Identification of Bovine Mastitis Associated Pathogens by Multiplex PCR
Anjali Gangwal and Kashyap SK*
Department of Microbiology and Biotechnology, India
Submission: September 09, 2017; Published: September 22, 2017
*Corresponding author: Kashyap SK, Department of Microbiology and Biotechnology, College of Veterinary & Animal Science, India, Email: gangwalanjali17@gmail.com
How to cite this article: Anjali G, Kashyap SK. Identification of Bovine Mastitis Associated Pathogens by Multiplex PCR. Dairy and Vet Sci J. 2017; 3(5): 555622. DOI: 10.19080/JDVS.2017.03.555622
Abstract
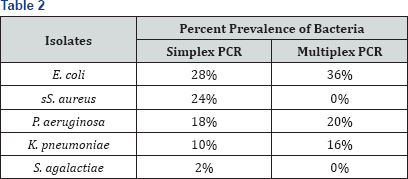
The present study was focused on development of a rapid and efficient method for detecting major pathogens of bovine clinical mastitis- E. coli, S. aureus, P. aeruginosa, K. pneumoniae, S. agalactiae, S. dysgalactiae and S. uberis simultaneously without the need of cultivation of bacteria.A total of 50 clinical mastitic milk samples were processed for isolation and biochemical identification of bacteria which were further confirmed geno typically by PCR. The prevalence of bacteria obtained was E. coli (28%), S. aureus (24%), P.aeruginosa (18%), K pneumoniae (10%) and S. agalactiae (2%). For rapid and accurate diagnosis, DNA was isolated directly from mastitic milk and subjected to multiplex PCR. On multiplexing of same DNA samples with 5 pairs of primers (of all 5 isolates), revealed simultaneous detection of E.coli and K pneumoniae (10%), E. coli, and P aeruginosa (8%), K pneumoniae and P aeruginosa (2%), & E. coli, K. pneumonia and P aeruginosa (2%) with an overall prevalence of E. coli (36%), P aeruginosa (20%)and K pneumonia (16%). However, S. aureus and S. agalactiae were not obtained by multiplexing that might be due to incompatibility of primers or differences in annealing temperatures of primers. Conclusively, molecular methods specifically multiplex PCR can be used as an efficient tool for detecting major pathogens of mastitis in one run with high accuracy and in short period of time.
Keywords: Multiplex; PCR; Diagnosis; Mastitis
Abbreviation: (PCR): Polymerase Chain Reaction
Introduction
Healthy udder of milking cows is one of the most important factors of successful breeding from the health as well as economic point of view. Mastitis is defined as an inflammation of the parenchyma of mammary gland, which not only reduces milk yield but alters milk composition [1] also. It is one of the most problematic diseases and continues to have a major economic impact on the dairy industry throughout the world [2]. It is a devastating disease haunting the dairy industry leading to reservoir of infection for human beings and the economic losses due to high morbidity, discarded milk, reduced milk production and increased antimicrobial resistance of the organisms in animals treated with antibiotics [3]. The genome sequences of many of the major mastitis causing pathogens are now available and can be utilized to develop nucleic acid-based testing methods. Nucleic acid testing by PCR is highly sensitive and specific. The identification of the bacteria using PCR at the species level is based on amplification of a target gene which is mostly rRNA gene, highly conserved in bacteria. It is the only assay that allows simultaneous screening for multiple pathogens that might be causing mastitis. To improve the diagnosis a rapid, sensitive and specific Multiplex polymerase chain reaction (mPCR) assay has been developed for detection of more than one pathogen in single PCR reaction causing mastitis at early stages. In multiplex PCR, multiple pairs of primers specific for different DNA segments are included in the same reaction to enable amplification of multiple target sequences in one assay [4]. This produces amplicons of variable sizes, allowing the identification of targeted bacterial pathogens in a milk sample.
Materials and Methods
Sampling and general microbiological analysis
A total of 50 milk samples from cattle each about 5-10 ml in amount were collected in sterilized test tubes. The milk samples were collected from Veterinary Clinical Complex and dairy farm in and around the Bikaner city, Rajasthan with clinical mastitis. The samples were collected in the morning and were immediately taken thereafter to the laboratory on ice for further processing.
