Antibiotic-Resistant Bacteria in Hospital Sewages and Treated Urban Wastewater: A Case Study from Marrakech (Morocco)
Fouzia El-Ogri1,2,3, Naaila Ouazzani2, Fatima Boraâm3, Nabila Soraa4 and Laila Mandi1,2*
1National Center for Research and Study on Water and Energy (CNEREE), Cadi Ayyad University, Morocco
2Laboratory of Hydrobiology, Faculty of Sciences Semlalia, Morocco
3Regional Laboratory of Epidemiology and Environmental Health, Delegation of Health, Morocco
4Laboratory of bacteriology-virology, Cadi Ayyad University, Morocco
Submission: January 29, 2019; Published: May 10, 2019
*Corresponding author:Laila Mandi, National Center for Research and Study on Water and Energy, Cadi Ayyad University, Marrakech, Morocco
How to cite this article: Fouzia El-Ogri, Naaila Ouazzani, Fatima Boraâm, Nabila Soraa, Laila Mandi. Antibiotic-Resistant Bacteria in Hospital Sewages and Treated Urban Wastewater: A Case Study from Marrakech (Morocco). Adv Res Gastroentero Hepatol. 2019; 12(5): 555850. DOI: 10.19080/ARGH.2019.12.555850.
Abstract
This study investigated the evaluation of the emergence of antibiotic resistance in bacteria isolated from untreated hospital wastewaters and their transmission to the receiving Wastewater Treatment Plant (WWTP) of Marrakech city. Water sampling was conducted from January 2015 to June 2015 at fourteen different points (each department of the hospital, main effluent outlet of the hospital, WWTP (inlet, secondary and tertiary treatment)). The study showed that hospital sewages and wastewater collected at the inlet and after secondary treatment by Marrakech WWTP were loaded with pathogenic bacteria such as Escherichia coli, Klebsiella pneumoniae, Salmonella spp, Aeromonas hydrophyla, Pseudomonas aeruginosa, Acinetobacter baumanii, Enterococcus faecalis, Enterococcus faecium and Staphylococcus aureus. In this study we isolated 285 strains: 227 from the sewers of CHU hospital effluents and 58 strains collected at the entrance and after secondary treatment at Marrakech WWTP, they were loaded with potential pathogenic bacteria such as E. coli (n=52), K. pneumoniae (n=29), A. hydrophyla (n=28), P. aeruginosa (n=16), A. baumanii (n=17), E. faecalis (n=23), E. faecium (n=7) and S. aureus (n=8). Most of the isolated strains (63.3%, n=180) were resistant to all the tested antibiotic families: beta-lactams (carbanims n=3; cephalosporins n=6; penicillins n=10; monobactams n=1), fluoroquinolones (n=3) and aminosides (n=4), except for the P. aeruginosa strain which was resistant to beta-lactam antibiotics, especially imipenem and ceftazidime. The majority of these bacterial strains were known to be involved in nosocomial infections. On the other hand, there is an absence of these antibiotic-resistant bacteria after advanced tertiary treatment of these effluents at the Marrakech WWTP. This means that advanced tertiary treatment plays an important role in removing pathogenic strains and preserving the environment. A proper use of antibiotics updated by regular programs to monitor the susceptibility of bacteria to antibiotics remains one of the key tools for reducing the magnitude of the bacterial resistance rising phenomenon. Therefore, the impact of hospital effluents on the diffusion of antibiotic resistance in the environment needs to be carefully considered.
Keywords: Antibiotic resistance; Pathogenic bacteria; Hospital effluents; WWTP effluents; Antibiotic resistant bacteria; Food industry; Water consumption; Bacteria; Sewage treatment plants; Infection; Antimicrobial therapy; Escherichia coli; Klebsiella pneumoniae; Salmonella spp; Aeromonas hydrophyla; Pseudomonas aeruginosa; Acinetobacter baumanii; Enterococcus faecalis; Enterococcus faecium
Abbrevations: ARB: Antibiotic Resistant Bacteria; WWTP: Sewage Treatment Plants; CLSI: Clinical and Laboratory Standard Institute; SFM: Society of Microbiology; EUCAST: European Committee on Antimicrobial Susceptibility Testing; FQs: Fluoroquinolones; KTG: Kanamycin, Tobramycin and Gentamycin
Introduction
The emergence and spread of infection-causing Antibiotic Resistant Bacteria (ARB) is considered as a major threat to human health due to excessive and abusive use of antibiotics [1] and their release into municipal wastewater following their elimination [2]. They can therefore reach surface water and groundwater through different routes, such as sewage treatment plants (WWTP) effluent (as they are not completely eliminated), runoff water or water infiltration used for agricultural purposes. Antibiotics pose a significant risk to human health and the environment, even at low concentrations [3]. Among the multiple reservoirs and pathways of antibiotic resistance dissemination, hospital liquid waste collectors, municipal wastewater, and sewage treatment plants play an important role [4]. In addition, the excessive and abusive use of antibiotics has led to the emergence of ARB, which undermines the effectiveness of antimicrobial therapy because infectious organisms become resistant to most antibiotics [5]. Food of animal origin can also be a source of ARB due to the intensive use of antimicrobials in the food industry in many parts of the world [6].
