Laser Micro Dissection Systems: An Alternative Method of Defined Cell Isolation for Molecular Profiling
Ratan K Choudhary*
School of Animal Biotechnology, Guru Angad Dev Veterinary and Animal Science University, Ludhiana- 141004, Punjab, IndiaSubmission: October 01, 2016; Published:January 03, 2017
*Corresponding author: Ratan K Choudhary, School of Animal Biotechnology, Guru Angad Dev Veterinary and Animal Science University, Ludhiana- 141004, Punjab, India; Email: vetdrrkc@gmail.com
How to cite this article: Ratan K C. Laser Micro Dissection Systems: An Alternative Method of Defined Cell Isolation for Molecular Profiling. Dairy and Vet Sci J. 2017; 1(1): 555552. DOI: 10.19080/JDVS.2017.01.555552
Abstract
Laser micro-dissection permits isolation groups of cells from heterogeneous tissue sections under direct microscopic visualization. This technology not only allows isolation of homogenous subpopulation of cells but also allows immunophenotypically similar cells for RNA/DNA or protein extraction for downstream applications like western blot, mass spectrophotometry, real time quantitative polymerase chain reaction, microarray and RNA sequencing. Isolation of dead cells unsuitable for functional assays, high cost of equipment and technical expertise required for operation are few limitations to laser microdissection systems.
Keywords: Laser microdissection; mammary tissue; carcinoma; immunolabelled cell; transcriptomics; proteomics
Abbreviations: LMD: Laser microdissection;LCM:Laser capture microdissection; UV: Ultraviolet; IR: Infrared rays; PEN:Polyethylene naphthalate; PPS:polyphenylenesulphide; RNAseq: RNA sequencing; BrdU:bromodeoxyuridine; qPCR: quantitative polymerase chain reaction; xMD:expression microdissection; 2D: Two dimensional; iTRAQ: Isobaric tag for relative and absolute quantitation; LREC: Label retaining epithelial cells
Introduction
Principles of laser microdissection systems
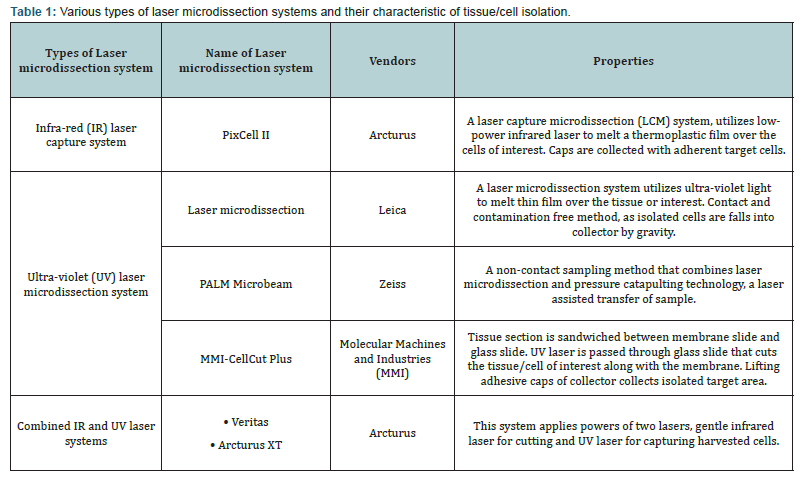
There are two types of laser based micro dissection systems, ultra-violet (UV) and Infrared (IR) laser based systems. IR laser micro dissection system transfers cells into thermoplastic polymer (called “capture) from which molecules are extracted for downstream applications. Therefore, this system is called laser capture micro dissection (LCM) system. The principle of LCM is depicted in which tissue sections are placed on the stage of microscope. Thermoplastic sheet (a transparent ethylene vinyl acetate layer, usually 6 mm in diameter) is placed onto the tissue section. When IR laser beam is focussed, thermoplastic sheet just above the tissue section melts and surrounds (i.e. captures) the tissue/cells of interest [1,2]. Later on thermoplastic sheet, embedded tissue/cells are harvested in a tube’s cap for nucleic acids or protein isolation for downstream applications. Initially, this technique was applicable on the frozen tissue sections [3]. Later on, this technique is used on formalin fixed, paraffin embedded tissue sections, smears, cytospins, live cells for RNA, DNA or protein isolations [4,5].
Laser micro-dissection (LMD) is UV laser based system where tissue sections are placed on glass membrane slides. Polyethylene naphthalate (PEN) or polyphenylene sulphide (PPS) membranes are cemented onto glass slides, commonly called glass membrane slides. Tissue sections placed on the top of the membrane and slides are kept inverted, facing tissue sections downwards towards collector. When UV laser passed from upward passes through the glass slides, it cuts the membrane. The areas of interest are selected on computer screen and membranes associated with area of interest are deposited into collector through gravity. Unfortunately, UV laser damages the molecules that comes in the cut perimeter especially when area of interests are very small like single cell.