Isolation and identification of bacteria
Culture characteristics: The milk samples were subjected to aerobic cultivation. Each milk sample was streaked on Nutrient agar, Blood agar and MacConkey (MCA) agar plates in primary, secondary and tertiary fashion in order to obtain isolated colonies of bacteria. These petri plates were incubated for 24hr at 37 °C. After 24hr incubation these isolated colonies were cultured on Mannitol salt agar (MSA), Edward's medium agar, Eosine methylene blue agar (EMB), Mac-conkeyand Cetrimide agar plates respectively for isolation of i>Staphylococcus aureus, Streptococcus spp., Escherichia coli, Klebsiella pneumoniae and Pseudomonas aeruginosa and incubated for 24 hours at 37 °C. The growth was examined for the colonial morphology and pigmentation, and different types of colonies were sub-cultured on separate nutrient agar plates in order to obtain pure cu.
Biochemical characteristics: Cultures were subjected to primary biochemical tests viz. Catalase test, Oxidase test,and secondary biochemical tests viz. Oxidation-Fermentation test, Mannitol fermentation, Growth on Mac-Conkey agar, 6.5% Sodium Chloride tolerance test, Aesculin Hydrolysis, Pigment production, Gelatinliquefaction, IMViC tests, growth on TSI slants, Sugar fermentation tests, Urease test, DNAse test, Arginine hydrolysis test, Nitrate reduction test and haemolysis on blood agar etc.
Genotyping: 23S rRNA spacer region gene based identification was carried out for S. aureus, S. agalactiae, S. dysgalactiae, S. uberis and E. coli as per the method described by [5]. 16S rRNA gene based identification as per the method described by [6] for P aeruginosa. 16S-23S rDNA internal transcribed spacer (ITS) region based identification as per the method described by [7] for K. pneumonia (Table1).

Results
Out of the 50 processed clinical mastitic milk samples, 33 (66%) samples were found positive for the presence of bacteria viz. E. coli, S. aureus, P. aeruginosa, K. pneumoniae, S. agalactiae. In remaining 17 (34%) samples of clinical mastitis, no aerobic bacteria could be isolated. A total of 41 bacterial isolates could be cultured and identified by biochemical and genotypic test which consist of 14(28%) isolates of E. coli, 12(24%) isolates of S. aureus, 9(18%) isolates of P aeruginosa, 5(10%) isolate of K. pneumonia and 1(2%) isolate of Streptococcus agalactiae.
All 14 isolates of E. coli on preliminary biochemical characterization revealed characteristic IMViC pattern, growth on TSI (Y/Y/-) and fermented different sugars. Preliminary biochemical characterization of all 12 isolates of S. aureus revealed characteristic growth pattern on mannitol salt agar, showed haemolysis on blood agar, DNAse test and fermented different sugars. All 9 P aeruginosa isolates revealed standard biochemical characterization as growth at 42 °C, pigment production, growth on triple sugar iron (TSI) agar (R/R/-), citrate utilization, arginine hydrolysis, gelatin liquefaction, haemolysis and nitrate reduction. Negative reaction was observed for aesculin hydrolysis and urease tests. All 5 isolates of K. pneumoniaeon preliminary biochemical characterization revealed characteristic IMViC pattern, growth on TSI (Y/Y/-) and fermented different sugars. One Streptococcus agalactiae isolate showed dew drop like colonies on Edward's medium, 6.5% sodium chloride medium, hydrolysedaesculin and fermented different sugars.
Molecular diagnosis of common bacterial pathogens directly from bovine clinical mastitic milk
All the 50 samples were subjected for DNA extraction directly. A standard protocol was followed for the isolation of DNA from milk sample which is simple and rapid method as per Nachimutu et al. [8] with slight modifications. The method was easy to perform and requires minimal sample manipulation.