Various studies have been carried out to evaluate the presence of these emerging pollutants in effluent discharges from WWTPs [7-9], and the presence of a number of antibiotics and the prevalence of certain antibiotic-resistant genes selected from environmental media [8].
In African countries, the liquid discharge produced by hospitals has been not yet considered at the same importance as the internal environment of the hospital. There are only few studies on hospital effluents and their behavior, in particular the risk of ARB.
In Morocco, most the studies that have been conducted to evaluate bio-resistance to antibiotics in bacteria were done particularly on the hospital environment (soil, air, surfaces, materials and tap water) and patients [10,11]. However, at our knowledge, very few studies have been done on this field directly on hospital effluents or in WWTPs [12].
The purpose of this study is to evaluate the level of antibiotic resistance of potential pathogenic bacteria isolated both from the hospital wastewater and the WWTP of Marrakech city (Morocco) at different treatment stages.
Materials and Methods
Study sites and sampling points
Our study was carried out in two sites located in the city of Marrakech: The Mohamed VI University Hospital Center (CHU) and the WWTP of the city. The first site of the CHU has different specialties with a capacity of 409 beds and generates several types of liquid wastes (Biological fluids « urine, feces, vomit, blood, sputum, water basins, etc. », chemical effluents « obsolete chemicals and liquids such as acids, bases, various reagents, solvents, etc.», effluents from the radiology department loaded with developers, fixers, and contrast products, disinfectants generated by the service of medical devices sterilization, detergents and disinfectants from cleaning and building maintenance).
The water consumption is estimated at 425 m3/d at a rate of 1.04 m3/d/bed, with a flow rate ranging from 1.63 to 12.67 m3/h [13]. The WWTP is located northwest of the city, at 7.4 km from the CHU hospital with a capacity equivalent to 1.3 inhabitants and an average daily flow of 100 000 m3/d. The WWTP is constructed in such a way as to ensure a tree-stage treatment of urban wastewaters: The first stage concerns pretreatment (by screening, desoldering and degreasing) and primary treatment (primary settling), the second stage concerns the secondary treatment (activated sludge) and the third stage is dedicated to advanced tertiary treatment (coagulation, flocculation, sand filtration and ultraviolet “UV” disinfection).
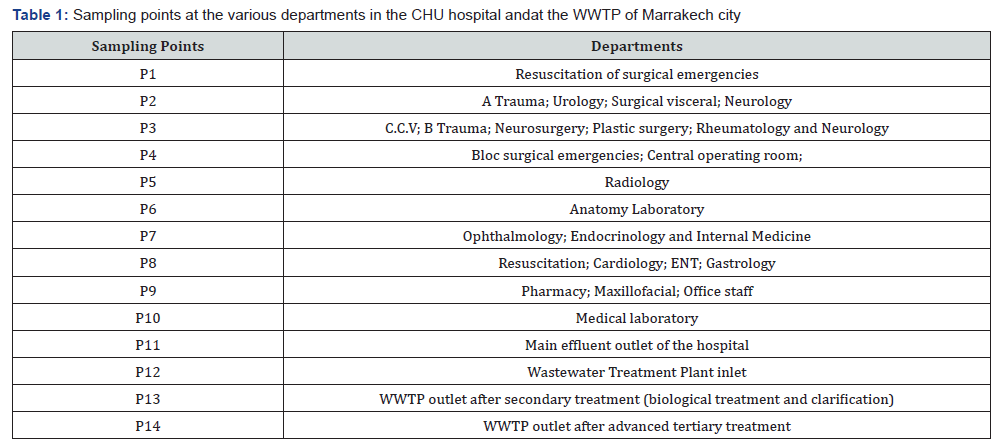
Water sampling was conducted from January 2015 to June 2015, at fourteen different points: P1 to P10 at each department of the hospital, P11 at the main effluent outlet of the hospital, which is discharged into the main sewerage network of the urban wastewater of Marrakech. The last three sampling points were done in Marrakech WWTP: P12 at the inlet of the system, P13 after the secondary treatment by activated sludge and P14 after the advanced tertiary treatment by coagulation, flocculation, followed by filtration on sand beds and then UV disinfection (Table 1). The choice of the WWTP is justified by the re-use of the treated water in the irrigation of green spaces, golf courses and the palm grove of the OULJA domain in Marrakech city.
Methods of analysis
Isolation and identification of bacteria Enterobacteriaceae Escherichia coli: E. coli detection was done according to the standard method NM ISO 9308-1/NM 03.7.003 using Tergitol7 Agar. After incubation at (44±0.5) °C for (21±3) hours, yellow to orange color, yellow halos under the membrane are developed. These colonies were confirmed by the oxidase test plus an indole test.
Klebsiella pneumoniae: K. pneumoniae develops on Tergitol7 Agar (standard method NM ISO 9308-1/NM 03.7.003). After incubation at (36 ± 2) °C, for 24 hours, the colonies (2mm to 3mm in diameter) of k. pneumoniae are lactose-positive, convex and mucous.
