in pets as well as other domestic and wild animals [3]. Various published scientific evidence revealed a strong homology between SARS-CoV-2 and bat coronaviruses, suggesting that the COVID19 pandemic is a zoonotic pandemic caused by a virus originating in bats. Additionally, pangolins have been counted as one of the possible intermediate hosts (Figure 1) [4,5].
Coronaviruses are one type of protein enveloped RNA viruses, which measure 80–220 nm in diameter. Their genome is the largest among RNA viruses and comprises a non-segmented,
single-stranded positive-sense genome of approximately 26–32 kilobases (kb) in size [6]. Structurally, the viral particle comprises four main proteins; 1) spike (S) glycoprotein, 2) small envelope (E) glycoprotein, 3) membrane (M) glycoprotein, and 4) nucleocapsid (N) protein (Figure 1) [7]. The M protein is the most abundant structural protein, specifies the shape of the viral particle and plays a major role in viral assembly [8]. The N protein is an RNA binding protein involved in the binding and packaging of the viral RNA genome into a long helical nucleocapsid structure. The S glycoprotein facilitates viral binding to a specific receptor on the animal and human host cells, and promotes fusion between the viral envelope and a cellular membrane [9].
The attachment of SARS-CoV-2 S glycoprotein to the ACE2 receptor starts the process of entry into target cells [10]. The SARS-CoV-2 can enter cells using either endosomal or non-endosomal pathways [11]. In the endosomal pathway, the binding of the S glycoprotein to ACE2 triggers receptor-mediated endocytosis which is followed by fusion between the viral envelope and an endosomal membrane to release the viral genome in the cytosol.
In the non-endosomal pathway, cleavage of the viral S glycoprotein
by transmembrane serine protease 2 (TMPRSS2) at the surface of
the host cell allows the S glycoprotein to mediate fusion of the
viral envelope directly with the cell plasma membrane. Both entry
pathways lead to the release and uncoating of the viral genome in
the cytosol of the host cell allowing it to be subsequently translated
and replicated [12].

The human ACE2 receptor, also termed as ACEH (ACE
homologue), is a dimeric, zinc-dependent metalloproteinase of the
ACE family that comprises germinal and somatic ACE [13,14]. ACE2
mRNA is highly detected in testes, heart and kidneys, however, it
is found at lower levels in an extensive variety of tissues [13,15].
As part of its natural function, ACE2 acts as a carboxypeptidase
which cleaves various substrates including angiotensin I and
II [10]. It is also involved in the function of the B0AT1-family of
amino acid transporters [13]. Due to these physiological functions,
ACE2 has been linked to various diseases including, heart disease
[16] as well as liver and lung fibrosis [17], although it has been
shown to have a protective effect in acute lung injury [18]. ACE 2
is a major factor in COVID-19, due to its key role as a receptor for
SARS-CoV-2.
Structurally, the ACE2 protein comprises three main regions:
1) the N-terminal part of the extracellular region that contains
the catalytic domain of the peptidase; 2) the C-terminal part of
the extracellular region contains a collectrin-like domain (CLD)
which extends through the transmembrane region; and 3) a
short cytoplasmic tail [19]. The N-terminal region is essential
for attachment to the SARS-CoV and SARS-CoV-2 spike proteins,
while the CLD contains a region that allows dimerization and
interaction with amino acid transporters [14]. The peptidase
domain comprises a long deep cleft that undergoes a large hingebending
movement at inhibitor and substrate attachment [19].
Pets as targets of SARS-CoV-2 infection
The first COVID-19 case in a domestic animal was reported in
a Pomeranian dog from China in February 2020 [20]. Afterwards,
in March 2020, COVID-19 was observed in a cat in the same
country [21]. These cases were detected after their owners were
positively diagnosed for COVID-19 [20,21]. The persistent positive
reverse transcription-polymerase chain reaction (RT-PCR) test
results of the Pomeranian dog was considered as a true positive
for an infection of the animal [22]. This conclusion was further
supported by the absence of contamination when the dog was
kept in quarantine at government kennels [22]. Genetic sequence
similarities of the SARS-CoV-2 genomes isolated from the pet
and the owners suggested the possibility of human-to-animal
transmission [23]. Additionally, both serological tests and viral
culture were performed to evaluate if the dog was contagious or
not. The negative results of these tests and the absence of any
symptom, led to the conclusion that the dog was not contagious to
humans and another animal [22].