All the established species primer pairs were used for evaluation of 5 bacterial species of bacteria in multiplex PCR. The results of multiplex PCR revealed simultaneous detection of E. coli and P. aeruginosa (8%), E. coli and K. pneumoniae (10%), aeruginosa (2%). However S. aureus and S. agalactiae were not P. aeruginosa and K. pneumoniae (2%) and E. coli, K. pneumoni, P detected by multiplex PCR.
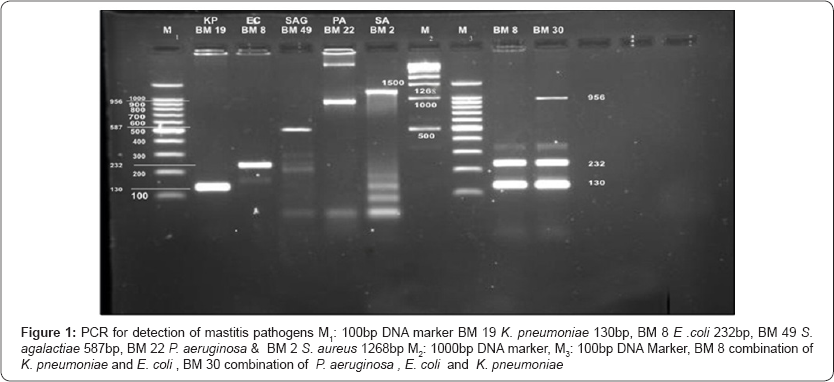
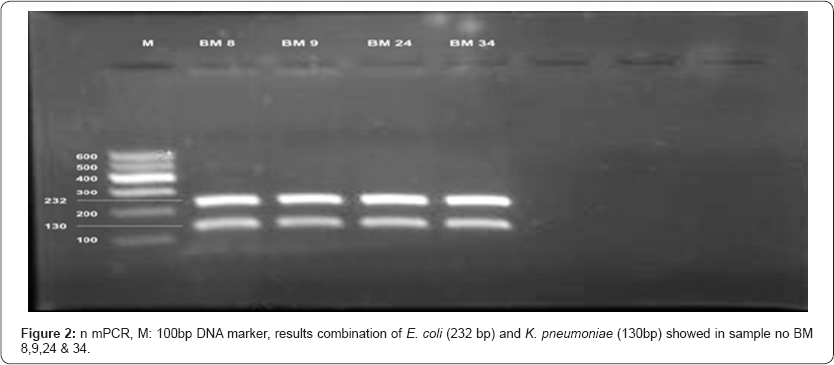
Comparative study of bacterial isolate, DNA with simplex PCR and DNA directly from milk with multiplex PCR
Discussion
Bovine mastitis is an important disease of cattle which causes economic losses in the country by affecting the milk quality and milk quantity. Mastitis is of multi-etiological in nature; loss is due to the major bacterial pathogens, Staphylococcus aureus, Streptococcus spp, Escherichia coli, Klebsiella pneumoniae and Pseudomon asaeruginosa. Diagnosis is more important in the case of mastitis. We can use conventional method as culture and advanced method as molecular. Compared to culture, molecular assays are less time consuming, takes less than 24 hours to complete, while identification of bacteria to the species levels by conventional microbiological and biochemical methods requires more than 72 hours. Conventional procedures for the identification of Bovine Mastitis pathogens are labor-intensive and most of the commercial identification systems are not designed to identify important mastitic pathogens [9,10]. In conventional method and molecular methods higher prevalence of E. coli found in the present study due to environment unhygienic condition which is somewhat similar with Bradely et al. [11]; Khaled et al. [12]; Sayed [13] who have also reported higher incidence. The biochemical characterization of isolates and fermentation of sugars are comparable with studies of Kruthi [14] and Pachaury [15]. In the present investigation all E. coli isolates were subjected to PCR amplification targeting 23S rRNA gene using species specific primers. Species specific amplicon of 232bp are similar with results of Riffon et al. [5] who found amplicon of this size. All 12 isolates of S. aureus showed standard biochemical characterization and sugar fermentation which similar with Upadhyay (2009) who carried out fermentation studies with S. aureus obtained from milk of mastitic cattle and goat. Streptococcus spp. isolate biochemically confirmed and were subjected to PCR amplification targeting 23S rRNA spacer region by using S. agalactiae, S. dysgalactiaeand S. uberisspecies specific primers with 587bp, 403bp and 95bp. 23S rRNA spacer region based studies were conducted by Riffon et al. [5] for genotypic confirmation of Streptococcus agalactiae isolated from bovine mastitic milk and they observed 587 bp PCR product. All 8 isolates which had been biochemically identified as P aeruginosa were subjected to genotyping using species specific primers reported earlier (Spilkeret al., 2004). In the present investigation all 5 isolates had been biochemically identified as K. pneumonia and were subjected to PCR amplification targeting 16S-23S rDNA internal transcribed spacer (ITS) region based specific primers [7]. Our results are completely in agreement with [7].