Salmonella spp: According to the standard method NM ISO 19250/ NM 03.7.050, the following steps were done: a) Pre-enrichment: The buffered peptone water is seeded at room temperature with a known volume of the sample or its dilutions and incubated at (36±2) °C for (18 ± 2) hours. Larger volumes can be concentrated by membrane filtration, after which the filter membrane is transferred to the buffered peptone water. b) Enrichment: Rappaport-Vassiliadis soy bean medium (RVS broth) and Muller-Kauffmann tetrathionate-novobiocin broth (MKTTn) are inoculated with the pre-enrichement culture. RVS broth is incubated at (41.5±1) °C for (24±3) hours and MKTTn broth at (36±2) °C for (24±3) hours. To detect Salmonella spp. with slow growth, incubate the enrichment broth for an additional (24±3) hours for a total incubation period of (48±4) hours at (41.5±1.0) °C. c) Isolation on selective growth medium: From the cultures obtained in enrichment, two solid selective medias are inoculated: lysine xylose deoxycholate agar (XLD agar) and Hechtoen agar. After incubation of both culture media at (36±2) °C and examine after (24±3) hours. Black-centered red colonies appear on XLD agar and black-centered green, which becomes completely black at the end of incubation on Hektoen agar.
Non enterobacteriaceae Pseudomonas aeruginosa: Standard method NM ISO 16266 was applied using cetrimide agar medium. After incubation at (41.5±0.5) °C for 48 hours, colonies (0.8 to 2.2 mm in diameter) showing flat appearance with blue or blue-green pigmentation surrounding the colonies light outer rims and brownish to greenish black-centers were selected and isolated as P. aeruginosa. These colonies were confirmed by three confirmation tests: Detection of oxidase, Pyocyanin production on King B medium, growth at 42°C in Tryptone-soy Broth.
Acinetobacter baumanii: Clinical and Laboratory Standard Institute (CLSI), using MacConkey Agar. After incubation at 30°C, colonies (2 to 3 mm in diameter) develop in 24 hours on MacConkey medium, they are convex, shiny and whitish. Aeromonas hydrophyla: The strains of A. hydrophila develop on nutrient Agar. After incubation at (36±2) °C, colonies develop in 24 hours; they are round, curved, smooth, regular and opaque.
Cocci
Staphylococcus aureus: Using Chapman-Mannitol salt agar medium (standard method NM 03.7.036/XP T 90-412). After incubation at (36±2) °C for 48 hours, Yellow colonies on a yellow background or surrounded by a yellow halo and confirmed by a tube coagulase test to give a coagulum occupies more than three quarters of the volume initially occupied by the liquid.
Enterococcus faecalis and Enterococcus faecium: According to the standard method NM ISO 7899-2/NM 03.7.006. Slanetz and Bartley Medium (Enterococcus Agar), colonies showing a red to pink color were isolated as presumptive Enterococcus, and confirmed by typical colonies will be transferred to the membrane and colonies by means of sterile tweezers, without turning, on a bile esculin-azide agar plate this has been preheated to 44 °C. After incubation at (44±0.5) °C for 2 hours, showing a brown to black color in the surrounding medium.
