Molecular Identification of Phyllanthus amarus Schumach. & Thonn. Phyllanthaceae
GO Alade1*, N Iyede2, KM Okpoji2 and KK Ajibesin1
1Department of Pharmacognosy & Herbal Medicine, Niger Delta University, Wilberforce Island, Nigeria
2Department of Pharmacognosy, Madonna University, Nigeria
Submission: March 11, 2023; Published: April 24, 2023
*Corresponding author: GO Alade, Department of Pharmacognosy & Herbal Medicine, Niger Delta University, Wilberforce Island, Nigeria, Email ID: aladegideon@yahoo.com;gideonalade@ndu.edu.ng
How to cite this article: GO Alade, N Iyede, KM Okpoji and KK Ajibesin. Molecular Identification of Phyllanthus amarus Schumach. & Thonn. Phyllanthaceae. Glob J Pharmaceu Sci. 2023; 10(4): 555793. DOI: 10.19080/GJPPS.2023.10.555793.
Abstract
The upswing in the exploitation of plants for prevention and treatment of diseases in human and animal cannot be overstressed. This upsurge has drawn the attention of the users, health care professionals and policy holders in health care delivery to the safety and therapeutic efficacy of these products. Every issue that has to do with these is initiated by identifying the plant(s) used as the raw material correctly. The aim of the present study is to identify the leaf of Phyllanthus amarus Schumach. & Thonn. Phyllanthaceae using DNA marker by employing standard method. Phyllanthus amarus is a very useful plant made into various preparations such as decoction, infusion and maceration, among others and used folklorically to treat miscellaneous ailments such as jaundice, hepatitis, dysentery, dropsy, diarrhoea, fevers, inflammation, urinary issues, diabetes, gonorrhoea, kidney ailments, measles, influenza and tuberculosis among many others.
Keywords: DNA; Safety; Identity; Efficacy; Barcoding
Introduction
Errors have been detected in using only morphological characters for plant identification [1-5]. The use of DNA-based technique, therefore, would provide proper authentication for discrimination of plant species [6]. It is a new trend for quick technique of identifying and evaluating organisms, including plants, animals and microorganisms based on the sequence of the DNA obtained from a small fragment of the individual organism [7]. This method also helps in detecting any species that are possibly novel [8-9], it is also a potential tool for detecting any error often faced in identifying species since this method often combines similarity-based method of DNA barcoding and morphology to solve taxonomical challenges and any stage plant’s growth can be employed [10-12]. The method is based on comparism of DNA sequence with established reference databases [13]. Phyllanthus amarus has its origin in the tropical parts of America and widely distributed throughout the tropical and subtropical regions globally, occurring as a weed and used for gonorrhea, jaundice, diabetes, stomachache, malaria, skin rashes, itches and dysentery [14] in ethnomedicine. Numerous biological activities have been carried out on the plant, some of which are anti-inflammatory [15], antioxidant [16], antinociceptive [17], immunostimulant [18], antiviral [19], antibacterial [20], antihyperglycemic [21], anti-diarrheal [22], gastro protective and antiulcer activity [23], contraceptive [24] among many others. Phyllanthin, hypohyllanthin, nirtetralin, nitanthin and phytetralin are some isolated chemical constituents of the plant [25]. The study is aimed at identifying P. amarus at molecularlevel.
Methodology
Collection of plant
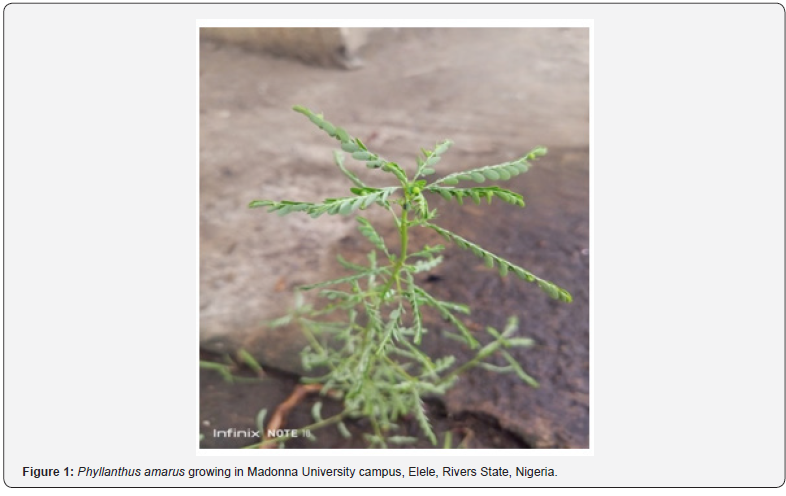
The leaves of the P. amarus (Figure 1) were harvested from the premise of Madonna University campus, Elele, Rivers State, Nigeria on the 10th of May 2022 after it has been properly identified at site by Dr. A. T. Oladele of the Department of Forestry and Wildlife Sciences, University of Port Harcourt, Rivers State, Nigeria. Mr, Frelix I. Nwafor of Department of Pharmacognosy, Faculty of Pharmacy, University of Nigeria, Nsukka authenticated the collected plant samples where a voucher specimen with voucher number PCG/UNN/0436 was thereafter deposited.