The combination of IR laser capture and UV laser cutting (Arcturus XT™ micro dissection system, from Life Technology, as an example) is really a promising one to perform both types of work under the same platform. Three different types of laser micro-dissection systems include, IR laser system, UV laser system and combined laser systems (Table 1). In this review, usefulness LMD on isolation of different type of cells, in particularmammary tissue, including isolation of pathogens from thetissue section, has been discussed. Author believes that there isnot much reviews focussed on application of LMD for isolationof various types of cells from complex tissue in veterinary and animal sciences are present. Furthermore, molecular profilingsuch as RNA seq or proteomics study of micro dissected cellswould certainly provide insights of biology of harvested cellsfrom their tissue.
In this review, usefulness LMD on isolation of different typeof cells, in particular mammary tissue, including isolation ofpathogens from the tissue section, has been discussed. Authorbelieves that there is not much reviews focussed on applicationof LMD for isolation of various types of cells from complex tissuein veterinary and animal sciences are present. Furthermore,molecular profiling such as RNAseq or proteomics study ofmicrodissected cells would certainly provide insights of biologyof harvested cells from their tissue.
Isolation and molecular profiling of defined cells fromtissue
Mammary epithelial cells and stroma
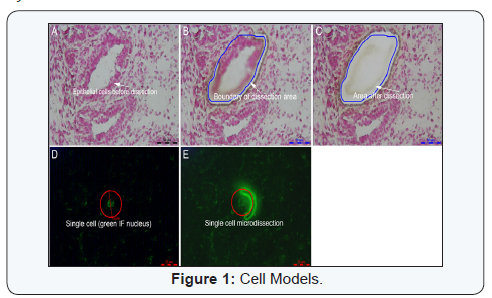
Information on use of laser micro dissection system tocharacterize mammary tissue is limited. Mammary tissue iscomposed of mammary epithelium and stroma embeddedin the adipose tissue. Isolation of epithelial cells and stromaindividually could provide unique and common characteristicsof mammary epithelial cells and stroma, respectively that couldprovide reliable cell-specific gene expression and functions.Considering that RNA quality decides transcriptomic analysisof tissue harvested after micro dissection, it is important toprocess frozen section quickly. Staining of frozen sections usinghaematoxylin or cresyl violet or fast red or toluidine blue stainsin nuclease free conditions provided isolation of good qualityRNA from mammary epithelium and stroma [6-10] (Figure 1AC).
Laser micro dissection also permitted investigators to studythe immune-modulatory role of mammary epithelial cells duringmastitis. In goats, mammary gland (one quarter) infected withpathogens while the other half gland (other quarter) served ascontrol. Mammary epithelial cells were dissected out leavingcontaminating inflammatory cells. Investigators examinedepithelial cells specific gene expression profiles [13]. In addition to the transcriptomic analysis of micro dissected mammarytissues using microarray and real time qPCR, RNA Sequencing(RNA seq) also examined gene profile [14]. RNA seq is the adventof next generation sequencing that offer more advantagesover microarray because of its ability to identify all genesincluding novel gene, splice variants, allele-specific expression,independent of genome annotation and absolute quantificationof copy numbers transcripts [15,16].
MNerve cells
Molecular analysis of class specific neuronal developmentrequires identification of pure population of class specificneurons. Identification and isolation of pure population ofperipheral neurons that were performed using magnetic beadsorting [17] and florescent activated cell sorting did not procureaccurate isolation of specific cells. However, laser capture microdissection provided accurate and precise isolation of dendriticneurons, motor nerves, dopaminergic, catecholamine neuronsand astrocytes for understanding molecular mechanism of classspecificdendrite morphogenesis and progression of disease likeParkinson’s [18-26]. Identification of similar cells that differsimmune phenol typically coupled with laser micro dissectionenabled RNA isolation [3,1][27-30] for gene expression analysislike qPCR. Alternatively, in thin serial sections, parvalabuminneurons were identified immune histo chemically in one sectionand adjacent sections simply stained with neutral red.
In neutral red sections, Neuronal profiles of parvalbuminimmune stained section identified cell bodies, subjected to lasermicro-dissection. This method located parvalbumin neuronswithout immune histo chemical processing and provided intactRNA [29]. Since neuronal cells are big in size, it was possibleto compare immune positive cells profile of one section to theserial section. Utility of this alternative method to identify cellsof interest among the other tissue needed to evaluate. Molecularsignature of glioma has been investigated where invading gliomatissue were harvested from rim of tumour cells mass [31-33].These studies investigated gene signature of glioblastoma ofbrains. However, these studies do not emphasize on the roleof tissue microenvironment on development of glioblastoma.Microenvironment plays an important role in orchestratinginvasion of cancerous cells into normal tissue. Laser microdissectionallows isolation of different types of tissue- gliomacancer cells, invading glioma cells and cells of normal cortex.This comprehensive type of tissue sampling permitted study oftumour microenvironment and gene regulatory networks andpathways responsive for glioma invasion [34].