Identification of bacteria by Analytical Profile (APIbiochemical) strips: The bacterial strains were identified by API biochemical system for identification kits (Marcy l’Étoile, France): Api 20E for enterobacteria, Api 20NE for non-enterobacteria, Api Staph for Staphylococcus, Api 20 Strepto for Enterococcus. The API strips consists of 20 micro-tubes containing dehydrated substrates. These tests are inoculated with a bacterial suspension that reconstitutes the media. The strips were incubated for 18–24 hours at (36±2) °C. During incubation, metabolism produces color changes that are either spontaneous or revealed by the addition of reagents. The reactions were read according to the reading table and the identification was obtained by referring to the Analytical Profile Index [14].
Antibiotic sensitivity test for bacterial isolates: The production and interpretation of the antibiogram were done according to the standards of the Antibiogram Committee of French Society of Microbiology / European Committee on Antimicrobial Susceptibility Testing (CA-SFM / EUCAST) [15]. The quality control of the antibiograms was carried out with the reference strains of Escherichia coli ATCC25922, Klebsiella pneumoniae ATCC700603, Staphylococcus aureus ATCC29213, Enterococcus faecalis ATCC29212 and Pseudomonas aeruginosa ATCC 27853. The antibiotics and their loads tested are described in Table 2.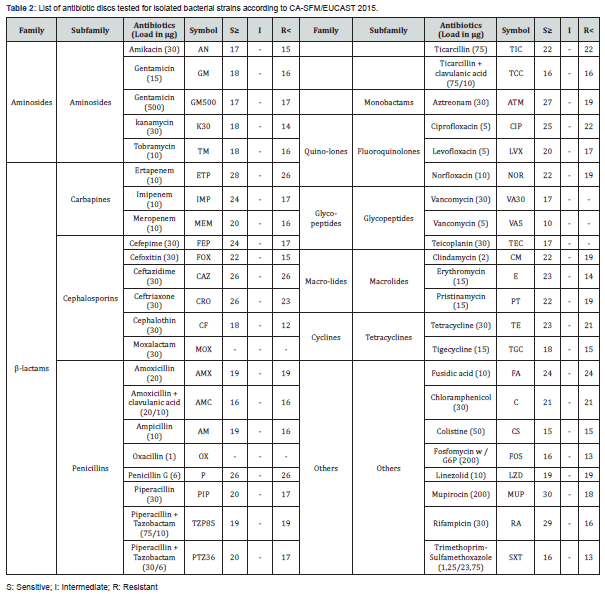
Results
Distribution of resistant bacteria
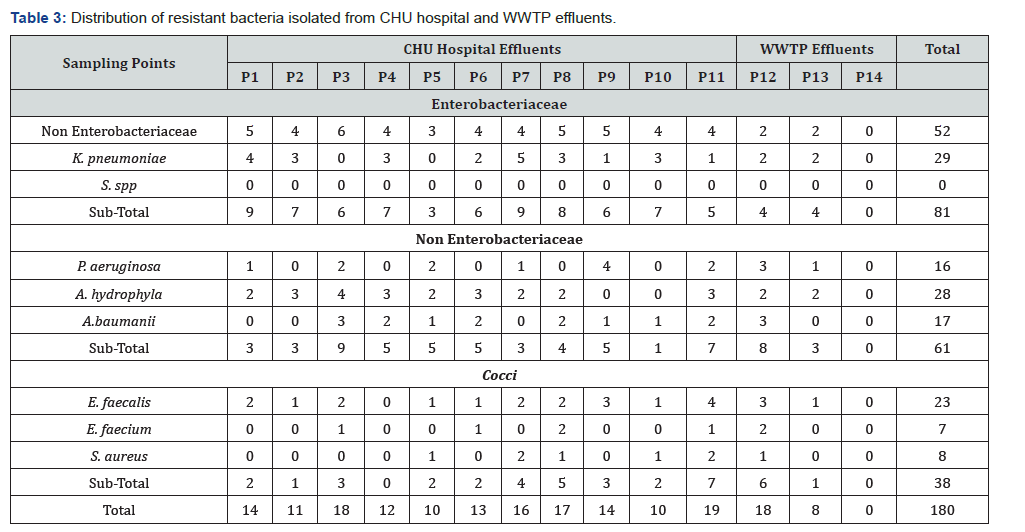
A total of 285 strains were isolated from the different sampling points distributed as follows: Escherichia coli (N = 68), Klebsiella pneumoniae (N = 37), Salmonella spp (N = 12), Pseudomonas aeruginosa (N = 30), Aeromonas hydrophyla (N = 48), Staphylococcus aureus (N = 15), Enterococcus faecalis (N = 44), and Enterococcus faecium (N = 10), of which 201 came from P1 to P10; 26 from P11 and 58 were isolated from P12, P13 and P14
According to Table 3, eight potentially pathogenic strains isolated at the P13 sampling point are resistant to antibiotics, these strains are distributed as follows: Three strains (E. coli, K.pneumonia and A. hydrophila) resistant to ciprofloxacin, two other strains (E. coli and K. pneumonaie) resistant to cephalothin, one strain of A. hydrophila resistant to cephtazidime, a strain of E. faecalis resistant to Amoxicillin + clavulanic acid and one strain of P. aeruginosa resistant to imipenem. The secondary treatment of municipal wastewater by activated sludge technology is probably insufficient and still constitutes a health risk. Two recent studies of UV-treated wastewater in a rural sewage treatment plant in southeast Louisiana (USA) confirmed the presence of antibiotic-resistant Gram-negative bacteria [16] and the presence of methicillin-resistant Staphylococcus aureus [17]. On the other hand, the present study demonstrated that at point P14, after advanced tertiary treatment (UV disinfection, preceded by coagulation-flocculation and sand filtration), pathogenic strains were absent. This means that advanced tertiary treatment plays an important role in removing pathogenic strains and preserving the environment.
Bacterial resistance and phenotypes
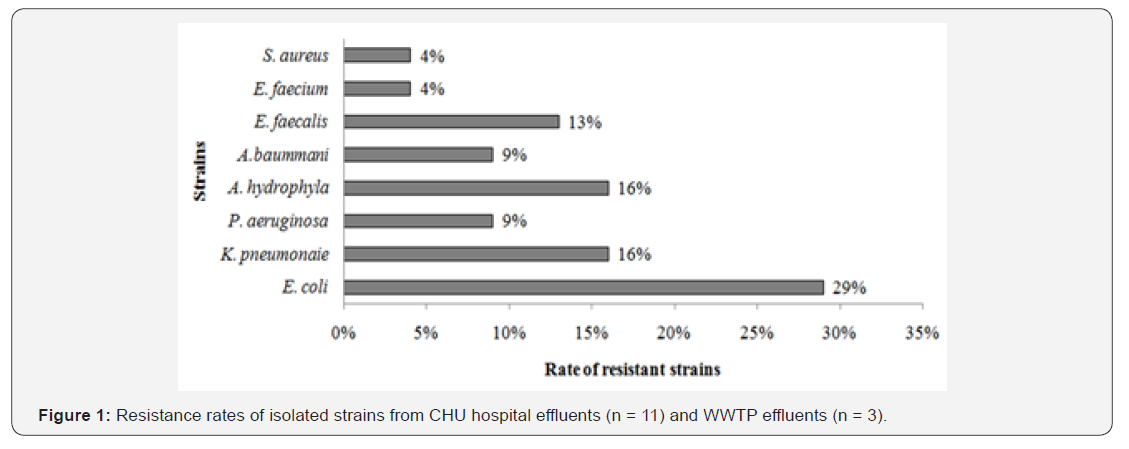
These results showed that bacterial resistance to all antibiotics is dominated by Gram-negative bacteria belonging to the Enterobacteriaceae family (E. coli, K. pneumonaie) with a rate of more than 45% followed by non-Enterobacteriaceae strains (P. aeruginosa, A. hydrophila and A. baumanii) with a rate of 34% then Cocci (E. faecalis, E. faecium and S. aureus) with a rate of 21% (Figure 1).
