Determination of Antibiotic Susceptibility Pattern of Clinical Isolates of Salmonella Typhi and Escherichia coli
Farouk Nas S1, Musa Diso AA2, Idris IS3 and Muhammad Ali4*
1Department of Biological Science, Bayero University, Nigeria
2Department of Science Laboratory Technology, School of Technology, Nigeria
3Department of Pharmaceutical Technology, School of Technology, Nigeria
4Microbiology Department, Kano University of Science and Technology, Nigeria
Submission: June 21, 2018; Published: August 08, 2018
*Corresponding author: Muhammad Ali, Microbiology Department, Kano University of Science and Technology Wudil, Kano, Nigeria, Tel: 2347032967252, Email: alimuhd4real@gmail.com
How to cite this article: Farouk Nas S, Musa Diso AA, Idris IS, Muhammad Ali. Determination of Antibiotic Susceptibility Pattern of Clinical Isolates of Salmonella typhi and Escherichia coli. Ann Rev Resear. 2018; 2(5): 555596. DOI: 10.19080/ARR.2018.02.555596
Abstract
The aim of the study is to determine the antibiotic sensitivity pattern of clinical isolates of Salmonella typhi and Escherichia coli. The isolates were obtained from stool sample of typhoid fever patients attending Muhammad Abdullahi wase Specialist Hospital Kano. Identification of the bacterial isolates was done using conventional standard laboratory methods including Gram staining, cultural characterization and biochemical test. The sensitivity pattern of Salmonella typhi and Escherichia coli was tested against commercially prepared antibiotics sensitivity disc using Kirby-Bauer method. Based on the finding of this study, most of the antibiotics are active against the isolates. Salmonella typhi was susceptible to Travid, Reflacin, Ciprofloxacin, Streptomycin and Gentamicin but resistance to Ampicillin. However, its sensitivity to Septrin Augmentin, Ceporex and Nalidixic acid is intermediate. On the other hand, Escherichia coli were found resistant to Augmentin, Ceporex and Ampicillin while sensitive to Reflacin, Taravid, Septri and Streptomycin while Escherichia coli were found intermediate sensitive to Ciprofloxacin and Gentamicin. Statistical analysis of the result showed that there is no significant different on the susceptibility of the organism to the antibiotics used at p<0.05.
Keywords: Antibiotics; Bacteria; Characterization; Escherichia coli and Salmonella typhi
Introduction
Presently, antimicrobial resistant among microorganism such as bacteria, virus, parasites and other disease-causing organisms is one of the serious threats to the management of infectious disease globally [1]. The increase in antibiotic resistance has been attributed to a combination of microbial characteristics, the selective pressure of antibiotic use and social and technical changes that enhance the transmission of resistant organisms. The growing threat from resistant organisms calls for concerted action to prevent the emergence of new resistant strains and the spread of existing ones [2]. Many procedures use and misuse of antibiotics in man have resulted in antibiotic-resistant bacteria. Antimicrobial resistance is a natural biological phenomenon which often enhanced as a consequence of infectious agents’ adaptation to exposure to antimicrobial agents used in human or Agriculture and the widespread use of disinfectant at the farm and the household levels [3]. It is now accepted that antimicrobial use is the single most important factor responsible for increased antimicrobial resistance [4]. Salmonella typhi is the etiological agent of typhoid fever; it is gram negative bacteria belonging to the family Enterobacteriaceae. Salmonella typhi are aerobic, non-spore-forming and flagellated bacilli of about 2-3 μm long and 0.4-0.6 μm diameter. Members of Genus Salmonella Members of this genus have variety of pathogenic effect [5]. The strains have evolved to infect wide variety of reptiles, birds and mammals resulting in many different syndromes ranging from colonization and chronic carriage to acute fatal diseases. Differences in lipopolysaccharide (LPS) in Salmonella generate antigenic variations, which also affect the virulence of the strain [6]. S. typhi is generally transmitted via food or water contaminated with feces or urine from a case or carrier. Direct person-to-person spread can also occur. Shellfish harvested from sewage contaminated water are potential vehicles, as well as fruits and vegetables grown in soil fertilized with human waste in developing countries. Sexual transmission from an asymptomatic carrier has been documented. Laboratory-acquired infections also have been reported including laboratory workers who do not directly handle Salmonella specimens [5].