Laser micro dissection can also be used to identify geneknockout cells at genomic level. Micro dissection of motorneurons from spinal cord for DNA isolation led identificationof neuron specific ablation of Mtmr2 gene inknock out miceusing PCR [35]. Identification of dopaminergic neurons fromsubstantia nigra pars compacta using laser captured microdissection determined transcriptional changes in aged-matched and young control individuals and revealed mitochondrial DNAdeletions in substantia nigra neurons in ageing [36]. One of thecauses for Parkinson’s disease was the substantial degradationof somatic mitochondrial DNA in aged-matched individual.In addition to finding a reason to prevent loss of somaticmitochondrial DNA loss in the disease, pathway analysis of generegulatory networks responsible for neuronal loss could resultin therapeutic interventions of Parkinson’s disease [36,24].
Underlying mechanisms of cell survival rheostat switchwhich controls dying or surviving cells after traumatic braininjury, was the result of single neuronal cell (injured vs healthycells from adjacent area) isolation using laser micro dissectionsystems [37]. Degeneration of motor neurons in amyotrophiclateral sclerosis has identified involvement of RNA bindingproteins, TDP43 and FUS. The genes involved with theseproteins are normally transcribed, evidenced by RNA sequencingof motor neurons of mutant vs. wild-type mice [38] that werepurely isolated using laser micro dissection. Molecular basisof diseases of nervous systems like Parkinson’s disease orschizophrenia, where particular types of neurons are vulnerablefor degeneration, can be studied in detail upon their isolationindividually in pure form [39].
Muscle cells
Skeletal muscle fibre is composed of various types of cellsincluding microsatellite stem cells, neurons and vascularendothelium enabling molecular profiling of specific cell typesat myofibril level difficult. Laser micro dissection systemsoffer expression profile of individual cell types like myofibrils,satellite stem cells, vascular elements and neuronal elementsfrom muscle fibres [40-43]. Identification of type I, type IIA, typeIIB and type IIC of myofibrils and then isolation of these differenttypes of myofibrils using laser micro dissection provided insightin understanding the biology of muscular atrophy [44].
Molecular signature of laser micro dissected smooth musclecells resulted in identification of gene involved in diseases likeasthma, atopic and hypoxia-induced hypertension [45-47].Asthma and atopy are the respiratory tract diseases havingsimilar symptoms except asthma obstructs airways whereasatopy does not. Out of one hundred differentially expressed genesbetween asthma and non-asthma (including atopy) patients, aset of eight genes distinguished asthmatic from non-asthmaticpatients [47].
Cancer cells
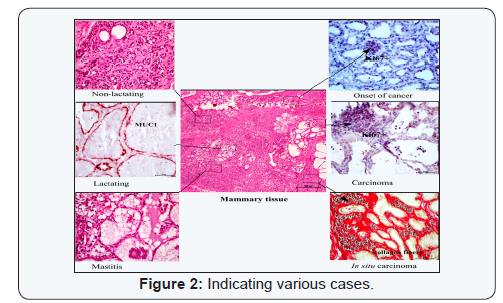
Cancer tissues are well known for tissue heterogeneityhaving different types of tissues e.g., healthy cells, cancerous cells,invasive cancerous cells, infiltrating immune cells, proliferatingcells of blood vessels and surrounding stroma (Figure 2). Breastcarcinoma, for example, is a mixture of different cells types asrevealed by molecular analysis of the tissue [48-50]. Becauseof such heterogeneity of tissues, the RNA or protein isolatedfrom cancer tissues actually represents transcripts or proteins of various cell types present in the cancerous tissue underexamination. Thus, pool of transcripts or protein derived fromvarious cell types of tumour, prohibits prognostic discovery ofcell-specific markers and drug discovery. Isolation of primarybreast tissue and corresponding metastatic lymph nodesusing laser micro dissection provided insights of metastaticprogression of breast cancer [51].
Protocol of performing staining, micro dissection of cancerspecific cells from heterogeneous mixture of breast tumours,protein isolation from micro dissected cells and protein biomarkerdiscovery using mass spectrophotometric are provided fromfrozen tissue (Braakman et al. 2011)[52]. Microenvironmentoften drives cells for normal or abnormal phenotypes. In breastcancer, understanding stromal component of tumour thus offerspromise to understand tumour-associated changes. Isolationof tumour cells and stromal cells surrounding cancer cellsusing laser micro dissection offered study of tumour associatedmicroenvironments [53]. Proteomic analysis of different stagesof colorectal cancers, it was found that tenascin-C was a potentialstromal biomarker for colorectal cancer metastasis [54]. It wasa combined methodology of LCM, iTRAQ labelling and liquidchromatography-tandem mass spectrometry (2D LC-MS/MS) ofstromal proteins from various stages of colorectal cancer.
