The Danger of Doing Power Calculations Using Only Descriptive Statistics
Steve Su*
Covance Pty Ltd, North Ryde, Australia
Submission: October 25, 2017; Published: March 12, 2018
*Corresponding author: Steve Su, Covance Pty Ltd, North Ryde, Australia, Email: allegro.su@gmail.com
How to cite this article:Steve Su. The Danger of Doing Power Calculations Using Only Descriptive Statistics. Biostat Biometrics Open Acc J. 2018; 5(4): 555670. DOI: 10.19080/BBOAJ.2018.05.555670
Keywords
Keywords:Normality; Power calculations; Clinical trial statistician; Generalised lambda distributions
Opinion
Power calculations are bread and butter in the daily life of a clinical trial statistician and necessary for the determination of sample sizes in clinical trials. However, the use of descriptive statistics and assumption of Normality in conducting power calculations, while prevalent in practice, is not necessarily correct. This article highlights the potential problems of traditional approach to power calculations and suggests some possible practical solutions. Power calculations are routinely done by statisticians to determine sample sizes in clinical trials and many sample size calculation software are available. However, most, if not all software rely on traditional statistical techniques, which may result in totally inappropriate sample size for a given situation.
As an illustration, consider the following parameters for power calculations involving a two-sample independent T-test, assuming equal variance:
o Mean for treatment 1: 3.7954545
o Mean for treatment 2: 3.8954545
o Standard deviation for both treatment groups is:0.2705155
o 90% power, 5% significance level
The standard power calculation will give n=155 per group. Obviously, the underlying assumption is that the data is symmetric and the average values represents the peaks of the distribution as shown in Figure 1a. However, if the true underlying distributions are actually skewed as shown in Figure 1b, then it is an open question as to whether the power calculation will still hold. The data for treatment groups were generated by FKML Generalized Lambda Distributions (GLDs) via GLDEX package in R with parameters λ1 = 3,λ2 = 2,λ3 = 10,λ4 = 3 for treatment 1 and λ1 = 3.1,λ2 = 2,λ3 = 10,λ4 = 3 for treatment 2 [1-4].
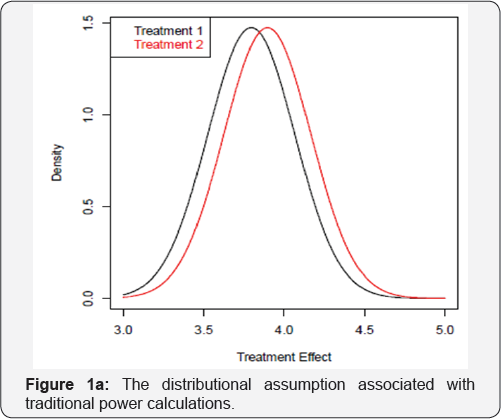
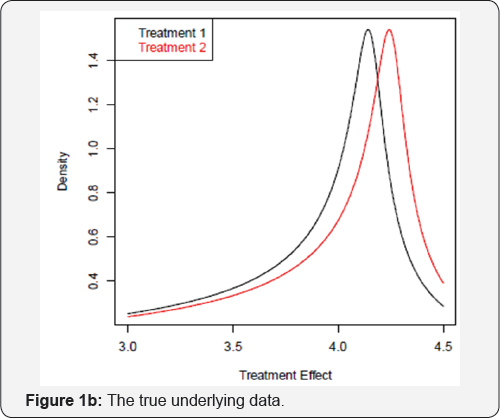
Rechecking the sample size of n=155 per group based on these distributions using simulations, the power is only approximately 40%, which is nowhere close to the desired level of 90%. To achieve approximate 90% power, a sample size of around 600 per group is needed instead. This highlights the problem with sample size calculations when the underlying distributional assumption is not met. Alternatively, a comparison of two treatment arms under Figure 1b using the mode (or the peak of the distributions) could be considered as a valid alternative. By simulation, using a sample size of approximately 350 per group, approximately 90% of the time we can declare that the effect of treatment 2 is greater than treatment 1 by comparing the modes of the distributions. The traditional approach to power calculation in this example would yield sample size with too little power. Yet if the true underlying distribution is modelled and sample size is re-estimated based on t-test, 600 per group is still quite large. A better approach is to consider comparing the modes of the distributions which will cut down the sample size to 350 per group, a 42% reduction which will translate into significant savings in terms of cost and time. To be able to obtain a sample size of 350, there are several technical issues that must be overcome.
It is imperative to be able to fit distribution to the underlying data first, and flexible distributions such as GLDs can be used for this purpose. The R package GLDEX contains a number of algorithms (maximum likelihood estimation, L moment matching and others) to fit GLDs to empirical data as well as goodness of fit statistics and graphics [2]. Once a sufficiently good distribution for each treatment arm is found, the trial statistician and clinician can then decide a parameter of interest for comparison and obtain sample size required for a given power via simulation from the fitted distribution. In the light of skewed data, the parameter of interest could be the modes of the distributions rather than the averages, as modes represent the most common values experienced by subjects among the treatment arms [3].
A further practical problem is that very often there may not be any data available other than the descriptive statistics, and in this case, it may be desirable to simulate from distributions with various degrees of skewness but similar mean and variance to assess the impact on statistical power with respect to sample sizes. An alternative is to run a pilot study to allow an initial assessment of the treatment effect, which can then be used to determine the final number of patients needed [4]. The main message is that statisticians should not be content with just a simple sample size calculation based on descriptive statistics, care and additional checks should always be carried out to ensure the sample size obtained is reasonably robust and that the trial has a reasonable chance of success.
References
- Freimer M, G Kollia, G Mudholkar, C Lin (1988) A study of the generalized Tukey lambda family. Commun Stat Theory Methods.
- Su S (2007) Fitting Single and Mixture of Generalized Lambda Distributions to Data via Discretized and Maximum Likelihood Methods: GLDEX in R. Journal of Statistical Software 21(9): 1-17.
- Su S (2007) Numerical maximum log likelihood estimation for generalized lambda distributions. Comput Stat Data Anal 51(8): 3983-3998.
- Su S (2010) Fitting GLD to data Using the GLDEX 1.0.4 in R. Handbook of Distribution Fitting Methods. Boca Raton, CRC Press/Taylor & Francis pp.585-608.






