The emergence of multi drug resistance S. typhi (MDRST) has been of major concern in recent years. MDRST is defined as strains of S. typhi resistant to all three first line antibiotics for typhoid fever. The number of reported multi resistant typhoid fever increased rapidly throughout the world from 1989 onwards with most of the cases from the Africa, Middle East and Asia especially in the Indian subcontinent, Pakistan and China. Resistance to these agents is associated with the plasmid present in the bacteria [7]. Escherichia coli are the most commonly encountered member of the family Enterobacteriaceae in the normal colonic flora and the most common cause of opportunistic infections [8]. All members of the family Enterobacteriaceae are facultative, all ferment glucose and reduce nitrates to nitrites and all are oxidase negative [9]. Escherichia coli typically colonize the gastrointestinal tract of human infants within a few hours after birth. Usually, E. coli and its human host coexist in good health and with mutual benefit for decades. These commensal E. coli strains rarely cause disease except in immune compromised hosts or where the normal gastrointestinal barriers are breached as in peritonitis, for example. The niche of commensal E. coli is the mucous layer of the mammalian colon. The bacterium is a highly successful competitor at this crowded site, comprising the most abundant facultative anaerobe of the human intestinal micro flora. Despite the enormous body of literature on the genetics and physiology of this species, the mechanisms whereby E. coli assures this auspicious symbiosis in the colon are poorly characterized. One interesting hypothesis suggests that E. coli might exploit its ability to utilize gluconate in the colon more efficiently than other resident species, thereby allowing it to occupy a highly specific metabolic niche. The aim of the study is to determine the antibiotic sensitivity pattern of clinical isolate of Salmonella typhi and Escherichia coli from stool sample of patients attending Muhammad Abdullahi waste Specialist Hospital, Kano State.
Materials and Methods
Ethical Approval
The study was conducted following ethical approval obtained from the Health Services Management Board, Kano State based on the consent of Ethical Committee of Muhammad Abdullahi waste Specialist Hospital Kano State.
Test Isolates
Clinical isolate of Salmonella typhi and Escherichia coli were used in this study. The isolates were obtained from Microbiology laboratory of Abdullahi waste Specialist, Kano. The bacteria were isolated from stool sample of typhoid fever patient attending the Hospital.
Isolation and Identification of Isolates
The clinical stool swab samples from infected patients were inoculated onto Nutrient agar (Life save Biotech, USA) and MacConkey agar (Life save Biotech, USA) plates and incubated aerobically at 370 C for 24 hours. After incubation bacterial growth was observed for colony appearance and morphology. Each colony was re-inoculated into freshly prepared agar plates until a pure colony was obtained. For identification, each pure colony was Gram stained and subjected to further biochemical tests Chessbrough [10]. Results were interpreted according to the guidelines of the Clinical and Laboratory Standards Institute [11].
Gram Staining
A drop of normal saline was placed on a well labeled clean grease-free glass slide using a sterile inoculating loop; a colony of an overnight culture of the bacterial isolate was emulsified with the normal saline to make a thin smear. The smear was air dried and then heat fixed. The slide was flooded with crystal violet (primary stain) for 30 seconds after which the stain was rinsed from the slide with water. The smear was flooded with Lugol’s iodine (mordant) to fix the primary stain. The iodine was rinsed with water after 60 seconds. The slide was then flooded with a decolorizer (acetone) and rinsed off almost immediately. The counter stain; safranin was added and left for 30 seconds before being rinsed off. The stained smear was air dried, and then observed under the microscope using X100 oil immersion objective lens of the microscope Sherman [12].
ABiochemical Characterization
The isolates were also characterized by biochemical tests viz. IMViC reactions i.e. Indole test, Methyl Red test, Vogues Proskauer test catalase test, oxidase test and Citrate utilization test, as well as Lactose and Mannitol fermentation reaction test by standard method given by Sherman & Holt [12,13].