Potential markers (AnnexinA2, HSP27, CK19 and 14-3-3σ)of lymph node metastasis of lung squamous cell carcinomaidentified using 2D-gel electrophoresis of laser captured microdissected cell in human lungs. Such differentially expressedproteins may serve as protein biomarker of metastatic lungsquamous cell carcinoma and provides molecular basis ofmetastasis that might provide a way for predictive diagnosis andoutcome of patient’s treatment [55]. A proteomic profiling ofvarious types of lung adenocarcinomas using LMD identified 840proteins in which early lepidic-types of lung adenocarcinomaswas associated with numerous tumour progression pathways[56].
Combination of LCM and mass spectrophotometry revealed14-3-3-mediated signalling pathways as the most relevantcanonical pathway of colorectal cancer [56]. Likewise, proteomicstudy of LCM islets, LCM acinar tissue and EIC islets proteomesenabled accurate and unbiased profiling of islet proteome. Such studies could enable study of progression of type1 diabetes ofpancreas and progression of other diseases where heterogeneoustypes of cells are involved [57]. Thus, laser micro dissection offersa reliable method to enrich specific cell population for proteomeanalysis. Proteome profiling of 38 breast cancer tissues inwhich, comparison of whole cell lysates and LCM enrichedtumour epithelial cells were compared, identified more numberof proteins in captured epithelial cells [58]. LCM and LMD alsoovercome limitation of working with small sample size such asthe early lesion of cancer of frozen or formalin fixed paraffinembedded tissue sections [59]. Thus, laser micro dissectiondisregards influence of tissue heterogeneity, low interferencefrom surrounding tissue and offers a better tool for biomarkerdiscovery in meagre of sample.
Microorganisms
Less known application of laser micro dissection is theisolation and characterization of selective microorganismsfrom biological samples. Bart De Spiegeleer and colleagues[60] described selective isolation of microorganisms frommilk using laser capture micro dissection. Investigators haveidentified Methylobacterium spp. using DNA sequencing of16SrRNA gene. Phylogenetic identification of unidentifiedand uncultured microorganisms has been done reliably on thebasis of ribosomal RNA sequence [61,62]. Likewise, a combinedapproach of fluorescent in situ hybridization and laser capturemicro dissection has identified Brachyspira spp from humangut. Bacteria were targeted with oligo nucleotide probe against16S RNA in paraffin embedded formalin fixed tissue sections,fluoresced green and regions of interest were micro dissectedfor DNA sequencing [63].
Use of LCM to harvest Lawsonia intracellularis infectedpigs enterocytes has enabled investigator to characterizehost-bacterial interactions. With the help of RNA sequencingof bacterial infected enterocytes, it was shown that bacteriaprevent differentiation of enterocytes by repressing membranetransporters related nutrient uptake and prevented themselvesfrom oxidative stress by utilizing host’s Zn/cu superoxidedismutase machinery [64]. In a recent study, LCM successfullyused to harvest intracellular malarial parasites from avianerythrocytes for whole genome sequencing [65]. LCM offerinvestigator to isolate parasitic DNA without contamination ofhost DNA. Thus, LCM might offer isolation of individual parasitesfrom co-infected hosts and also help in isolation of rear speciesof parasites from museum/achieved samples [65].
Molecular profiling of immune-selected cells
In majority of the situations, precise identification ofphenotypically similar but immune phenotypically dissimilarcell types are not useful by standard histo chemical stainingprocedures. Only immune staining with immune specificmarkers offer unequivocal identification of precise cell types.Mammary stem/progenitor cells of bovine mammary gland havebeen identified using bromodeoxy uridine (BrdU) label retention study in which BrdU label retaining epithelial cells (LREC)were micro dissected and expression study were performedusing microarray analysis [13]. In this study, LREC (mammarystem/progenitor cells) and non-LREC (differentiated cells fromadjacent locations as control cells) were harvested using lasermicro dissection and total RNA isolated. Microarray analysisrevealed up-regulation of stem cells related genes in LREC incomparisons to non-LREC, provided the evidence of LREC asputative stem/progenitor cells. Immuno staining protocol usingglial fibrillary acidic protein marker permitted precise andaccurate identification of astrocytes and enabled expressionanalysis of laser micro dissected astrocytes using quantitativePCR [66].
To understand biology of different cells of pancreas namely,β-cells, alpha-cells and islets, laser micro dissections employedin normal and pathological conditions to harvest cells of interest.Isolating β-cells of pancreas by identifying fluorescence of antiinsulinantibody has enriched β-cells isolation by LCM [67,68].This protocol prevented contamination of isolated cells fromexocrine cells of pancreas. Protocols for obtaining pancreaticβ-cells [69] and pancreatic duct glands [70] for transcriptomeanalysis have been proposed. Immuno-selected specificcells could also use for comprehensive protein profiling. Inhuman pancreatic adenocarcinoma cells, CD24+ cells weremicro dissected and protein profiles were compared withadjacent normal tissue that were CD24- cells [71]. In this study,investigators have discovered 27 candidate proteins expressedsignificantly different in CD24+ cells than normal tissue andsuggested these proteins as potential biomarkers for pancreaticcancers as potential therapeutic targets.