After the first reported case of COVID-19 in a cat in China
[21], other COVID-19 cases were reported from Belgium [24],France [25], Germany [26], Russia [27], and the United States
[28]. According to these reports, it is currently accepted that cats
and dogs are two companion animals susceptible to SARS-CoV-2.
Cats are more susceptible than dogs and may have the potential
to transmit the illness to other naive cats [29]. In addition, in
laboratory experiments golden Syrian hamsters have been shown
to be susceptible to SARS-CoV-2 and effectively transmit infection
to naïve hamsters via aerosols or direct contact [30].
According to Dodds et al. [31] animals that are susceptible to
SARS-CoV-2 under experimental conditions, include cats, dogs,
and ferrets, but not horses, pigs, or poultry [31]. The results of
experimental studies in cattle are conflicting, however it does not
appear that cattle can easily be infected [31]. Additionally, another
study of caged cats demonstrates that transmission between cats
is possible with prolonged close contact. It does claim how likely
this is to occur in natural settings [32]. Under natural conditions,
dogs and cats are not considered easily infected, and infected
cats or dogs are not easily to spread the virus to other animals
or humans. However, according to the Ohio State University
College of Veterinary Medicine, dogs appear to be more resistant
than cats, and ferrets in a laboratory setting [33]. In summary,
pets living in the household of COVID-19 diagnosed people are in
danger of contracting the disease and can spread the virus to the
nearby naive pets. Therefore, owners need to strictly protect their
companion animals from getting infected. Understanding how
COVID-19 affects companion animals is essential given that 84.9
millions of U.S households’ own pets (Figure 2A) [34]. Notably,
as of July 2021, the USDA reports that the highest number of
positively tested animals in the USA are in Texas (Figure 2B) [35].
Although, the numbers of infected pets remain low overall, these
pets may have the potential to transmit the virus and potentially
generate variants which may adapt to better use animal ACE2
receptors and spread through new species [30].

High sequence similarity among human and pets ACE2
proteins
To understand the properties that allow SARS-CoV-2 infection
of pets, we considered primarily the interaction of SARS CoV-2 S
glycoprotein with the cellular receptor ACE2. Although species
specificity of viral infections is defined by numerous factors that
may affect multiple stages of the replication cycle of a virus,
effective interaction with a receptor that allows entry is an
essential, albeit not sufficient, step to ensure infection. Thus, we
modelled and analysed the interaction of the S glycoprotein with
ACE2 proteins from different species by phylogenetic analysis and
protein docking method [36]. We first compared the amino acid
sequences of ACE2 proteins from human and 27 animal species,
including major household animals (Table 1). The sequences were
retrieved from the National Center for Biotechnology Information
(NCBI) protein sequence database and the Sequence Alignment
was performed using the online tool “Clustal Omega” (https://
www.ebi.ac.uk/Tools/msa/clustalo/) [37]. Noticeably, pair-wise
alignments showed that all 27 ACE2 receptor sequences displayed
high similarities with human ACE2 protein, ranging from 99%
(chimpanzee) to 66.38% (white-ruffed manakin). We observed
that 5 species, chimpanzee, Sumatran orangutan, Macaca
nemestrina, sooty mangabey and rat, have an overall amino-acid
identity larger than 90%, suggesting that ACE2 is highly conserved
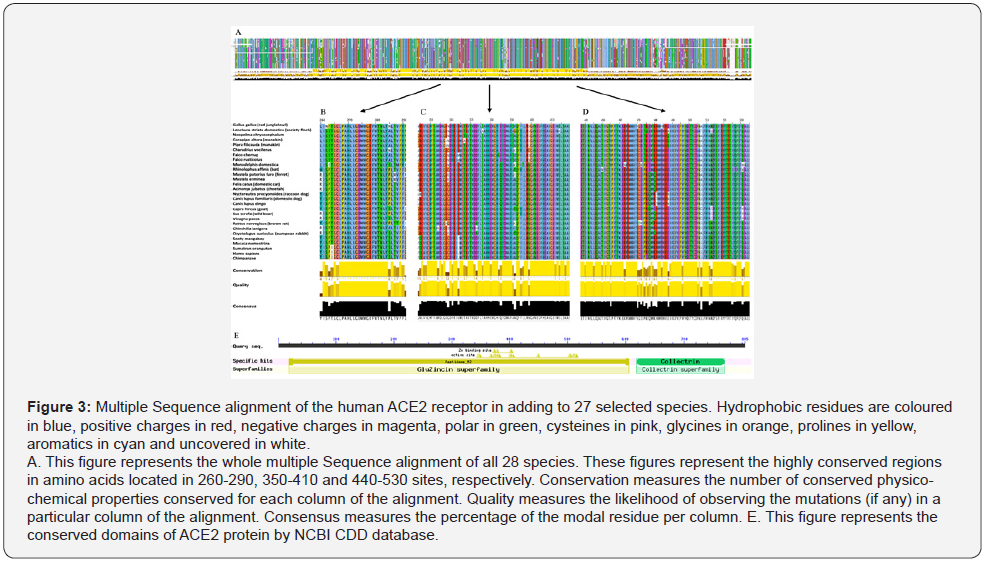
in structure and function between these species. We conducted