Molecular Identification
The extraction of the deoxyribonucleic acid (DNA) of P. amarus leaf sample was effected by employing Zymo Research (ZR) fungal/ bacterial deoxyribonucleic acid (DNA) Mini-Prep™ isolation/ recovery kit obtained from ‘Inqaba’, South Africa and carried out using the method described by Alade et al. [26]. It was thereafter quantified and subjected to Maturase K region amplication with the use of PCR [26]. Chromatography of the amplified-product was run by using Gel electrophoresis technique with agarose-gel (1%) as the stationary phase at 120V for 15 minutes. It was finally visualized and removed in UV transilluminator light. ZR fungal/ bacterial DNA mini prep recovery kit was then used to recover the DNA for sequencing, which was carried out at Inqaba, South Africa. The resulting amino acid sequences of the DNA were engineered (edited) by employing the ‘bioinformatics algorithm Trace edit’ in which sequences that were related or showed similarities were downloaded from the ‘National Center for Biotechnology Information’ (NCBI) data-base by blasting the DNA sequence and aligning [27]. This was used to synthesize evolution trend of the P. amarus in the Phyllanthaceae family.
Result
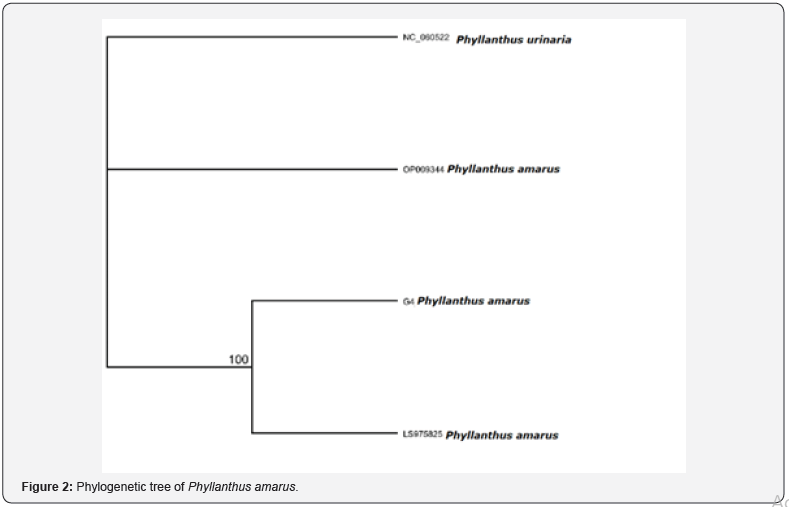
The phylogenetic tree of P. amarus is as shown in Figure 2.