Expression micro dissection (xMD)
It is an immune-based cell or sub cellular isolation tool whencombined with laser micro dissection system provides rapidprocurement of target of interest from cytological/histologicaltissue samples. This is very useful in where immune positivecells are harvested from the tissue section with increased speedof dissection in multiple magnitude [72,73]. In one recentstudy, tumour cells with estrogen receptor-positive nucleiharvested for large region of tissue in contiguous fashion thusoffered precise sub cellular dissection of targets [74]. In cancerresearch, this tool is going to provide information of neoplastictransformation of cells from normal to cancer type. It has beenobserved that within the same tissue section, various types oftissues are found (Figure 2). Especially in carcinoma, wherecells are invasive carcinoma, hyperplasia, partially transformedcells to normal healthy cells is present. xMD is going to offerenrichment of scanty cells from admixture of cell populationincluding pathogens for next generation sequencing, proteomicsand genomics study.
Limitations of laser micro dissection system
Major consideration during immune staining and lasermicro dissection of tissue or cells is the integrity of RNA for geneexpression analysis and quality of protein for protein analysis.Reduction in immune staining protocol by short incubationtime, maintaining RNase free environment by addition of RNaseinhibitor and nuclease free water and buffers are imperative forsuccessful isolation of intact RNA. In addition to that, testing ofRNA quality using standard procedures like Nano Drop readingand RNA denaturing gel electrophoresis to visualize 28S and18S band might not be applicable as the quantity of RNA is someagre. Only capillary-based electrophoresis system like Agilentbioanalyzer or Bio Rad experion systems could determinequantity of RNA in nano- and pico-grams and quality in terms ofRNA integrity number.
Working with limited amount of RNA introduces problemsrelated to signal to noise ratio in gene expression analysis.Quantity of RNA obtained from laser micro dissected cells rangebetween nano- to pico-grams and the amount of RNA needed formicroarray or RNA sequencing is in microgram. Therefore, it isrequired to amplify total RNA from pico- to micro-gram quantity.Various techniques exists to amplify picogram of RNA intomicrogram quantity that are required for expression analysis[75] indicating that it is feasible and reliable to accuratelyperform expression profiling of sub-optimal quantity of RNA.
Another limitation is the ineffective isolation of homogenouscell types due to lack of distinct cell types of haematoxylinstained sections. Adoption of immune histo chemical stainingprior to LCM will assist researchers for immune phenotypicidentification of cells within tissue. In such cases, identified cellsare dead in nature and hence could not be propagated to testtheir characteristics and also studying of expression dynamicsof the immune stained protein is limited due to binding ofantibody with the protein of interest. Another limitation of lasermicro dissection is that a skilful person and therefore, it usuallyidentify various cell types of tissue could cyto architecture oftissue of interest should be known to the investigators [76].
Conclusion
Laser micro dissection system is a powerful and useful toolin many areas of biological and medical research. The techniqueis now able to produce meaningful, accurate and reproduciblebiology at cellular and molecular level. In cell biology, it is usedin neuroscience research to study various types of nerve cells.In cancer biology, laser micro dissection is used for study themicroenvironment of cancer cells, biology of various cell typesand cancer metastasis. Profiling of various cell types withinepithelial and stromal compartment of mammary tissue ishelpful in delineating their molecular phenotypes. Isolationof microorganisms from tissue is a novel research towardsunderstanding host-pathogen interactions and hence sheds lightinto progression of the disease.
Laser micro dissection could be the most beneficial andadvantageous tool in the context that the area of transcriptomicsand proteomics of selective cells with almost nil contaminationof other cell types. It would be interesting to see advancement inthis technique to isolate live cells so that isolated cells would besuitable for studying cell kinetics and propagation. Additionally,isolation of cell organelles, low cost system and high throughputmicro dissection like expression micro dissection system (xMD)would like help in studying progress of neoplastic transformationof cells and their microenvironments.
Acknowledgment
Author would like to thank Department of Biotechnology,New Delhi (project reference number BT/AAQ/01/AB-I/TFAP/2014) for fund. Author would also like to thank USDA,Beltsville lab during which author gets the expertise of handlinglaser micro dissection under the guidance of Dr. Anthony Capuco.Author would like to acknowledge Dr Devendra Pathak forhelping in capturing microscopic images (Figure 2).
References
- Fend F, Kremer M, Quintanilla-Martinez L (2000) Laser capturemicrodissection: methodical aspects and applications with emphasison immuno-laser capture micro dissection. Pathobiology 68(4-5): 209-214.
- Curran S, McKay JA, McLeod HL, Murray GI (2000) Laser capturemicroscopy. Mol Pathol 53(2): 64-8.