Indole test
Tryptophan broth was inoculated with an isolate of the test organism and incubated at 370C for 24 hours. About 0.5 ml of Kovack’s reagents was added to the broth culture [10].
Methyl red test
MR-VP broth was inoculated with an isolate of the test organism using sterile inoculating loop and incubated at 370C for 24 hours. About 5 drops of Methyl-red reagent was added to the broth culture [10].
Voges Proskauer
MR-VP broth was inoculated with an isolate of the test organism using sterile inoculating loop and incubated at 370C for 24 hours. Six millilitres (6ml) of 5% alpha naphthol was added followed by 0.2 ml of KOH. The tube was shaken gently and remained undisturbed for 5min [10].
Citrate utilization test
Simmons’s citrate agar was streaked back and forth with inoculums of the test organism and incubated aerobically at 370C for 24 hours [10].
Catalase test
Few colonies of the test organisms were emulsified in distilled water on a clean slide. 1-2 drops of 3% H2O2 were added and cover with cover slip [13].
Nitrate reduction test
Nitrate broth was inoculated with an isolate of the test organism using sterile inoculating loop and incubated at 370C for 24 hours. A dropper full of sulfonic acid and that of α- naphthylamine were added to the broth [13].
Motility test
A semi solid medium in a test tube was inoculated with an isolate of the test organism using straight sterile wire and making a single stab at the center of the test tube. The test tube was incubated at 370C and examined the stab line at various intervals to determine motility [13].
Antibiotic Sensitivity Test
The bacteria isolates (Salmonella typhi and Escherichia coli) were subjected to antibiotic susceptibility testing using the agar disc diffusion method as described by Bauer, et al. [14] Mueller Hilton agar (MHA) plates were inoculated with overnight culture of each isolate by streak plating method. The standard antibiotic sensitivity discs were then aseptically placed at equidistance on the plates and allowed to stand for 1 hour. The plates were then incubated at 37°C for 24 hours. Sensitivity pattern of the isolates to Augmentin (30 μg/disc), Taravid (10 μg/disc), Streptomycin (30 μg/disc), Ampicillin (30 μg/disc), Gentamicin (20 μg/disc), Reflacin (10 μg/disc), Ceporex (30 μg/disc), Nalixidic acid (20 μg/disc), Ciprofloxacin (10 μg/disc) and Septrin (30 μg/disc), produced by Abtek pharmaceutical limited, were determined. Isolates were divided into three groups based on the zone of inhibition produced by the antibiotic disc; susceptible, intermediately susceptible and resistant according to the Clinical and Laboratory Standards Institute (CLSI) guideline; Performance Standards for Antimicrobial Susceptibility Testing [11]. The experiment was conducted in triplicate and the average value was calculated.
Statistical Analysis
The data of average zone of inhibition produced by the isolates against the antibiotics used was analyzed using paired T-test. Significance level for the differences was set at p<0.05.
Result
Cultural and Biochemical Characteristics of the Isolates