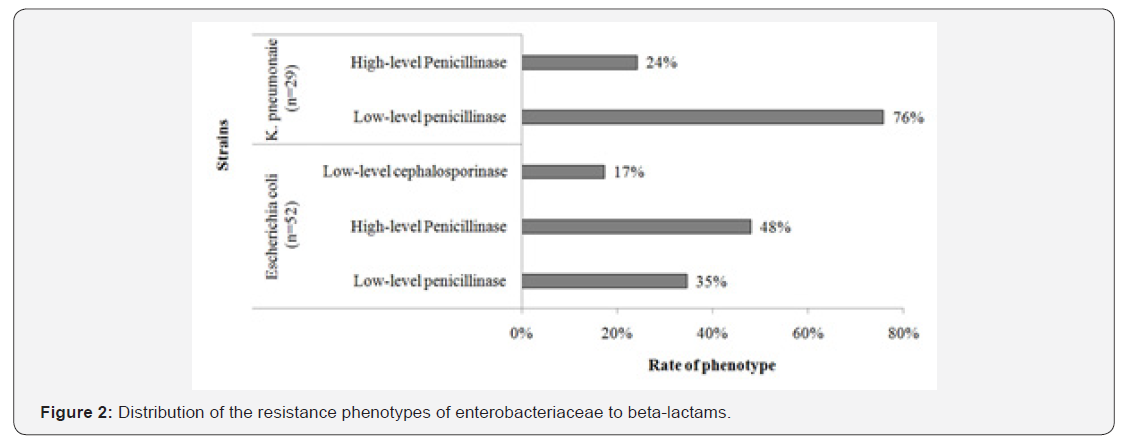
Enterobacteriaceae isolates produced 76% high penicillinases for K. pneumoniae and 48% for E. coli and were susceptible to imipenem (Figure 2).
These results are comparable to those obtained by Guessennd et al. [18] in the Ivory Coast on hospital effluents from the university hospital centre of the city of Abidjan.
Concerning the strain of Aeromonas hydrophila which has been frequently isolated from our effluents, numerous studies showed that Aeromonas strains isolated from water and sediments were resistant to cefoxitin with a rate of 80 % [19]. In the present study, Aeromonas hydrophila strains were quinolone-resistant with a rate over 90% against 41% obtained by Varela et al. [20].
The species Acinetobacter baumannii is a frequent cause of nosocomial pneumonia and other infections, such as skin and wound infections, infective endocarditis, bacteremia, urinary tract infections, and meningitis [21]. Several studies have demonstrated that A. baumannii is resistant to many of the antibiotics used in the hospital context [22,23]. All strains of A. baumanii were resistant to beta-lactams with 82.35% resistance to imipenem, 88.23% to ceftazidime and 70.60% to gentamycin (Table 4). This bacterium is distinguished by its multi-resistant character [24].
The presence of fluoroquinolones (FQs) residues in various effluents (households, hospitals and pharmaceutical effluents) is a major cause of acute and chronic toxicity, as well as the appearance of resistant bacteria [25].
Therefore, the removal of FQs residues from the environment is a crucial problem. Regarding our study and among the eight (8) strains that we have isolated, four (4) were resistant to ciprofloxacin. The strain of E. coli alone has a rate of 88.46% (Table 4) against a rate of 34.4% found by Conte et al. [26]. Ciprofloxacin is frequently prescribed to treat urinary tract infections and it is excreted unchanged in urine and feces and remains adsorbed in sludge even after treatment in WWTPs [27]. Therefore, it is a good indicator of the presence of antimicrobials in the environment [28].