- Fend F, Emmert-Buck MR, Chuaqui R, Cole K, Lee J, et al (1999) Immuno-LCM: laser capture micro dissection of immuno stained frozen sectionsfor mRNA analysis. Am J Pathol 154(1): 61-66.
- Kaneko T, Cordeiro MMR, Okiji T, Kaneko R, Suda H, et al. (2010) Lasercapture microdissection from formaldehyde-fixated and demineralizedparaffin embedded tissues. Microscopy: Science, Technology,Applications and Education. pp 2111-2116.
- Burgemeister R (2011) Laser capture micro dissection of FFPE tissuesections bridging the gap between microscopy and molecular analysis.Methods Mol Biol 724: 105-15.
- Bevilacqua C, Makhzami S, Helbling JC, Defrenaix P, Martin P (2010)Maintaining RNA integrity in a homogeneous population of mammaryepithelial cells isolated by Laser Capture Micro dissection. BMC CellBiol 11: 95.
- McGinley JN, Zhu Z, Jiang W, Thompson HJ (2010) Collection of epithelialcells from rodent mammary gland via laser capture microdissectionyielding high-quality RNA suitable for microarray analysis. Biol ProcedOnline 12: 31-43.
- Daniels KM, Choudhary RK, Evock-Clover CM, et al (2011) Comparitivetranscriptome analysis of laser micro dissected cells from bovinemammary gland.
- Casey T, Dover H, Liesman J, DeVries L, Kiupel M, et al. (2011)Transcriptome analysis of epithelial and stromal contributions tomammogenesis in three week prepartum cows. PLoS One 6(7): e22541.
- Velayudhan BT, Mcgilliard ML, Jiang H, et al (2013) Gene expressionprofile in prepubertal bovine mammary parenchyma and epithelialcells in response to ovariectomy. j Vet Anim Sci 44: 8-14.
- Choudhary RK, Daniels KM, Evock-Clover CM, Garrett W, Capuco AV, etal. (2010) Technical note: A rapid method for 5-bromo-2’-deoxyuridine(BrdU) immunostaining in bovine mammary cryosections that retainsRNA quality. J Dairy Sci 93(6): 2574-9.
- Choudhary RK, Li RW, Evock-Clover CM, Capuco AV (2013) Comparisonof the transcriptomes of long-term label retaining-cells and controlcells microdissected from mammary epithelium: an initial study tocharacterize potential stem/progenitor cells. Front Oncol 3: 21.
- Brenaut P, Lefèvre L, Rau A, Laloe D, Pisoni G, et al. (2014) Contributionof mammary epithelial cells to the immune response during earlystages of a bacterial infection to Staphylococcus aureus. Vet Res 45(1):16.
- Cánovas A, Rincón G, Bevilacqua C, Islas Trejo A, Brenaut P, et al. (2014)Comparison of five different RNA sources to examine the lactatingbovine mammary gland transcriptome using RNA-Sequencing. Sci Rep4: 5297.
- Marioni JC, Mason CE, Mane SM, Stephens M, Gilad Y (2008) RNA-seq:an assessment of technical reproducibility and comparison with geneexpression arrays. Genome Res 18(9): 1509-17.
- Marioni JC, Mason CE, Mane SM, Stephens M, Gilad Y (2008) RNA-seq:an assessment of technical reproducibility and comparison with geneexpression arrays. Genome Res 18(9): 1509-17.
- Iyer EPR, Iyer SC, Sulkowski MJ, Cox DN (2009) Isolation andpurification of Drosophil peripheral neurons by magnetic bead sorting.J Vis Exp 34: e1599.
- Huynh HT, Robitaille G, Turner JD (1991) Establishment of bovinemammary epithelial cells (MAC-T): an in vitro model for bovinelactation. Exp Cell Res 197(2): 191-9.
- Kawahara Y, Kwak S, Sun H, Ito K, Hashida H, et al. (2003) Humanspinal motoneurons express low relative abundance of GluR2 mRNA:an implication for excitotoxicity in ALS. J Neurochem 85(3): 680-689.
- Elstner M, Morris CM, Heim K, Lichtner P, Bender A, et al. (2009) Singlecellexpression profiling of dopaminergic neurons combined withassociation analysis identifies pyridoxal kinase as Parkinson’s diseasegene. Ann Neurol 66(6): 792-8.
- Brown AL, Smith DW (2009) Improved RNA preservation forimmunolabeling and laser microdissection. RNA 15(12): 2364-74.
- Iyer EPR, Cox DN (2010) Laser capture microdissection of Drosophilaperipheral neurons. J Vis Exp 39: 2016.
- Hideyama T, Yamashita T, Suzuki T, Tsuji S, Higuchi M, et al. (2010)Induced loss of ADAR2 engenders slow death of motor neurons fromQ/R site-unedited GluR2. J Neurosci 30(36): 11917-25.