multiple sequence alignment for the 28 species sequences by using
the ClustalX method [38]. Results show that all these sequences
have either high amino acid sequence similarity or high physical/
structural features (Figure 3). As shown in figure 3B, 3C and 3D,
these regions are highly conserved [39]. In particular, the 362-366
region has been annotated to be important for natural substrate
binding and for interaction with SARS-CoV spike glycoprotein
(Protein Data Bank, https://www.rcsb.org), (Figure 3C).
Additionally, phylogenetic and molecular evolutionary
analyses were conducted using neighbour-joining method
by MEGA software [40]. A rooted phylogenetic tree was then
created by using the amino acid sequences to demonstrate the
evolutionary relationships between the 28 species (Figure 4). The
branching pattern of the phylogenetic tree reflects evolutionary
relationships. The phylogenetic analysis (Figure 4) shows that the
human ACE2 protein is most closely related to the orangutan and
chimpanzee proteins, but much less related to ACE2 from the 8
bird species analysed, which is consistent with general taxonomic
analysis. Overall, these sequence-based analyses demonstrate a
high genetic and structural conservation of ACE2 proteins among
the major pet species and humans. This suggests that ACE2 from
numerous species may be functional receptors for SARS-CoV-2
S glycoprotein and allow entry of the virus into cells from these
species.
High docking affinity of SARS‐CoV‐2 spike protein
with pet ACE2 protein
The interaction between ACE2 and SARS‐CoV‐2 S glycoprotein
is of utmost importance for SARS CoV-2 infection and spread in
humans [41]. To determine the potential binding affinity of the S
glycoprotein for ACE2 proteins from pet species, we performed a
protein–protein docking analysis using the HDOCK server [36].
This approach generates docking scores to represent the binding
free energy of two interacting proteins. Smaller negative docking
scores suggest higher docking affinity strength. For each of the
28 ACE2 proteins, the top ten models and their docking scores
were outputted and then the smallest negative docking scores
were selected (Figure 5). In surprise, we found there are 7 species
(Chimpanzee, Canis Lupus, Canis Lupus Dingo, Nyctereutes
Procyonoides, Oryctolagus Cuniculus, Sus Scrofa and Mustela
Putorius Furo) whose docking scores are smaller than human’s score. Among them, Chimpanzee achieves a smallest docking score
of -665.81 kcal/mol, indicating it may be the most suspectable
to Covid-19 infection. Meanwhile, humans and cats (Felis Catus)
have similar docking scores (-527.72 kcal/mol vs -523.15 kcal/
mol), which is in agreement with the published data suggests cats
can spread the infection among animals and humans [31]. Shen
et al. [42] also assessed the receptor utilization capacity of ACE2
from different species and noticed that cats were most susceptible
to infection by SARS-CoV-2 [42].


Pets as players in COVID-19 transmission
The protein-protein docking data and phylogenetic analysis of
ACE2 indicate that certain animal species express cellular ACE2
receptors that are highly similar to the human protein and are
likely to be used by SARS-CoV-2 with similar efficiency to penetrate
cells. Despite this similarity, infections of animals by SARS-CoV-2
remain very limited in number and severity, due to other speciesspecific
restriction to infection. Nevertheless, the data suggest
that if the essential step that initiate infection of e cell is conserved
between humans and several animal species, the virus may target
such animals and possibly evolve into variants more adapted to
infect those new species. This is relevant since 53% of US families
own dogs, 35.7% own cats, and 4.8% own birds (Figure 2B) [34].