Discussion
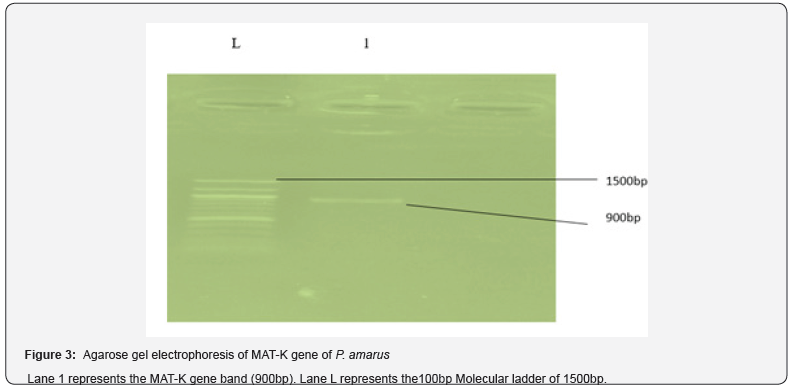
The success of amplication of mat k gene was deduced from the 900 bp obtained for the extracted DNA of the plant (Figure 3). Issues of concern or challenges arising from Plant taxonomy are mostly tacked with the use of mat-K gene complex [8]. A size of 800 – 900 base pair indicates successful amplication which in turns shows uncontaminated DNA. Maturase K genes are employed for the study of ‘sequence variations’ and ‘evolutionary trends’ at the level of the genus [2]. The upsurge in the use of the gene is as a result of its fast evolution at the nucleotide and analogous amino acid levels [10]. The measure of similarity of a plant species to an already identified one in the data base is known as its maximum identity. Accuracy of identification is specified by a maximum similarity that is not less than 95%. One hundred percent (100%) maximum identity was observed for P. amarus (Figure 2). The use of DNA sequence data has been seen to be informative in characterizing and synthesizing phylogenetic relationship of different medicinal plant species. This trend contributed immensely to adequate plant classification [28]. Information obtained from sequenced genes is crucial in molecular systematic studies [29] Analysis of obtained DNA sequence can provide taxonomic information of medicinal plants [13]. Similarity of DNA sequence indicates that the various sequences have the same homologs [30]. DNA sequences with percentage identity above 70 means that they have ≥ 90% probability to share the same biological processes and functions. Data gathered from sequenced genes is crucial in molecular systematic studies. Analysis of sequenced DNA of medicinal plant species provides vital taxonomic data. Doubtlessly, genomic regions differ significantly in their prospective phylogenetic instructiveness and their beneficence in solving a taxonomy challenges [31].

Conclusion
The findings from this study have shown that medicinal plants and cogenerics can be differentiated with the use of DNA barcoding technique with Mat-K region which is a known DNA marker for plants. Plant characterization with DNA markers is a good method for identification of plant medicines and should be recommended for plants especially those with closely related species. This does not jettison the expertise of taxonomists but will go a long way at validating their work.
References
- Kress WJ (2017) Plant DNA barcodes: Applications today and in the future. Journal of Systematics and Evolution 55(4): 291-307.
- Li X, Yang Y, Henry RJ, Rossetto M, Wang Y, et al. (2015) Plant DNA barcoding: from gene to genome. Biol Rev Camb Philos Soc 90(1): 157-166.
- Techen N, Parveen I, Pan Z, Khan IA (2014) DNA barcoding of medicinal plant material for identification. Curr Opin Biotechnol 25: 103-110.
- Ganie SH, Upadhyay P, Das S, Sharma MS (2015) Authentication of medicinal plants by DNA markers. Plant Gene 4: 83-99.
- Zhu S, Liu Q, Qiu S, Dai J, Gao X (2022) DNA barcoding: an efficient technology to authenticate plant species of traditional Chinese medicine and recent advances. Chinese Medicine 17: 112.
- Kress WJ, Wurdack KJ, Zimmer EA, Weigt LA, Janzen DH (2005) Use of DNA barcodes to identify flowering plants. Proc Natl Acad Sci USA 102(23): 8369-8374.
- Singh HK, Parveen I, Raghuvanshi S, Babbar SB (2012) The loci recommended as universal barcodes for plants on the basis of floristic studies may not work with congeneric species as exemplified by DNA barcoding of Dendrobium species. BMC Res Notes5:
- Selvaraj D, Sarma RK, Sathishkumar R (2008) Phylogenetic analysis of chloroplast matK gene from Zingiberaceae for plant DNA barcoding. Bioinformation 3(1): 24-27.
- Ghorbani A, Saeedi Y, de Boer HJ (2017) Unidentifiable by morphology: DNA barcoding of plant material in local markets in Iran. PLoS One12(4).
- Koonin EV, Novozhilov AS (2009) Origin and evolution of the genetic code: the universal enigma. IUBMB Life 61(2): 99-111.
- Hebert PDN, Cywinska A, Ball S, de Waard J (2003) Biological identifications through DNA barcodes. Proc Biol Sci B 270 (1512): 313-322.
- Hajibabaei M, Singer GA, Hebert PD, Hickey DA (2007) DNA barcoding: how it complements taxonomy, molecular phylogenetics and population genetics. Trends Genet 23(4): 167-172.
- Koonin EV, Galperin MY (2003) Sequence - Evolution - Function: Computational Approaches in Comparative Genomics. Boston: Kluwer Academic; Chapter4, Principles and Methods of Sequence Analysis.