- Elstner M, Morris CM, Heim K, Bender A, Mehta D, et al. (2011)Expression analysis of dopaminergic neurons in Parkinson’s diseaseand aging links transcriptional dysregulation of energy metabolism tocell death. Acta Neuropathol 122(1): 75-86.
- Hideyama T, Teramoto S, Hachiga K, Yamashita T, Kwak S (2012) Cooccurrenceof TDP-43 mislocalization with reduced activity of an RNAediting enzyme, ADAR2, in aged mouse motor neurons. PLoS One 7(8):e43469.
- Tagliafierro L, Bonawitz K, Glenn OC, Chiba-Falek O (2016) GeneExpression Analysis of Neurons and Astrocytes Isolated by LaserCapture Microdissection from Frozen Human Brain Tissues. Front MolNeurosci 9:72.
- Murakami H, Liotta L, Star RA (2000) IF-LCM: laser capture microdissection of immuno fluorescently defined cells for mRNA analysisrapid communication. Kidney Int 58(3): 1346-53.
- Smolinski D, Blessenohl M, Neubauer C, Kalies K, Gebert A (2006)Validation of a Novel Ultra-Short Immuno labeling Method for High-Quality mRNA Preservation in Laser Microdissection and Real-TimeReverse Transcriptase-Polymerase Chain Reaction. J Mol Diagnostics8(2): 246-253.
- Kase M, Houtani T, Sakuma S, Tsutsumi T, Sugimoto T, et al. (2007)Laser micro dissection combined with immune-histo chemistry onserial thin tissue sections: a method allowing efficient mRNA analysis.Histochem Cell Biol 127:215-9.
- Gründemann J, Schlaudraff F, Haeckel O, Liss B (2008) Elevatedalpha-synuclein mRNA levels in individual UV-laser-micro dissected dopaminergic substantia nigra neurons in idiopathic Parkinson’sdisease. Nucleic Acids Res 36: e38 .
- Hoelzinger DB, Mariani L, Weis J, Woyke T, Berens TJ (2005) Geneexpression profile of glioblastoma multiforme invasive phenotypepoints to new therapeutic targets. Neoplasia 7(1): 7-16.
- Li A, Walling J, Ahn S, Kotliarav Y, Su Q, et al. (2009) Unsupervisedanalysis of transcriptomic profiles reveals six glioma subtypes. CancerRes 69(5): 2091-9.
- Kislin KL, McDonough WS, Eschbacher JM, et al (2009) NHERF-1:modulator of glioblastoma cell migration and invasion. Neoplasia11(4): 377-87.
- Nevo I, Woolard K, Cam M, et al (2014) Identification of molecularpathways facilitating glioma cell invasion in situ. PLoS One 9:e111783
- Bolis A, Coviello S, Bussini S, et al (2005) Loss of Mtmr2 phosphatasein Schwann cells but not in motor neurons causes Charcot-Marie-Toothtype 4B1 neuropathy with myelin outfoldings. J Neurosci 25(37):8567-77.
- Bender A, Krishnan KJ, Morris CM, Taylor GA, Reeve AK, et al. (2006)High levels of mitochondrial DNA deletions in substantia nigra neuronsin aging and Parkinson disease. Nat Genet 38: 515-7.
- Boone DR, Sell SL, Hellmich HL (2013) Laser capture micro dissectionof enriched populations of neurons or single neurons for geneexpression analysis after traumatic brain injury. J Vis Exp (74): 50308.
- Bandyopadhyay U, Cotney J, Nagy M, et al (2013) RNA-Seq profiling ofspinal cord motor neurons frBandyom a presymptomatic SOD1 ALSmouse. PLoS One 8:e53575.
- Dalmas DA, Scicchitano MS, Chen Y, Kane J, Mirabile R, et al. (2008)Transcriptional profiling of laser capture micro dissected rat arterialelements: fenoldopam-induced vascular toxicity as a model system.Toxicol Pathol 36(3): 496-519.
- Dalmas DA, Scicchitano MS, Chen Y, Kane J, Mirabile R, et al. (2008)Transcriptional profiling of laser capture micro dissected rat arterialelements: fenoldopam-induced vascular toxicity as a model system.Toxicol Pathol 36(3): 496-519.
- Huang ZS, Malek JY, Bhasin M (2009) Selective isolation of veingraft endothelial and smooth muscle cells using laser capture microdissection for microarray gene analysis. FASEB J 23:LB322-.
- Greenberg SA, Salajegheh M, Judge DP, Feldman MW, Kunci RW, et al(2012) Etiology of limb girdle muscular dystrophy 1D/1E determinedby laser capture microdissection proteomics. Ann Neurol 71(1): 141-5.
- Ikeda A, Kai H, Kajimoto H, Yasuoka S, Kage Masayoshi, et al (2012)Selective gene expression analysis of muscular and vascularcomponents in hearts using laser microdissection method. Int J VascMed 2012: 863410.
- Vanderburg CR, Clarke MS (2013) Laser capture micro dissection ofmetachromatically stained skeletal muscle allows quantification offiber type specific gene expression. Mol Cell Biochem 375(1-2): 159-70.