Quarantine has affected many pets throughout the pandemic,
similarly to humans, pets can be stressed by change to their daily
lives. The Ohio State College of Veterinary Medicine has also
pointed out that there is no reason to harm wildlife or abandon
family pets out of fear [33]. Many families have actually opted to
adopt a pet during quarantine, increasing the percentage of U.S.
families who own pets. The risk of COVID-19 infection in pets can
be contained by treating pets as if they were a human member of
the family, keeping them away from people and animals who are
ill [33].
The phylogenetic analysis (Figure 4) shows a high degree of
conservation between orangutan and human ACE2 proteins. This
suggests that the SARS-CoV-2 S glycoprotein interacts with both
receptors in a very similar fashion and that the same region of the
S glycoprotein is involved in the binding to both receptors. This
supports the idea that a SARS-CoV-2 vaccine would be effective
to protect orangutans and human equally well, despite minor
differences in ACE2 proteins. The phylogenetic comparisons are
supportive of the fact that pets can also be infected by and spread
SARS-CoV-2. Indeed, infection has been confirmed in various
species, including 115 cats, 81 dogs, 27 big cats, 3 gorillas, 1
domestic ferret and 1 mink [7]. Our phylogenetic and sequence
alignment analysis (Figures 3 and 4) further demonstrates that
ACE2 proteins have highly conserved functional domains which
may be responsible for the docking process of the S-glycoprotein.
Vaccination of zoo animals has been undertaken in various zoos.
For instance, four orangutans at the San Diego Zoo received
an experimental COVID-19 vaccine, receiving two doses each
[43]. The vaccines developed to protect animals uses the same
approaches as human COVID-19 vaccines. As of July 2021, Zoetis
has donated approximately 11,000 doses of the experimental
veterinary use COVID-19 vaccine with hopes of protecting the
animals residing in nearly 70 zoos [44]. According to the Oakland
Zoo in Northern California, their veterinary team has started
early to vaccinate the animals deemed most at risk by their team,
including tigers, black bears, grizzly bears, mountain lions, ferrets
and chimpanzees [44]. The vaccines used in these studies were
safe and efficient in eliciting an immune response as judged by the
generation of serum neutralizing antibodies in-vitro.
Conclusion
The main purpose of this review is to contribute to our
understanding of how COVID-19 affects various family pets. This
is a relevant issue since humans have frequently maintained
close contacts with household pets and other animal species
during the current pandemic. To understand how pets and other animals may become infected by SARS-CoV-2, we investigated the
binding of the viral S glycoproteins to the cellular ACE2 receptor,
since this interaction is necessary to initiate infection of host
cells. In particular, we performed a protein-protein molecular
docking analysis of SARS-CoV-2 S glycoprotein with the ACE2
receptor from different species to better understand the potential
of those species to serve as intermediate hosts or simply to be
susceptible to SARS-CoV-2 infection. Our modeling data on ACE2
and S-glycoprotein docking provides a molecular and structural
basis for comparing how the virus infects animals to humans. This
approach provides an important tool to understand fundamental
aspects of SARS-CoV-2 infection that can translate to better
prevention and treatment and advance the fields of veterinary
medicine and public health. Functional studies have emerged to
address testing animals and reporting cases, case management for
minimizing exposure and handling recommendations for equine
species, food animals and other farm animals [45]. People are
primarily contracting COVID-19 from close contact with infected
individuals. Based on limited data available, the risk of animals
contributing to the spread of COVID-19 to humans remains very
low [45]. We learn more about the biology of SARS-CoV-2 every
day, but obviously more research must be conducted assess
how animals contribute to and suffer from the spread of this
virus. In addition, structural and functional analyses of SARSCoV-
2 infection, epidemiological studies will provide remarkable
information about outbreaks in humans and animals. For instance,
compartmental models have been applied in several countries to
analyses the compound transmission pattern of the COVID-19
pandemic, using delay differential equations, ordinary differential
equations, fractional order Caputo derivative and stochastic
differential equations [46-51]. Such mathematical model with
pets added as contributors may help understanding SARS-CoV-2
transmission patterns that include family pets and humans. As
the contribution of animals, especially pets, to viral spread in
households remains undetermined, it is important to have a plan
of care in place if a family pet tests positive for COVID-19.