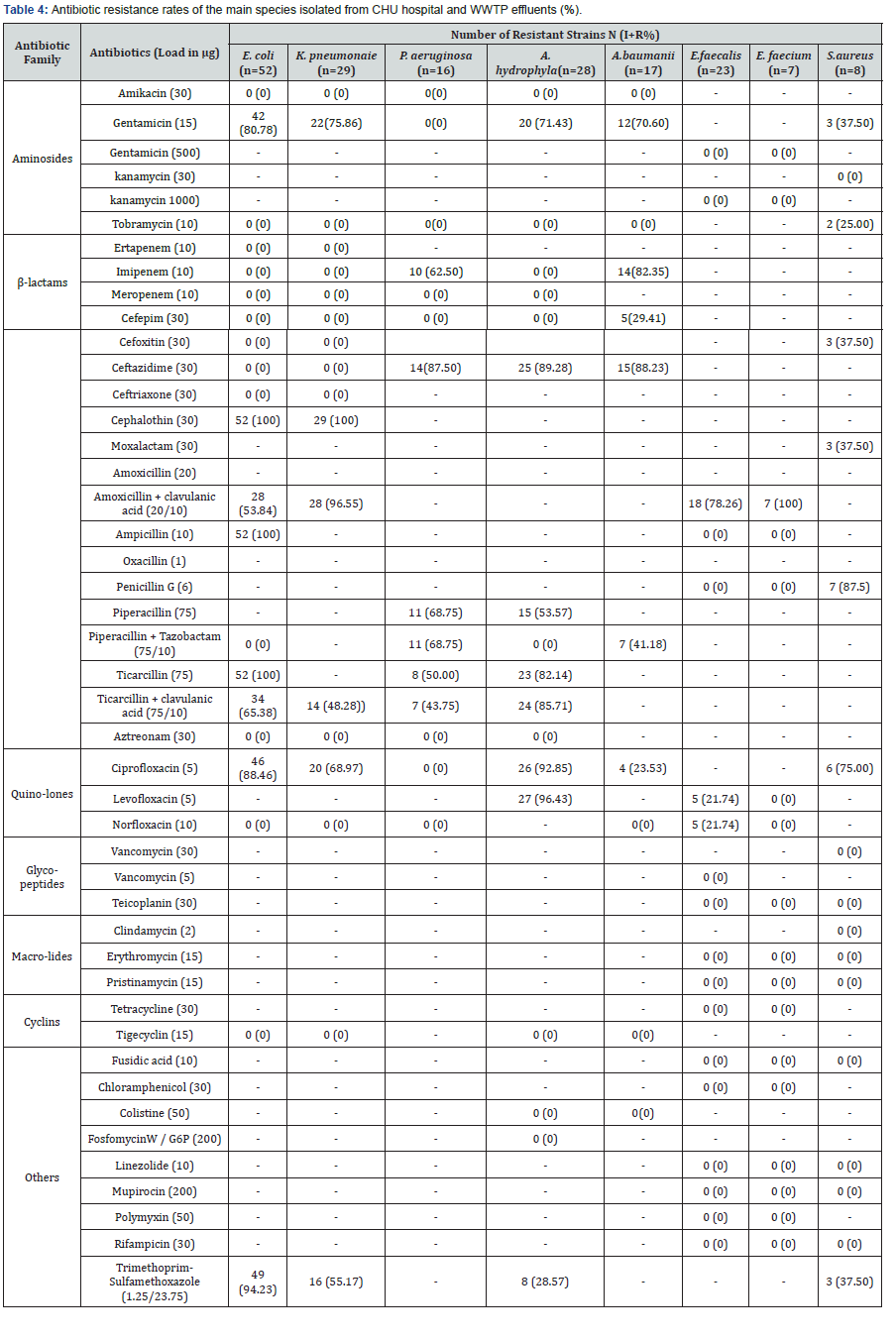
The global epidemiological situation of the resistance of E. coli strains to FQs remains variable with resistance rates of 10% in the United States and 50% in China [29]. This is probably due to the presence of emerging FQs residues in the investigated effluents which favors the acquisition of resistance of the E. coli strains and/or other strains of K. pneumoniae, A. baumanii and A. hydrophyla, which exhibits high resistance levels: 68.97%, 23.53% and 92.85% (Table 4), respectively.
The observation of this bio-resistance clearly reflects the presence of potentially pathogenic bacteria resistant to antibiotics in CHU hospital effluents and in the WWTP effluents (Raw urban wastewater and after secondary treatment) of Marrakech city. This could be explained by the selection pressure exerted by the practitioners and by the presence of low concentrations of non-metabolized antibiotics discharged from hospital effluents [30]. These different multi-resistance phenotypes could also be explained by the fact that antibiotics are among the most important molecules in Africa; and among these antibiotics, betalactams were the most frequently prescribed [31]. An 87.5% of the Pseudomonas aeruginosa were resistant to ceftazidime, 50% to ticarcillin, 43.75% to ticarcillin-clavulanic acid, 68.75% to piperacillin + tazobactam, and 62.50% to imipenem as certain strains of the hospital environment, P. aeruginosa is a ubiquitous microorganism capable of persisting in water, soil and plants, and is also responsible for a large number of common infections acquired in hospitals [32].
A study has confirmed the presence of resistant strains of P. aeruginosa in surface water bodies that received wastewaters from a hospital in Brazil [32]. All isolated Acinetobacter strains were resistant to beta-lactams and also resistant to ciprofloxacin and gentamycin with a rate of 27.3%.
The strains of Staphylococcus were recognized as having a high capacity of acquisition of resistance genes [18]. Among the aminoglycosides, the most active was gentamycin. The results obtained in this work (37.50%) are lower than those of Guessennd et al. [18] where the proportion of kanamycin, tobramycin and gentamycin (KTG) phenotype was 100% in hospital effluents. The resistance of S. aureus to many antibiotics was also observed in several studies with a rate of 90% to 95% for aminoglycosides and 90% for FQs [33]. We found a very high rate of resistance to methicillin of 37.50% compared to results obtained in hospitals 6.64% in the United States (2010), 5.97% in Canada (1999) and 0.8% in Japan (2003) [34], and in surface waters [35]. Also, the resistance rate of 75% for ciprofloxacin obtained in the present study is similar to the results obtained by Salou et al. [36] in a hospital-based study (on patients urine and pus) who demonstrated that S. aureus was resistant to ciprofloxacin 1.85 times more likely to be resistant to Methicillin resistance; methicillin resistance is also correlated with an increase in resistance to ciprofloxacin. The S. aureus resistant to methicillin is 2.54 times more likely to be resistant to ciprofloxacin. This could be explained by the emergence of resistant strains and the increase in acquired infections in hospitals resulting from the absence of good care practices and infection control guidelines.