As the present study is focused on development of a rapid and early detection method, a multiplex PCR was devised to allow the simultaneous detection of multiple pathogens of mastitis in a single reaction, which requires smaller amounts of reagents and even lesser time than simplex PCR [5,16]. This multiplex PCR method appeared as a useful tool to resolve bacterial aetiology of mastitis milk samples simultaneously and at the earliest [12,17,18].
Multiplexing is alternative method however, the compatibility of the primer pairs is important for the reaction to work efficiently with high sensitivity of detection. All the established species primer pairs were used for evaluation of 5 bacterial species in multiplex PCR. The results of multiplex PCR revealed simultaneous detection of E. coli and P. aeruginosa (8%), E. coli and K. pneumoniae (10%), P aeruginosa and K. pneumoniae (2%) and E. coli, K. pneumoniae and P aeruginosa (2%) which is compare with Pradhan 2011 who used multiplex PCR as routine detection of S. aureus, Streptococcus spp. and E. coli in milk samples and However, S. aureus and S. agalactiae were not detected by multiplex PCR but various previous studies as done by Shome et al. [19] were able to detect 10 bacterial strains simultaneously by multiplex PCR. This disparity in results of simplex and multiplex PCR might be due to incompatibility of primers or differences in annealing temperatures. The multiplex PCR assay developed in this study provides a convenient means of accurately identifying important bacterial pathogens of mastitis simultaneously.
In cultural isolation, colonies are selected based on phenotypic characteristics like size, color, shape etc. There can be possibility of missing out colonies that are having similar phenotype to the predominant ones but very few in numbers. Similarly, possible reasons for no growth in milk samples can include low concentration of bacteria in the milk samples, pathogens not growing in standard culture media, or presence of substances in the milk decreasing the viability of bacteria in culture [20]. When used for the detection of pathogens in subclinical mastitis milk samples, mPCR was found to be more efficacious than microbiological studies. The mPCR successfully detected all bacteria, as isolated in culture method. Besides, mPCR detected organisms in samples found culture-negative. Culture-negative milk samples in conventional bacteriology represent a large proportion of positive samples in molecular methods [20].
Conclusion
mPCR assay can be used for rapid, sensitive, specific and reliable identification of the major mastitis pathogens from milk. As several bacterial species are involved in mastitis infection, inadequate pathogen detection or identification techniques often delay timely interventions in disease treatment and control.
References
- Souto LI, Minagawa CY, Telles EO, Garbuglio MA, Amaku M, et al. (2010) Correlation between mastitis occurrence and the count of microorganisms in bulk raw milk of bovine dairy herds in four selective culture media. J Dairy Rese 77(1): 63-70.
- Bachaya HA, Raza MA, Murtaza S, Akbar IUR (2011) Subclinical bovine mastitis in MuzaffarGarh district of Punjab (Pakistan). Journal of Animal and Plant Sciences 21(1): 16-19.
- Sumathi BR, Veeregowda BM, Gomes AR (2008) The occurrence and anti biogram of bovine mastitis. Veterinary World 1(8): 237-238.