References
- Timeline of WHO’s response to COVID-19 [Internet].
- Center for Systems Science and Engineering, Johns Hopkins University. COVID Dashboard. Johns Hopkins University. 2021.
- Michelitsch A, Wernike K, Ulrich L, Mettenleiter TC, Beer M (2021) SARS-CoV-2 in animals: From potential hosts to animal models. In: Advances in Virus Research.
- XL, JZ, QZ, QN, YL, et al. (2020) Evolutionary history, potential intermediate animal host, and cross-species analyses of SARS-CoV-2. J Med Virol 92(6): 602-611.
- Latinne A, Hu B, Olival KJ, Zhu G, Zhang L, et al. (2020) Origin and cross-species transmission of bat coronaviruses in China. Nat Commun 11: 4235.
- Helmy YA, Fawzy M, Elaswad A, Sobieh A, Kenney SP, et al. (2020) The COVID-19 pandemic: A comprehensive review of taxonomy, genetics, epidemiology, diagnosis, treatment, and control. J Clin Med 9(4): 1225.
- Astuti I, Ysrafil (2020) Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2): An overview of viral structure and host response. Diabetes Metab Syndr Clin Res Rev 14(4): 407-412.
- Schoeman D, Fielding BC (2019) Coronavirus envelope protein: Current knowledge. Virol J 16(1): 69.
- Luan J, Lu Y, Jin X, Zhang L (2020) Spike protein recognition of mammalian ACE2 predicts the host range and an optimized ACE2 for SARS-CoV-2 infection. Biochem Biophys Res Commun 526(1): 165-169.
- Jinghua Lu, Peter D Sun (2020) High affinity binding of SARS-CoV-2 spike protein enhances ACE2 carboxypeptidase activity. bioRxiv.
- Mahmoud IS, Jarrar YB, Alshaer W, Ismail S (2020) SARS-CoV-2 entry in host cells-multiple targets for treatment and prevention. Biochimie 175: 93-98.
- Mousavizadeh L, Ghasemi S (2020) Genotype and phenotype of COVID-19: Their roles in pathogenesis. J Microbiol Immunol Infect 54(2): 159-163.
- Kuba K, Imai Y, Ohto-Nakanishi T, Penninger JM (2010) Trilogy of ACE2: A peptidase in the renin-angiotensin system, a SARS receptor, and a partner for amino acid transporters. Pharmacol Ther 128(1): 119-128.
- Yan R, Zhang Y, Li Y, Xia L, Guo Y, et al. (2020) Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 367(6485): 1444-1448.
- S R Tipnis, N M Hooper, R Hyde, E Karran, G Christie, et al. (2000) A human homolog of angiotensin-converting enzyme. Cloning and functional expression as a captopril-insensitive carboxypeptidase. J Biol Chem 275(43): 33238-33243.
- Huang L, Sexton DJ, Skogerson K, Devlin M, Smith R, et al. (2003) Novel peptide inhibitors of angiotensin-converting enzyme 2. J Biol Chem 278(18):15532-15540.
- Østergaard ME, Nichols J, Dwight TA, Lima W, Jung ME, et al. (2017) Fluorinated Nucleotide Modifications Modulate Allele Selectivity of SNP-Targeting Antisense Oligonucleotides. Mol Ther - Nucleic Acids 7: 20-30.
- Imai Y, Kuba K, Rao S, Huan Y, Guo F, et al. (2005) Angiotensin-converting enzyme 2 protects from severe acute lung failure. Nature 436: 112-116.
- Towler P, Staker B, Prasad SG, Menon S, Tang J, et al. (2004) ACE2 X-Ray Structures Reveal a Large Hinge-bending Motion Important for Inhibitor Binding and Catalysis. J Biol Chem 279(17): 17996-8007.
- Hong Kong (2020) Detection of low level of COVID-19 virus in pet dog. The Government of the Hong Kong Special Administrative.
- Pet cat tests positive for COVID-19 virus (2021).
- Almendros A (2020) Can companion animals become infected with Covid-19? Veterinary Record.
- Almendros A (2019) Can pets transmit Covid-19 infection? Open Vet J.