- Patel JR, Tripathi P, Sharma V, Chauhan NS, Dixit VK (2011) Phyllanthus amarus: ethnomedicinal uses, phytochemistry and pharmacology: a review. J Ethnopharmacol 138(2): 286-313.
- Harikrishnan H, Jantan I, Haque M, Kumolosasi E (2018)Anti-inflammatory effects of Phyllanthus amarus Schum. & Thonn. through inhibition of NF-κB, MAPK, and PI3K-Akt signaling pathways in LPS-induced human macrophages. BMC Complement Altern Med 18(1):
- Karuna R, Reddy SS, Baskar R, Saralakumari D (2009) Antioxidant potential of aqueous extract of Phyllanthus amarus in rats. Indian J Pharmacol 41(2): 64-67.
- Adedapo AA, Ofuegbe SO (2015) Anti-inflammatory and antinociceptive activities of the aqueous leaf extract of Phyllanthus amarus Schum (Euphorbiaceae) in some laboratory animals. J Basic Clin Physiol Pharmacol 26(1): 89-94.
- Muthulakshmi M, Subramani PA, Michael RD (2016) Immunostimulatory effect of the aqueous leaf extract of Phyllanthus niruri on the specific and nonspecific immune responses of Oreochromis mossambicus Iran J Vet Res 17(3): 200-202.
- Yeh SF, Hong CY, Huang YL, Liu TY, Choo KB, et al. (1993) Effect of an extract from Phyllanthus amarus on hepatitis B surface antigen gene expression in human hepatoma cells. Antiviral Res 20(3): 185-192.
- Ribeiro AMB, de Sousa JN, Costa LM, Oliveira FA, dos Santos RC, et al. (2019) Antimicrobial activity of Phyllanthus amarus & Thonn and inhibition of the NorA efflux pump of Staphylococcus aureus by Phyllanthin. Microb Pathog 130: 242-246.
- Okoli CO, Obidike IC, Ezike AC, Akah PA, Salawu OA (2011) Studies on the possible mechanisms of antidiabetic activity of extract of aerial parts of Phyllanthus niruri. Pharm Biol 49(3): 248-255.
- Odetola AA, Akojenu SM (2000) Anti-diarrhoeal and gastro-intestinal potentials of the aqueous extract of Phyllanthus amarus (Euphorbiaceae). Afr J Med Med Sci 29(2): 119-122.
- Mostofa R, Ahmed S, Begum MM, Sohanur RM, Begum T, et al. (2017) Evaluation of anti-inflammatory and gastric anti-ulcer activity of Phyllanthus niruri L. (Euphorbiaceae) leaves in experimental rats. BMC Complement Altern Med 17(1): 267.
- Rao MV, Alice KM (2001) Contraceptive effects of Phyllanthus amarus in female mice. Phytother Res 15(3): 265-267.
- Wu W, Li Y, Jiao Z, Zhang L, Wang X, et al. (2019) Phyllanthin and hypophyllanthin from Phyllanthus amarus ameliorates immune-inflammatory response in ovalbumin-induced asthma: role of IgE, Nrf2, iNOs, TNF-α, and IL's. Immunopharmacol Immunotoxicol 41(1): 55-67.
- Alade GO, Olamiju O, Ajibesin KK (2022) Molecular identification of Peperomia pellucida (L.) Kunth Piperaceae.International Journal of Science and Technology Research Archive 2(2): 107-111.
- Wheeler DL, Barrett T, Benson DA, Bryant SH, Canese K, et al. (2006) Database resources of the National Center for Biotechnology Information. Nucleic Acids Res 1: 34.
- Hao DC, Xiao PG (2020) Pharmaceutical resource discovery from traditional medicinal plants: Pharmacophylogeny and pharmacophylogenomics. Chin Herb Med 12(2): 104-117.
- Brown TA (2002) Genomes. 2nd Oxford: Wiley-Liss Chapter 16, Molecular Phylogenetics.
- Frazer KA, Elnitski L, Church DM, Dubchak I, Hardison RC (2003) Cross-species sequence comparisons: a review of methods and available resources. Genome Res 13(1): 1-12.
- Neves SS, Forrest LL (2011) Plant DNA sequencing for phylogenetic analyses: from plants to sequences. Methods Mol Biol 781: 183-235.






