- Kwapiszewska G, Wilhelm J, Wolff S, Laumanns I, Koenig IR, et al.(2005) Expression profiling of laser-micro dissected intrapulmonaryarteries in hypoxia-induced pulmonary hypertension. Respir Res 6(1):109.
- Fink L, Kwapiszewska G, Wilhelm J, Bohle RM (2006) Lasermicrodissectionfor cell type- and compartment-specific analyses ongenomic and proteomic level. Exp Toxicol Pathol 57 (Suppl 2): 25-9.
- Yick CY, Zwinderman AH, Kunst PW, GrunbergK, Mauad T, et al. (2014)Gene expression profiling of laser microdissected airway smoothmuscle tissue in asthma and atopy. Allergy 69(9): 1233-40.
- Wild P, Knuechel R, Dietmaier W, et al (2000) Laser micro dissectionand microsatellite analyses of breast cancer reveal a high degree oftumor heterogeneity. Pathobiology 68:180-90.
- Stingl J, Caldas C (2007) Molecular heterogeneity of breast carcinomasand the cancer stem cell hypothesis. Nat Rev Cancer 7: 791-9..
- Polyak K (2011) Heterogeneity in breast cancer. J Clin Invest 121(10):3786-8.
- Ellsworth RE, Seebach J, Field LA, Heckman C, Kane J, et al. (2009) Agene expression signature that defines breast cancer metastases. ClinExp Metastasis 26(3): 205-13.
- Braakman RB, Luider TM, Martens JW, Foekens JA, Umar A (2011) Lasercapture micro dissection applications in breast cancer proteomics.Methods Mol Biol 755:143-54.
- Bertos NR, Park M (2016) Laser Capture Micro dissection as a Tool toStudy Tumor Stroma. Methods Mol Biol 1458: 13-25.
- Zhang Y, Liu Y, Ye Y, et al (2016b) Quantitative proteome analysis ofcolorectal cancer-related differential proteins. J Cancer Res Clin Oncol.
- Yao H, Zhang Z, Xiao Z, Chen Yongheng, Li C, et al (2009) Identificationof metastasis associated proteins in human lung squamous carcinomausing two-dimensional difference gel electrophoresis and laser capturemicro dissection. Lung Cancer 65(1): 41-8.
- Kato Y, Nakamura H, Tojo H, Nomura M, Nagao T (2015) A proteomicprofiling of laser-micro dissected lung adenocarcinoma cells of earlylepidic-types. Clin Transl Med 4: 24.
- Zhang L, Lanzoni G, Battarra M, Inverardi L, Zhang Q, et al. (2016a)Proteomic profiling of human islets collected from frozen pancreatausing laser capture microdissection. J Proteomics 150: 149-159.
- De Marchi T, Braakman RB, Stingl C, Van Duijn MM, Smid M, et al. (2016)The advantage of laser-capture micro dissection over whole tissueanalysis in proteomic profiling studies. Proteomics 16(10): 1474-85.
- Longuespée R, Alberts D, Pottier C, Smargiasso N, Mazzucchelli G, et al.(2016) A laser micro dissection-based workflow for FFPE tissue microproteomics: Important considerations for small sample processing.Methods 104:154-62.
- Bracke N, Van Poucke M, Baert B, Wynendaele E, Bels LD, et al. (2014)Identification of a microscopically selected microorganism in milksamples. J Dairy Sci 97(2): 609-15.
- Olsen GJ, Lane DJ, Giovannoni SJ, Pace NR, Stahl DA (1986) Microbialecology and evolution: a ribosomal RNA approach. Annu Rev Microbiol40: 337-65.
- Schmidt TM, Relman DA (1994) Phylogenetic identification ofuncultured pathogens using ribosomal RNA sequences. MethodsEnzymol 235: 205-22.
- Klitgaard K, Mølbak L, Jensen TK, Lindboe CF, Boye M (2005) Lasercapture micro dissection of bacterial cells targeted by fluorescence insitu hybridization. Biotechniques 39(6): 864-8.
- Vannucci FA, Foster DN, Gebhart CJ (2013) Laser micro dissectioncoupled with RNA-seq analysis of porcine enterocytes infected withan obligate intracellular pathogen (Lawsonia intracellularis). BMCGenomics 14:421.
- Lutz HL, Marra NJ, Grewe F, Carlson JS, Palinauskas V, et al. (2016)Laser capture micro dissection microscopy and genome sequencingof the avian malaria parasite, Plasmodium relictum. Parasitol Res115(12): 4503-4510.
- Burbach GJ, Dehn D, Nagel B, Turco DD, Deller T (2004) Lasermicrodissection of immunolabeled astrocytes allows quantification ofastrocytic gene expression. J Neurosci Methods 138: 141-8.






