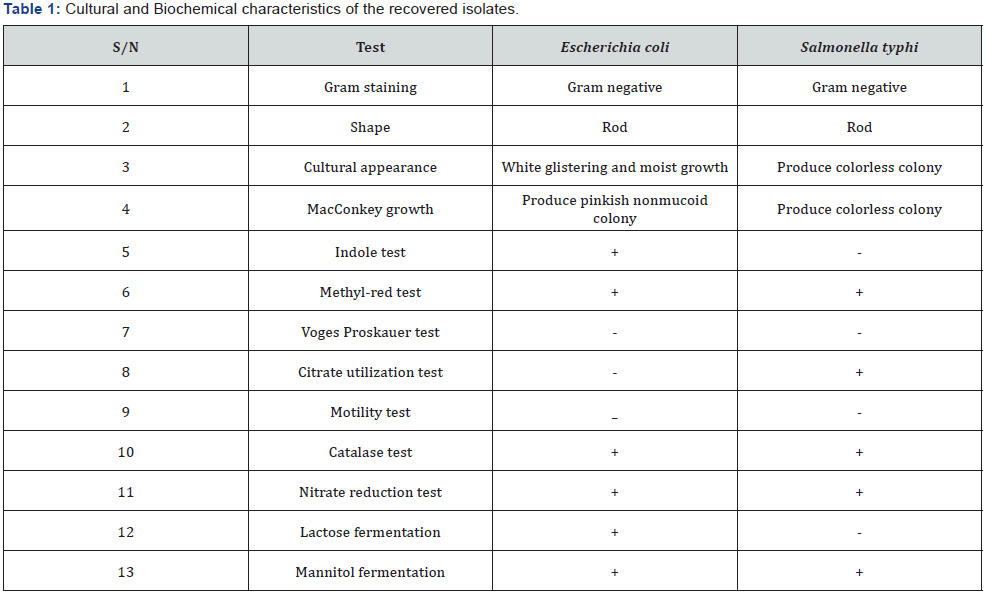
The cultural and morphological characteristics of the isolates are presented in Table 1. The result shows that E. coli was found to be Gram negative rod with White glistering and moist growth appearance on Nutrient agar plate and pinkish non-mucoid colony in MacConkey agar plate while S. typhi was found to be Gram negative rod which produce colorless colony on Nutrient and MacConkey agar plates. The isolates were characterized based on indole, methyl-red, Vougues Proskeaur, citrate utilization, catalase and motility tests. Nitrate reduction test, lactose and mannitol fermentation were also conducted.
Antibiotic Sensitivity Test
The zone of inhibition of antibiotic sensitivity disc against Salmonella typhi and Escherichia coli isolates is presented in Table 2. Most of the antibiotics are active against the isolates with zone of inhibition greater than 18 mm.

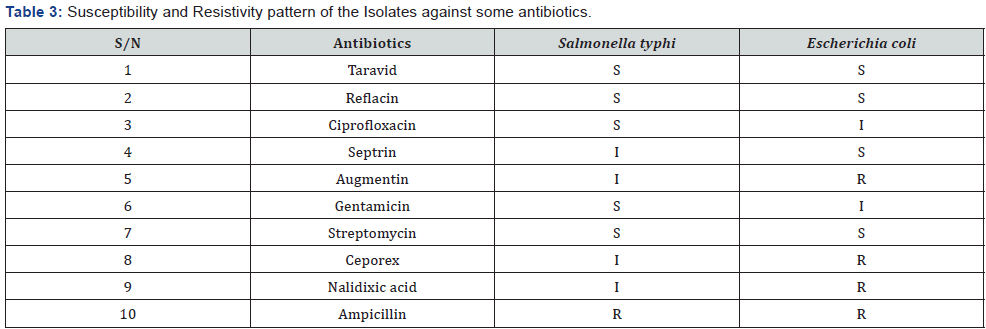
Susceptibility and Resistivity Status of the Isolates
The sensitivity pattern of the isolates against the antibiotics used is presented in Table 3. Isolates were divided into three groups based on the zone of inhibition produced by the antibiotic disc; Susceptible S (above 18mm), intermediately susceptible I (11-17mm) and Resistant R (below 11mm).
Discussion
Antimicrobial resistance has increased rapidly in the recent years in both developing and developed countries and it has rapidly becomes a leading health concern [15]. In the presence study, sensitivity pattern of Salmonella typhi and Escherichia coli was tested against commercially prepared antibiotics. Based on the finding of this study, most of the antibiotics are active against the isolates. Salmonella typhi was susceptible to Travid, Reflacin, Ciprofloxacin, Streptomycin and Gentamicin but resistance to Ampicillin. However, its sensitivity to Septrin Augmentin, Ceporex and Nalidixic acid is intermediate. The result of this study was in conformity with that of Shehu, et al who found Salmonella typhi susceptible Ciprofloxacin, Erythromycin, Gentamicin and Streptomycin but resistant to Rifampicin and Ampicillin [16]. In another work, all the Salmonella typhi isolates were less sensitive to ciprofloxacin but no resistance was seen, whereas 76% of the same isolates showed resistant to nalidixic acid. In this work, it was shown that nalidixic acid resistant isolates had decreased susceptibility to ciprofloxacin [17]. Similar work was carried out by John, et al. [18] which reveals same pattern of result as in the above. Changing trends of antibiogram of S. typhi and S. paratyphi A says that 67% of S. typhi in the study were sensitive to chloramphenicol and the sensitivity of S. typhi isolates to cephalosporin was found to have increased from 2001-2004 [19]. Resistance to amoxicillin, chloramphenicol, ampicillin and cotrimoxazole were significantly high in a study conducted in most part of the world [20]. Plasmid mediated multi-drug resistance to ampicillin, chloramphenicol and cotrimoxazole has been described in various parts of Asia [21].
