Hierarchical Classification F Directed Graph Nodes and Application to Protein networks
Tsitsiashvili G Sh1*, Bulgakov VP2and Losev AS1
1Institute for Applied Mathematics FEB RAS, Russia
2Institute of Biology and Soil Science FEB RAS, Russia
Submission: March 15, 2017; Published: May 08, 2017
*Corresponding author: Tsitsiashvili G Sh, Institute for Applied Mathematics FEB RAS, Russia, Spain, Email: guram@iam.dvo.ru
How to cite this article: Tsitsiashvili G S, Bulgakov V, Losev A. Hierarchical Classification F Directed Graph Nodes and Application to Protein networks. Biostat Biometrics Open Acc J. 2017;1(4): 555567. DOI: 10.19080/BBOAJ.2017.01.555567
Abstract
Algorithm of hierarchical classification for nodes of directed graph is constructed and is applied to sub network of secondary metabolism in Arabidopsis protein network
Main Algorithm
In [1,2] an algorithm of directed graph nodes dividing into cyclically equivalent classes (clusters) is constructed. A cyclical equivalence of two nodes is an existence of a cycle containing these nodes in the directed graph. A partial order on a set of all clusters is defined and a sequential algorithm of zero-one matrix characterizing this partial order is suggested. However, this algorithm does not allow to analyze detailed a structure of single cluster which may be sufficiently complicated. Such analysis is important in some problems of the bioengineering. For example in the Arabidopsis protein network sub networks of secondary metabolism, light, hormonal, and signal sub networks are defined in [3]. Another approach is a hierarchical classification of undirected weighed graph G .
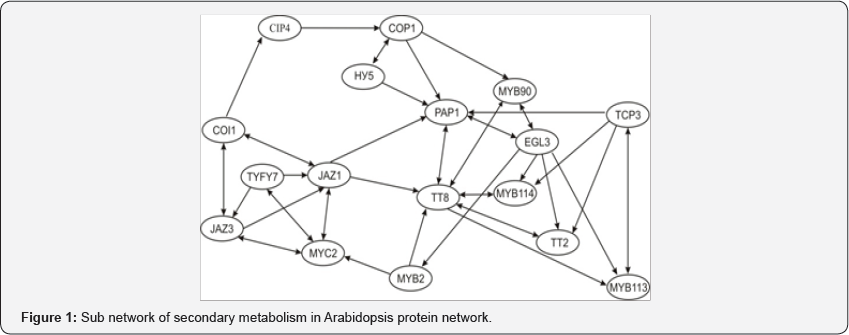
This approach [4] is based on a choice of some critical value а and on a transformation of the weighed graph G into unweighed graph Ga using a following rule. If a weight of some edge sЄG is smaller than critical value а then the edge s remains in the graph Ga else it does not remain in Ga . After this transformation, it is possible to find connectivity components of the un weighed and undirected graph Ga. These components create clusters in the nodes set of the graph G To find connectivity components in the graph Ga . it is convenient to use following algorithm. Assume that at step 1 we have information about node 1. Then at this step there is single cluster {1} Suppose that at step t we have clusters k1,...,kpt, and there are edges which connect directly new node (t+1) with some nodes in clusters k1,...,kqt,0 ≤ qt ≤ pt (if qt=0 then node (t+1) is not connected directly with nodes of clusters k1,...,kpt,). As a result at step t+1 we obtain new set of clusters: 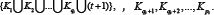
Successfully decreasing critical value, а we divide earlier obtained clusters into sub clusters and so on. This algorithm allows constructing the classification of undirected weighed graph G nodes. In this paper, an algorithm of the hierarchical classification of nodes in the cluster G is based on a concept of cycles with minimal length. Each pair of nodes in the graph G is connected by some cycle. So it is possible to define minimal length of cycles connected this pair of nodes. Minimal length of cycle connecting nodes 1,2 equals a sum of minimal path length from node 1to node 2and minimal path length from node 2 to node1. Well-known Deikstra or Floid algorithms [5] may calculate minimal path lengths. As a result, directed graph G is transformed into complete undirected and weighed graph. Then it is possible to apply procedure of its nodes hierarchical classification. This algorithm allows constructing the classification of undirected weighed graph S nodes. This algorithm was applied to make a hierarchical classification of plants depending on a presence or on an absence of some organic matters in them [6].
Numerical Experiment and Its Interpretation
Consider the sub network of secondary metabolism in the protein Arabidopsis network [3] (Figure 1). Direct test shows that this sub network is a directed graph with cyclically equivalent nodes. Using suggested algorithm calculate constants defining hierarchical classification on steps 1, 2, 3: a1= 4, a2 = 6, a3= 8. . On the step 3, this procedure ends. In Figure 2, we may see how considered sub network of the secondary metabolism is divided into classes of circles mainly with the length four. It is worthy to say that the proteins JAZ1 is an inhibitor and the protein TT8 is an activator, which effects on the proteins MYB113, MYB114 that make the secondary metabolism products. Therefore, the inhibitor JAZ1 decelerates a work of the activator TT8 and consequently decreases an amount of the secondary metabolism products.

References
- Tsitsiashvili G Sh (2013) Sequential algorithms of graph nodes factorization. Reliability: Theory & Applications 8(4): 30-33.
- Tsitsiashvili G Sh, Osipova MA, Losev AS (2016) Graphs clusterization algorithms. Vestnik of Voronej state university, pp. 145-149.
- Bulgakov VP, Avramenko TV, Tsitsiashvili G Sh (2016) Critical analysis of protein signalling networks involved in the regulation of plant secondary metabolism: focus on anthocyanins. Crit Rev Biotechnol 24: 1-6.
- Tsitsiashvili G Sh, Osipova MA (2015) Construction of hierarchical classification by similarity matrix. Reliability: Theory and Applications 10(3): 16-19.
- Cormen TH, Leiserson CE, Rivest RL (1990) Introduction to Algorithms (1st edn.). MIT Press and McGraw-Hill.
- Tarbeeva DV, Fedoreyev SA, Mironova LN, Tsitsiashvili G Sh (2015) A comparative study on polyphenolic compounds composition of i. pseusa-corus and its cell culture. Korea-Russia Seminar on Science and Technology, Khabarovsk, Russia, pp. 19-20.






