E. faecalis and E. faecium are the most common species in hospital and urban wastewaters [37]. In this study, all strains of E. faecalis and E. faecium (Table 4) resistant to beta-lactam (amoxicillin + clavulanic acid), has very high rates of 78.26% and 100%, respectively. This may be explained by the fact that their spread in the hospital environment is due in part to their survival at high temperatures and to their resistance to chemical disinfectants such as chlorine, glutaraldehyde and alcohol [38], and in the past three decades, Enterococci has become the third most frequent nosocomial human pathogens, with a mortality rate of up to 23% [39].
Conclusion
The present study confirmed the resistance to antibiotics of the majority of bacterial strains isolated from CHU hospital sewages and the Marrakech receiving WWTP effluents (raw wastewater and even after secondary treatment). This antibiotic resistance is dominated by Gram-negative bacteria belonging to the Enterobacteriaceae family (45%) followed by non- Enterobacteriaceae strains (34%) then Cocci (21%). In addition, almost 63% of isolated bacterial strains were resistant to at least 3 families of antibiotics (beta-lactams, aminoglycosides and fluoroquinolones) except for P. aeruginosa, which revealed resistance to imipenem and ceftazidime (family of betalactams). However, the study demonstrated that the advanced tertiary treatment (coagulation flocculation + sand filtration + UV disinfection) at Marrakech WWTP removed pathogenic strains completely from the municipal wastewater. Therefore, advanced tertiary treatment is strongly recommended to ovoid dissemination of multiple resistant bacterial strains to the environment.
Acknowledgements
This work was supported by the Pole of Competences on Water and Environment (PC2E); the National Centre for Research and Studies on Water and Energy (CNEREE); the Regional Laboratory of Epidemiology and Environmental Health, Direction of Health and the Mohamed VI University Hospital Center (CHU) of Marrakech city.
References
- (2017) Antimicrobial resistance. Fact sheet N°194, World Health Organization (WHO).
- Rodriguez-Mozaz S, Weinberg HS (2010) Meeting report: Pharmaceuticals in water–An interdisciplinary approach to a public health challenge. Environ. Health Perspect 118(7): 1016-1020.
- Kümmerer K (2009) Antibiotics in the aquatic environment – A review – Part II. Chemosphere 75(4): 435-441.
- Rizzo L, Manaia CM, Merlin C, Schwartz T, Dagot C, et al. (2013) Urban wastewater treatment plants as hotspots for antibiotic resistant bacteria and genes spread into the environment: a review. Sci Total Environ 447: 345-360.
- Marti E, Variatza E, Balcázar JL (2014b) Bacteriophages as a reservoir of extended-spectrum β-lactamase and fluoroquinolone resistance genes in the environment. Clin Microbiol Infecti 20(7): 456-459.
- Ghosh A, Zurek L (2015) Antibiotic Resistance in Enterococci: A Food Safety Perspective Chapter 9. Methods and Techniques, pp. 155-180.
- Hirsch R, Ternes T, Haberer K, Kratz KL (1999) Occurrence of antibiotics in the aquatic environment. Sci Total Environ 225(1-2): 109-118.
- Li JZ, Guang-Guo Y, Liu S, Zhao JL, Yang B, et al. (2013) Occurrence and fate of eleven classes of antibiotics in two typical wastewater treatment plants in South China. Sci Total Environ 452-453: 365-376.
- Rodríguez-Mozaz S, Chamorro S, Marti E, Huerta B, Gros M, et al. (2015) Occurrence of antibiotics and antibiotic resistance genes in hospital and urban wastewaters and their impact on the receiving river. Water Res 69: 234-242.
- Bekkari H, Benchemsi N, Touijer H, Berrada S, Maniar S, et al. (2016) Microbial Analysis of Air in a Public Hospital in the City of Fez, Morocco. Int J Pharmaceuti Clin Res 8(6): 533-537.
- Bouzid J, Bouhlal A, Chahlaoui A, Aababou S, Aarab M, et al. (2015) Determining the prevalence of Surgical Site Infections of operated patients in Mohamed V hospital in Meknes. Int J Inno Sci Res 14(2): 198-207.
- Imziln B (1990) Treatment of wastewater by anaerobic and aerobic lagoons in Marrakech: Bacteriological quantitative and qualitative study; antibio-resistance of bacteria of health interest. Thèse de Doctorat de 3ème cycle Université cadi Ayyad, Faculté des Sciences Semlalia de Marrakech, pp. 122.
- El-Ogri F, Ouazzani N, Boraâm F, Mandi L (2016) A survey of wastewaters generated by a hospital in Marrakech city and their characterization. Desalin. Water Treat 57(36): 17061-17074.
- Juang DF, Morgan JM (2001) The application of the API 20 E and API rapid NET systems for the identification of bacteria from activated sludge. Electron J Biotechnol 4(1): 1-7.
- CA-SFM / EUCAST (2015) Comité d’antibiogramme de la société française de microbiologie/European Committee on Antimicrobial Susceptibility Testing.
- Naquin T, Robichaux K, Belding C, Bergeron S, Boopathy R (2017) Presence of sulfonamide and carbapenem resistance genes in a sewage treatment plant in southeast Louisiana, USA. Int Biodeterior Biodegradation 124: 10-16.
- Boopathy R (2017) Presence of Methicillin Resistant Staphylococcus aureus (MRSA) in sewage treatment plant. Bioresour Technol 240: 144-148.