- PRO/AH/EDR> COVID-19 update (58): Belgium, animal, cat, clinical case, RFI [Internet]. 2020.
- PRO/AH/EDR> COVID-19 update (149): France (IF) animal, cat, owned [Internet]. 2020.
- PRO/AH/EDR> COVID-19 update (181): Germany (BY), France (AC), cat, OIE animal case defin. [Internet]. 2020.
- SARS-CoV-2, Russia [Internet].
- SARS-CoV-2/COVID-19, United States of America.
- Shi J, Wen Z, Zhong G, Yang H, Wang C, et al (2020) Susceptibility of ferrets, cats, dogs, and other domesticated animals to SARS-coronavirus 2. Science 368(6494): 1016-1020.
- Sia SF, Yan LM, Chin AWH, Fung K, Choy KT, et al. (2020) Pathogenesis and transmission of SARS-CoV-2 in golden hamsters. Nature 583: 834-838.
- Dodds WJ (2020) Coronavirus SARS-CoV-2 (COVID-19) and Companion Animal Pets. J Immunol Allergy.
- Halfmann PJ, Hatta M, Chiba S, Maemura T, Fan S, et al. (2020) Transmission of SARS-CoV-2 in Domestic Cats. N Engl J Med 383(6): 592-594.
- What you need to know about COVID-19 and Pets and Other Animals. 2020.
- S. pet ownership statistics | American Veterinary Medical Association.
- USDA APHIS | Cases of SARS-CoV-2 in Animals in the United States. 2021
- Yan Y, Tao H, He J, Huang SY (2020) The HDOCK server for integrated protein–protein docking. Nat Protoc 15: 1829-1852.
- Sievers F, Higgins DG (2014) Clustal omega, accurate alignment of very large numbers of sequences. Methods Mol Biol 1079: 105-116.
- Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, et al. (2007) Clustal W and Clustal X version 2.0\r10.1093/bioinformatics/btm404. Bioinformatics 23(21): 2947-2948.
- National Center for Biotechnology Information (2021).
- Stecher G, Tamura K, Kumar S (2020) Molecular evolutionary genetics analysis (MEGA) for macOS. Mol Biol Evol 37(4): 1237-1239.
- Datta PK, Liu F, Fischer T, Rappaport J, Qin X (2020) SARS-CoV-2 pandemic and research gaps: Understanding SARS-CoV-2 interaction with the ACE2 receptor and implications for therapy. Theranostics 10(16): 7448-7464.
- Shen M, Liu C, Xu R, Ruan Z, Zhao S, et al. (2020) Predicting the Animal Susceptibility and Therapeutic Drugs to SARS-CoV-2 Based on Spike Glycoprotein Combined With ACE2. Front Genet 11: 575012.
- Orangutans Receive COVID-19 Vaccine (2021).
- Zoetis Donates COVID-19 Vaccines to Help Support the Health of Zoo Animals (2021).
- One Health Toolkit for Health Officials Managing Companion Animals with SARS-CoV-2.
- Iboi EA, Sharomi O, Ngonghala CN, Gumel AB (2020) Mathematical modeling and analysis of COVID-19 pandemic in Nigeria. Math Biosci Eng 17(6): 7192-7220.
- Aslan IH, Demir M, Wise MM, Lenhart S (2020) Modeling COVID-19: Forecasting and analyzing the dynamics of the outbreak in Hubei and Turkey. medRxiv.
- Lahiri A, Suman Jha S, Bhattacharya S, Ray S, Chakraborty A (2020) Effectiveness of preventive measures against COVID-19: A systematic review of In Silico modeling studies in indian context. Indian J Public Health 64(Supplement): S156-S167.
- Zhao S, Chen H (2020) Modeling the epidemic dynamics and control of COVID-19 outbreak in China. Quant Biol 1-9.
- Lu Z, Yu Y, Chen YQ, Ren G, Xu C, et al. (2020) A fractional-order SEIHDR model for COVID-19 with inter-city networked coupling effects. Nonlinear Dynamics 101: 1717-1730.
- Atangana A (2020) Modelling the spread of COVID-19 with new fractal-fractional operators: Can the lockdown save mankind before vaccination? Chaos, Solitons and Fractals 136: 109860.




