On the other hand, Escherichia coli were found resistant to Augmentin, Ceporex and Ampicillin while sensitive to Reflacin, Taravid, Septri and Streptomycin. According to the present study, Escherichia coli were found intermediate sensitive to Ciprofloxacin and Gentamicin. The result of this study supported the result of a study conducted by Dessalegn et al. [22]. They found that E. coli were resistant to ampicillin (100%) but susceptible to ciprofloxacin. Within the community, Escherichia coli strains are commonly susceptible to all agents active against the Enterobacteriaceae. However, because of the frequent occurrence of R plasmids, strains acquired in hospitals may be resistant to any combination of potentially effective antibiotics and therapy must therefore guided by susceptibility testing [8]. Based on the finding of this study, Salmonella typhi were found to be more sensitive to the antibiotics used covering an average zone of inhibition of 17.9 mm while Escherichia coli has an average zone of inhibition of 15.8 mm. Statistical analysis of the result revealed that the table value (p value at p < 0.05) is greater than the calculated value for paired t-test; therefore, there is no significant different on the susceptibility of the organism to the antibiotics used hence null hypothesis is accepted (Figure 1).
Conclusion
The study was aimed to determine the antibiotic sensitivity pattern of clinical isolate of Salmonella typhi and Escherichia coli against some selected antibiotics. Salmonella typhi was susceptible to Travid, Reflacin, Ciprofloxacin, Streptomycin and Gentamicin but resistance to Ampicillin. However, its sensitivity to Septrin Augmentin, Ceporex and Nalidixic acid is intermediate. On the other hand, Escherichia coli were found resistant to Augmentin, Ceporex and Ampicillin while sensitive to Reflacin, Taravid, Septri and Streptomycin. According to the present study, Escherichia coli were found intermediate sensitive to Ciprofloxacin and Gentamicin.
References
- Umaru IJ, Badruddin FA, Assim ZB, Umaru HA (2018) Antibacterial and Cytotoxic Actions of Chloroform Crude Extract of Leptadenia hastata (Pers) Decnee. Clin Med Biochemistry 4(1): 1-4.
- Thiel M, Gutow, L (2005) The ecology of rafting in the marine environment. Oceanography and Marine Biology: An Annual Review 42: 181-263.
- Tsou CH, Mori SA (2002) Seed coat anatomy and its relationship to seed dispersal in subfamily Lecythiodoideae of the Lecythideae (The Brazil Nut Family). Botanical Bulletin of Academia Sinica 43: 37-56.
- Cannon JG, Burton RA, Wood SG, Owen NL (2004) Naturally occurring fish poisons from plants. Journal of chemical education 81(10): 1457- 1461.
- Meyer BN, Ferrigni, NR, Putnam, JE, Jacobson, LB, Nichols, DE, et al. (1982) Brine shrimp: A convenient general bioassay for active plant constituents. Planta Med 45: 31-34.
- Pundir RK, Jain P (2010) Comparative studies on the antimicrobial activity of black pepper (Piper nigrum) and turmeric (Curcuma longa) extracts. International Journal of Applied Biology and Pharmaceutical Technology 1: 491-501.
- Mahesh B, Satish S (2008) Antimicrobial activity of some important medicinal plant against plant and human pathogens. World Journal of Agricultural Sciences 4: 839-843.
- Vandepitte J, Engback K, Piot P, Heuck CC (1995) Basic Microbiology Procedures in Clinical Bacteriology. World Health Organization, Geneva, p. 85.






