- Guessennd NK, Ouattara MB, Ouattara ND, Nevry RK, Gbonon V (2013) Study of multi-resistant bacteria of hospital effluents from university hospital center (CHU) of Abidjan city (Ivory Coast). J appl Biosci 69: 5456-5464.
- Dellali M (2001) Use of microbiological and biochemical indicators in Ruditapes decussatus and Mytilus galloprovincialis in the biosurveillance of the Bizerte lagoon: Validation of certain biomarkers. Thèse de doctorat. Faculté des Sciences de Bizerte, pp: 218.
- Varela AR, Olga C, Nunes OC, Manaia CM (2016) Quinolone resistant Aeromonas spp. as carriers and potential tracers of acquired antibiotic resistance in hospital and municipal wastewater. Sci Total Environ 542: 665-671.
- Wang Q, Wang P, Yang Q (2017) Occurrence and diversity of antibiotic resistance in untreated hospital wastewater. Sci Total Environ 621: 990-999.
- Fournier PE, Richet H (2006) The epidemiology and control of Acinetobacter baumannii in health care facilities. Clin Infect Dis 42(5): 692-699.
- Hu Q, Hu Z, Li J, Tian B, Xu H, et al. (2011) Detection of OXA-type carbapenemases and integrons among carbapenem-resistant Acinetobactor baumannii in a teaching hospital in China. J Basic Microbiol 51(5): 467-472.
- Berche P, Gaillard JL, Simonet M (1988) Bactériologie: bactéries des infections humaines. Paris, Flammarion pp. 267-275.
- Prutthiwanasan B, Phechkrajang C, Suntornsuk L (2016) Fluorescent labeling of ciprofloxacin and norfloxacin and its application for residues analysis in surface water. Talanta 159: 74-79.
- Conte D, Palmeiro JK, da Silva Nogueira K, de Lima TMR, Cardoso MA (2017) Characterization of CTX-M enzymes, quinolone resistance determinants, and antimicrobial residues from hospital sewage, wastewater treatment plant, and river water. Ecotoxicol Environ Saf 136: 62-69.
- Osińska A, Harnisz M, Korzeniewska E (2016) Prevalence of plasmid-mediated multidrug resistance determinants in fluoroquinolone-resistant bacteria isolated from sewage and surface water. Environ Sci Pollut Res 23(11): 10818-10831.
- Tello A, Austin B, Telfer TC (2012) Selective Pressure of Antibiotic Pollution on Bacteria of Importance to Public Health. Environ Health Perspect 120(8): 1100-1106.
- Ling TK, Xiong J, Yu Y (2006) Multicenter antimicrobial susceptibility survey of Gram-negative bacteria isolated from patients with community-acquired infections in the People’s Republic of China. Antimicrob Agents Chemother 50(1): 374-378.
- Thomas S, Holger V, Slike K, Wolfagang K, Katja S, et al. (2007) Detection of antibiotic- resistant bacteria and their resistance genes in waster, surface water and drinking water biofilms. FEMS Microb Ecol 43(3): 325-335.
- Dosso M, Bissagnene E, Coulibaly M (2000) Acquired Resistances and Prescriptions of Antibiotics in Africa: What Adaptations? Medecine et Maladies Infectieuses 30: 197-204.
- Fuentefria DB, Ferreira AE, Corção G (2011) Antibiotic-resistant Pseudomonas aeruginosa from hospital wastewater and superficial water: Are they genetically related? J Environ Manage 92(1): 250-255.
- Ekhaise FO, Omavwoya BP (2008) Influence of wastewater discharged from university of Benin reaching Hospital (UBTH), Benin City on its receiving environment. Am Eurasian J Agric Environ Sci 4(4): 484-488.
- Jarvis WR, Jarvis AA, Chinn RY (2012) National Prevalence of Methicillin-Resistant Staphylococcus aureus in Inpatients at United States Health Care Facilities, 2010 40(3): 194-200.
- Hatcher SM, Myers KW, Heaney CD, Larsen J, Hall D, et al. (2016) Occurrence of methicillin-resistant Staphylococcus aureus in surface waters near industrial hog operation spray fields. Sci Total Environ 565: 1028-1036.
- Salou M, Yehadji D, Ekouevi K, Dossim S, Tsogou C, Nyasenu YT (2014) Ciprofloxacin Sensitivity of Staphylococcus Strains Isolated at the Sylvanus Olympio University Hospital, Togo. Pharmacology & Pharmacy 5(13): 1143-1147.
- Blanch AR, Caplin JL, Iversen A, Ku¨hn I, Manero A, et al. (2003) Comparison of enterococcal populations related to urban and hospital wastewater in various climatic and geographic European regions. J Appl Microbiol 94(6): 994-1002.
- Bradley CR, Fraise AP (1996) Heat and chemical resistance of enterococci. J Hosp Infect 34(3): 191-196.
- Hidron AI, Edwards JR, Patel J, Horan TC, Sievert DM, et al. (2008) NHSN annual update: antimicrobial-resistant pathogens associated with healthcare-associated infec-tions: annual summary of data reported to the National Healthcare Safety Network at the Centers for Disease Control and Prevention, 2006-2007. Infect Control Hosp Epidemiol 29(11): 996-1011.






