- Henegariu O, Heerema NA, Dlouhy SR, Vance GH, Vogt PH (1997) Multiplex PCR-critical parameters and step-by-step protocol. Biotechniques 23(3): 504-511.
- Riffon R, Sayasith K, Khalil H, Dubreuil P, Drolet M, et al. (2001) Development of a Rapid and Sensitive Test for Identification of Major Pathogens in Bovine Mastitis by PCR. Journal of Clinical Microbiology 39(7): 2584.
- Spilker T, Coenye T, Vandamme P, LiPuma JJ (2004) PCR-Based Assay for Differentiation of P.aeruginosa from other Pseudomonas Species Recovered from Cystic Fibrosis Patients. Journal of Clinical Microbiology 42(5): 2074-2079.
- Liu Y, Liu C, Zheng W, Zhang X, Yu J, et al. (2008) PCR detection of Klebsiella pneumonia in infant formula based on 16S-23S internal transcribed spacer. International Journal of Food Microbiology.125: 230-235.
- Nachimuttu K, Ramadas P, Thiagarajan V, Dhinakar Raj G, Kumanam K (2001) Laboratory manual on Polymerase chain reaction based methods for diagnosis. Animal Science University India. pp 5-13.
- Jayarao BM, Dore JJ, Baumbach GA, Matthews KR, Oliver, SP (1991) Differentiation of Streptococcus uberis from Streptococcus parauberis by polymerase chain reaction and restriction fragment length polymorphism analysis of 16S ribosomal DNA. Journal of Clinical Microbiology 29: 2774-2778.
- Watts JL (1989) Evaluation of the Minitek Gram-Positive Set for identification of streptococci isolated from bovine mammary glands. Journal of Clinical Microbiology 27(5): 1008-1010.
- Bradley AJ (2002) Bovine mastitis: evolving diseases. The Veterinary journal 163:1-13.
- Khaled A, Abd El-Razik K, Abdelrahman A, Yousse FA, Alaa MG, et al. (2010) Direct Identification of Major Pathogens of the Bubaline Subclinical Mastitis in Egypt using PCR. Journal of American Sciences 6(10).
- Sayed SM (2014) A contribution on Coliforms causing mastitis in cows with reference to serotypes and virulence factors of E. coli isolates. Assiut University Bulletin for Environmental Researches 17, Egypt.
- Kruthi M (2006) Phenotypic characterization of E. coli strainsisolated from human intestinal and urinary tracts. Thesis submitted for MSc Biotechnology, Wright state university, Fairborn, USA.
- Pachaury R (2011) Genotypic and phenotypic characterization of Escherichia coli isolated from diarrhoeic calves, lambs and kids. MVSc Thesis submitted to RAJUVAS, Bikaner, Rajasthan, India.
- Phuektes P, Mansell PD, Browning G (2001) Multiplex Polymerase Chain Reaction Assay for Simultaneous Detection of Staphylococcus aureus and Streptococcal Causes of Bovine Mastitis Journal of Dairy Sciences 84: 1140-1148.
- Forsman P, Tilsala-Timisjarvi A, Alatossava T (1997) Identification of staphylococcal and streptococcal causes of bovine mastitis using 16S-23S rRNA spacer regions. Microbiology 143(11): 3491- 3500.
- Gangwal A, Kashyap SK, Katiyar S, Meena D, Boyal P, et al. (2017) Isolation and Identification of Common Mastitis Causing Pathogens from Clinical Bovine Mastitis Milk. Journal of Pure and Applied Sciences 11(1): 329-333.
- Shome BR, Das Mitra S, Bhuvana M, Krithiga N, Velu D, et al. (2011) Multiplex PCR assay for species identification of bovine mastitis pathogens. J App Microbiology 111(6): 1349-1356.
- Taponen S, Salmikivi L, Simojoki H, Koskinen MT, Pyorala S, (2009) Real Time Polymerase chain reaction based identification of bacteria in milk samples from bovine clinical mastitis with no growth in conventional culturing. Journal of Dairy Science 92(6): 2610-2617.






























